Abstract
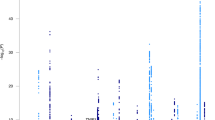
We undertook a two-stage genome-wide association study (GWAS) of Alzheimer's disease (AD) involving over 16,000 individuals, the most powerful AD GWAS to date. In stage 1 (3,941 cases and 7,848 controls), we replicated the established association with the apolipoprotein E (APOE) locus (most significant SNP, rs2075650, P = 1.8 × 10−157) and observed genome-wide significant association with SNPs at two loci not previously associated with the disease: at the CLU (also known as APOJ) gene (rs11136000, P = 1.4 × 10−9) and 5′ to the PICALM gene (rs3851179, P = 1.9 × 10−8). These associations were replicated in stage 2 (2,023 cases and 2,340 controls), producing compelling evidence for association with Alzheimer's disease in the combined dataset (rs11136000, P = 8.5 × 10−10, odds ratio = 0.86; rs3851179, P = 1.3 × 10−9, odds ratio = 0.86).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Accession codes
Change history
28 September 2009
NOTE: In the version of this article initially published, the name of the first author of reference 12 was stated incorrectly in the reference list. The correct reference is: “Lambert, J.-C. et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer's disease. Nat. Genet. advance online publication, doi:10.1038/ng.439 (6 September 2009).” The error has been corrected in the HTML and PDF versions of the article.
09 May 2013
In the version of this article initially published, Reinhard Heun was not included in the author list. This has been corrected in the HTML and PDF versions of the article.
References
Avramopoulos, D. Genetics of Alzheimer's disease: recent advances. Genome Med. 1, 34 (2009).
Abraham, R. et al. A genome-wide association study for late-onset Alzheimer's disease using DNA pooling. BMC Med. Genomics 1, 44 (2008).
Beecham, G.W. et al. Genome-wide association study implicates a chromosome 12 risk locus for late-onset Alzheimer disease. Am. J. Hum. Genet. 84, 35–43 (2009).
Bertram, L. et al. Genome-wide association analysis reveals putative Alzheimer's disease susceptibility loci in addition to APOE. Am. J. Hum. Genet. 83, 623–632 (2008).
Carrasquillo, M.M. et al. Genetic variation in PCDH11X is associated with susceptibility to late-onset Alzheimer's disease. Nat. Genet. 41, 192–198 (2009).
Coon, K.D. et al. A high-density whole-genome association study reveals that APOE is the major susceptibility gene for sporadic late-onset Alzheimer's disease. J. Clin. Psychiatry 68, 613–618 (2007).
Grupe, A. et al. Evidence for novel susceptibility genes for late-onset Alzheimer's disease from a genome-wide association study of putative functional variants. Hum. Mol. Genet. 16, 865–873 (2007).
Li, H. et al. Candidate single-nucleotide polymorphisms from a genomewide association study of Alzheimer disease. Arch. Neurol. 65, 45–53 (2008).
Reiman, E.M. et al. GAB2 alleles modify Alzheimer's risk in APOE epsilon4 carriers. Neuron 54, 713–720 (2007).
WTCCC. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Lambert, J.-C. et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer's disease. Nat. Genet. advance online publication, doi:10.1038/ng.439 (6 September 2009).
Tycko, B. et al. Polymorphisms in the human apolipoprotein-J/clusterin gene: ethnic variation and distribution in Alzheimer's disease. Hum. Genet. 98, 430–436 (1996).
Cousin, M.A. & Robinson, P.J. The dephosphins: dephosphorylation by calcineurin triggers synaptic vesicle endocytosis. Trends Neurosci. 24, 659–665 (2001).
Hiesberger, T. et al. Direct binding of Reelin to VLDL receptor and ApoE receptor 2 induces tyrosine phosphorylation of disabled-1 and modulates tau phosphorylation. Neuron 24, 481–489 (1999).
Jenne, D.E. & Tschopp, J. Clusterin: the intriguing guises of a widely expressed glycoprotein. Trends Biochem. Sci. 17, 154–159 (1992).
Jones, S.E. & Jomary, C. Clusterin. Int. J. Biochem. Cell Biol. 34, 427–431 (2002).
Calero, M. et al. Apolipoprotein J (clusterin) and Alzheimer's disease. Microsc. Res. Tech. 50, 305–315 (2000).
Giannakopoulos, P. et al. Possible neuroprotective role of clusterin in Alzheimer's disease: a quantitative immunocytochemical study. Acta Neuropathol. 95, 387–394 (1998).
Liang, W.S. et al. Altered neuronal gene expression in brain regions differentially affected by Alzheimer's disease: a reference data set. Physiol. Genomics 33, 240–256 (2008).
McGeer, P.L., Kawamata, T. & Walker, D.G. Distribution of clusterin in Alzheimer brain tissue. Brain Res. 579, 337–341 (1992).
