Abstract
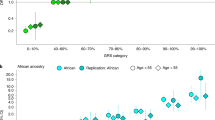
We report a prostate cancer genome-wide association follow-on study. We discovered four variants associated with susceptibility to prostate cancer in several European populations: rs10934853[A] (OR = 1.12, P = 2.9 × 10−10) on 3q21.3; two moderately correlated (r2 = 0.07) variants, rs16902094[G] (OR = 1.21, P = 6.2 × 10−15) and rs445114[T] (OR = 1.14, P = 4.7 × 10−10), on 8q24.21; and rs8102476[C] (OR = 1.12, P = 1.6 × 10−11) on 19q13.2. We also refined a previous association signal on 11q13 with the SNP rs11228565[A] (OR = 1.23, P = 6.7 × 10−12). In a multivariate analysis using 22 prostate cancer risk variants typed in the Icelandic population, we estimated that carriers in the top 1.3% of the risk distribution are at a 2.5 times greater risk of developing the disease than members of the general population.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat. Genet. 39, 631–637 (2007).
Haiman, C.A. et al. Multiple regions within 8q24 independently affect risk for prostate cancer. Nat. Genet. 39, 638–644 (2007).
Gudmundsson, J. et al. Two variants on chromosome 17 confer prostate cancer risk, and the one in TCF2 protects against type 2 diabetes. Nat. Genet. 39, 977–983 (2007).
Eeles, R.A. et al. Multiple newly identified loci associated with prostate cancer susceptibility. Nat. Genet. 40, 316–321 (2008).
Thomas, G. et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat. Genet. 40, 310–315 (2008).
Gudmundsson, J. et al. Common sequence variants on 2p15 and Xp11.22 confer susceptibility to prostate cancer. Nat. Genet. 40, 281–283 (2008).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat. Genet. 39, 645–649 (2007).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Amundadottir, L.T. et al. A common variant associated with prostate cancer in European and African populations. Nat. Genet. 38, 652–658 (2006).
Tomlinson, I. et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat. Genet. 39, 984–988 (2007).
Zanke, B.W. et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on chromosome 8q24. Nat. Genet. 39, 989–994 (2007).
Haiman, C.A. et al. A common genetic risk factor for colorectal and prostate cancer. Nat. Genet. 39, 954–956 (2007).
Kiemeney, L.A. et al. Sequence variant on 8q24 confers susceptibility to urinary bladder cancer. Nat. Genet. 40, 1307–1312 (2008).
Ghoussaini, M. et al. Multiple loci with different cancer specificities within the 8q24 gene desert. J. Natl. Cancer Inst. 100, 962–966 (2008).
Duggan, D. et al. Two genome-wide association studies of aggressive prostate cancer implicate putative prostate tumor suppressor gene DAB2IP. J. Natl. Cancer Inst. 99, 1836–1844 (2007).
Kote-Jarai, Z. et al. Multiple novel prostate cancer predisposition loci confirmed by an international study: the PRACTICAL Consortium. Cancer Epidemiol. Biomarkers Prev. 17, 2052–2061 (2008); erratum 17, 2901 (2008).
Rafnar, T. et al. Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat. Genet. 41, 221–227 (2009).
Sun, J. et al. Evidence for two independent prostate cancer risk-associated loci in the HNF1B gene at 17q12. Nat. Genet. 40, 1153–1155 (2008).
Sun, J. et al. Sequence variants at 22q13 are associated with prostate cancer risk. Cancer Res. 69, 10–15 (2009).
Kutyavin, I.V. et al. A novel endonuclease IV post-PCR genotyping system. Nucleic Acids Res. 34, e128 (2006).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Gretarsdottir, S. et al. The gene encoding phosphodiesterase 4D confers risk of ischemic stroke. Nat. Genet. 35, 131–138 (2003).
Falk, C.T. & Rubinstein, P. Haplotype relative risks: an easy reliable way to construct a proper control sample for risk calculations. Ann. Hum. Genet. 51, 227–233 (1987).
Mantel, N. & Haenszel, W. Statistical aspects of the analysis of data from retrospective studies of disease. J. Natl. Cancer Inst. 22, 719–748 (1959).