Ghiso, J. et al. The cerebrospinal-fluid soluble form of Alzheimer's amyloid beta is complexed to SP-40,40 (apolipoprotein J), an inhibitor of the complement membrane-attack complex. Biochem. J. 293, 27–30 (1993).
Golabek, A., Marques, M.A., Lalowski, M. & Wisniewski, T. Amyloid beta binding proteins in vitro and in normal human cerebrospinal fluid. Neurosci. Lett. 191, 79–82 (1995).
Zlokovic, B.V. et al. Brain uptake of circulating apolipoproteins J and E complexed to Alzheimer's amyloid beta. Biochem. Biophys. Res. Commun. 205, 1431–1437 (1994).
Zlokovic, B.V. et al. Glycoprotein 330/megalin: probable role in receptor-mediated transport of apolipoprotein J alone and in a complex with Alzheimer disease amyloid beta at the blood-brain and blood-cerebrospinal fluid barriers. Proc. Natl. Acad. Sci. USA 93, 4229–4234 (1996).
Bales, K.R. et al. Apolipoprotein E is essential for amyloid deposition in the APP(V717F) transgenic mouse model of Alzheimer's disease. Proc. Natl. Acad. Sci. USA 96, 15233–15238 (1999).
Castano, E.M. et al. Fibrillogenesis in Alzheimer's disease of amyloid beta peptides and apolipoprotein E. Biochem. J. 306, 599–604 (1995).
Ma, J., Yee, A., Brewer, H.B. Jr., Das, S. & Potter, H. Amyloid-associated proteins alpha 1-antichymotrypsin and apolipoprotein E promote assembly of Alzheimer beta-protein into filaments. Nature 372, 92–94 (1994).
Boggs, L.N. et al. Clusterin (Apo J) protects against in vitro amyloid-beta (1–40) neurotoxicity. J. Neurochem. 67, 1324–1327 (1996).
DeMattos, R.B. et al. Clusterin promotes amyloid plaque formation and is critical for neuritic toxicity in a mouse model of Alzheimer's disease. Proc. Natl. Acad. Sci. USA 99, 10843–10848 (2002).
Lambert, M.P. et al. Diffusible, nonfibrillar ligands derived from Abeta1–42 are potent central nervous system neurotoxins. Proc. Natl. Acad. Sci. USA 95, 6448–6453 (1998).
Matsubara, E., Soto, C., Governale, S., Frangione, B. & Ghiso, J. Apolipoprotein J and Alzheimer's amyloid beta solubility. Biochem. J. 316, 671–679 (1996).
Oda, T. et al. Clusterin (apoJ) alters the aggregation of amyloid beta-peptide (A beta 1–42) and forms slowly sedimenting A beta complexes that cause oxidative stress. Exp. Neurol. 136, 22–31 (1995).
DeMattos, R.B. et al. ApoE and clusterin cooperatively suppress Abeta levels and deposition: evidence that ApoE regulates extracellular Abeta metabolism in vivo. Neuron 41, 193–202 (2004).
Bell, R.D. et al. Transport pathways for clearance of human Alzheimer's amyloid beta-peptide and apolipoproteins E and J in the mouse central nervous system. J. Cereb. Blood Flow Metab. 27, 909–918 (2007).
Bertrand, P., Poirier, J., Oda, T., Finch, C.E. & Pasinetti, G.M. Association of apolipoprotein E genotype with brain levels of apolipoprotein E and apolipoprotein J (clusterin) in Alzheimer disease. Brain Res. Mol. Brain Res. 33, 174–178 (1995).
Dreyling, M.H. et al. The t(10;11)(p13;q14) in the U937 cell line results in the fusion of the AF10 gene and CALM, encoding a new member of the AP-3 clathrin assembly protein family. Proc. Natl. Acad. Sci. USA 93, 4804–4809 (1996).
Tebar, F., Bohlander, S.K. & Sorkin, A. Clathrin assembly lymphoid myeloid leukemia (CALM) protein: localization in endocytic-coated pits, interactions with clathrin, and the impact of overexpression on clathrin-mediated traffic. Mol. Biol. Cell 10, 2687–2702 (1999).
Yao, P.J., Petralia, R.S., Bushlin, I., Wang, Y. & Furukawa, K. Synaptic distribution of the endocytic accessory proteins AP180 and CALM. J. Comp. Neurol. 481, 58–69 (2005).
Harel, A., Wu, F., Mattson, M.P., Morris, C.M. & Yao, P.J. Evidence for CALM in directing VAMP2 trafficking. Traffic 9, 417–429 (2008).
Masliah, E. et al. Altered expression of synaptic proteins occurs early during progression of Alzheimer's disease. Neurology 56, 127–129 (2001).
Fitzjohn, S.M. et al. Age-related impairment of synaptic transmission but normal long-term potentiation in transgenic mice that overexpress the human APP695SWE mutant form of amyloid precursor protein. J. Neurosci. 21, 4691–4698 (2001).