Styrkarsdottir, U. et al. Multiple genetic loci for bone mineral density and fractures. N. Engl. J. Med. 358, 2355–2365 (2008).
Pe'er, I. et al. Evaluating and improving power in whole-genome association studies using fixed marker sets. Nat. Genet. 38, 663–667 (2006).
Nicolae, D.L. Testing untyped alleles (TUNA)-applications to genome-wide association studies. Genet. Epidemiol. 30, 718–727 (2006).
Zaitlen, N., Kang, H.M., Eskin, E. & Halperin, E. Leveraging the HapMap correlation structure in association studies. Am. J. Hum. Genet. 80, 683–691 (2007).
Zheng, S.L. et al. Cumulative association of five genetic variants with prostate cancer. N. Engl. J. Med. 358, 910–919 (2008).
Stefansson, H. et al. A common inversion under selection in Europeans. Nat. Genet. 37, 129–137 (2005).
Acknowledgements
We thank the individuals who participated in the study and whose contribution made this work possible. This project was funded in part by contract number 202059 (PROMARK) from the Seventh Framework Program of the European Union to deCODE Genetics (T.R. and L.A.K.), in part by FP7-MC-IAPP Grant agreement no. 218071 (CancerGene) to deCODE Genetics, in part by a V Foundation award and US Department of Veterans Affairs grants to J.R.S., and in part by Academy of Finland, Sigrid Juselius Foundation, Finnish Cancer Organisations and the Competitive Research Funding of the Pirkanmaa Hospital District, Tampere University Hospital, to J.S.
Author information
Authors and Affiliations
Contributions
The study was designed and results were interpreted by J. Gudmundsson, P.S., A.K., U.T., T.R. and K.S. Statistical analysis was carried out by P.S., D.F.G., J. Gudmundsson, M.L.F. and A.K. Subject recruitment, biological material collection and handling along with genotyping were supervised and carried out by J. Gudmundsson, B.A.A., K.R.B., T.B., A.G., D.N.M., G.O., M.J., S.N.S., A.S., T.W., T.T., J.P.B., K.M.M., K.M.B., B.S., J. Godino, S.N., F.F., L.M., E.P., K.K.A., I.M.v.O., B.K.S., B.T.H., D.K., C.Z., K.K., J.R.G., G.V.E., E.J., W.J.C., J.I.M., L.A.K., J.R.S., J.S., R.B.B., U.T. and T.R. Authors J. Gudmundsson, P.S., D.F.G., T.R. and K.S. drafted the manuscript. All authors contributed to the final version of the paper. Principal investigators and corresponding authors for the respective replication study populations are: The Netherlands, L.A.K.; Spain, J.I.M.; Chicago, W.J.C.; Nashville, Tenn., USA, J.R.S.; Finland, J.S.
Corresponding authors
Ethics declarations
Competing interests
The authors from deCODE in Reykjavik, Iceland are shareholders in deCODE Genetics, Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–6, Supplementary Figure 1 and Supplementary Note. (PDF 223 kb)
Rights and permissions
About this article
Cite this article
Gudmundsson, J., Sulem, P., Gudbjartsson, D. et al. Genome-wide association and replication studies identify four variants associated with prostate cancer susceptibility. Nat Genet 41, 1122–1126 (2009). https://doi.org/10.1038/ng.448
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.448
This article is cited by
-
Association between interleuleukin-1β polymorphism (rs16944) and biomarkers levels in Iraqi patients with prostate cancer
Molecular Biology Reports (2023)
-
Genetic variant located on chromosome 17p12 contributes to prostate cancer onset and biochemical recurrence
Scientific Reports (2022)
-
Extensive germline-somatic interplay contributes to prostate cancer progression through HNF1B co-option of TMPRSS2-ERG
Nature Communications (2022)
-
Prostate cancer risk variants of the HOXB genetic locus
Scientific Reports (2021)
-
HNF1B-mediated repression of SLUG is suppressed by EZH2 in aggressive prostate cancer
Oncogene (2020)