Nordstedt, C., Caporaso, G.L., Thyberg, J., Gandy, S.E. & Greengard, P. Identification of the Alzheimer beta/A4 amyloid precursor protein in clathrin-coated vesicles purified from PC12 cells. J. Biol. Chem. 268, 608–612 (1993).
Carey, R.M., Balcz, B.A., Lopez-Coviella, I. & Slack, B.E. Inhibition of dynamin-dependent endocytosis increases shedding of the amyloid precursor protein ectodomain and reduces generation of amyloid beta protein. BMC Cell Biol. 6, 30 (2005).
Koo, E.H. & Squazzo, S.L. Evidence that production and release of amyloid beta-protein involves the endocytic pathway. J. Biol. Chem. 269, 17386–17389 (1994).
Cirrito, J.R. et al. Endocytosis is required for synaptic activity-dependent release of amyloid-beta in vivo. Neuron 58, 42–51 (2008).
Göring, H.H., Terwilliger, J.D. & Blangero, J. Large upward bias in estimation of locus-specific effects from genomewide scans. Am. J. Hum. Genet. 69, 1357–1369 (2001).
Zeggini, E. et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 40, 638–645 (2008).
Barrett, J.C., Fry, B., Maller, J. & Daly, M.J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Teo, Y.Y. et al. A genotype calling algorithm for the Illumina BeadArray platform. Bioinformatics 23, 2741–2746 (2007).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Moskvina, V., Craddock, N., Holmans, P., Owen, M.J. & O'Donovan, M.C. Effects of differential genotyping error rate on the type I error probability of case-control studies. Hum. Hered. 61, 55–64 (2006).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Price, A.L. et al. Long-range LD can confound genome scans in admixed populations. Am. J. Hum. Genet. 83, 132–139, author reply 135–139 (2008).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Schlesselman, J.J. Case-Control Studies: Design, Conduct, Analysis (Oxford University Press, Oxford, UK, 1982).
Moskvina, V. & Schmidt, K.M. On multiple-testing correction in genome-wide association studies. Genet. Epidemiol. 32, 567–573 (2008).
Conde, L. et al. PupaSuite: finding functional single nucleotide polymorphisms for large-scale genotyping purposes. Nucleic Acids Res. 34, W621–W625 (2006).
Acknowledgements
We thank the individuals and families who took part in this research. Cardiff University was supported by the Wellcome Trust, Medical Research Council (MRC, UK), Alzheimer's Research Trust (ART) and the Welsh Assembly Government. ART supported sample collections at the Institute of Psychiatry, the South West Dementia Bank and the Universities of Cambridge, Nottingham, Manchester and Belfast. The Belfast group acknowledges support from the Alzheimer's Society, Ulster Garden Villages, Northern Ireland Research and Development Office and the Royal College of Physicians–Dunhill Medical Trust. The MRC and Mercer's Institute for Research on Ageing supported the Trinity College group. The South West Dementia Brain Bank acknowledges support from Bristol Research into Alzheimer's and Care of the Elderly. The Charles Wolfson Charitable Trust supported the Oxford Project to Investigate Memory and Ageing (OPTIMA) group. A.A.-C. and C.E.S. thank the Motor Neurone Disease Association and MRC for support. D.C.R. is a Wellcome Trust Senior Clinical Research Fellow. Washington University was funded by US National Institutes of Health (NIH) grants, the Barnes Jewish Foundation and the Charles and Joanne Knight Alzheimer's Research Initiative. The Mayo GWAS was supported by NIH grants, the Robert and Clarice Smith and Abigail Van Buren AD Research Program, and the Palumbo Professorship in AD Research. Patient recruitment for the MRC Prion Unit/University College London Department of Neurodegenerative Disease collection was supported by the UCL Hospital/UCL Biomedical Centre. London and the South East Region (LASER)-AD was funded by Lundbeck. The Bonn group was supported by the German Federal Ministry of Education and Research (BMBF), Competence Network Dementia and Competence Network Degenerative Dementia, and by the Alfried Krupp von Bohlen und Halbach-Stiftung. The Kooperative gesundheitsforschung in der region Augsburg (KORA) F4 studies were financed by Helmholtz Zentrum München, the German Research Center for Environmental Health, BMBF, the German National Genome Research Network and the Munich Center of Health Sciences. The Heinz Nixdorf Recall cohort was funded by the Heinz Nixdorf Foundation (G. Schmidt, chairman) and BMBF. Coriell Cell Repositories is supported by the US National Institute of Neurological Disorders and Stroke and the Intramural Research Program (IRP) of the National Institute on Aging (NIA). Work on this sample was supported in part by the IRP of the NIA, National Institutes of Health, Department of Health and Human Services; Z01 AG000950-06. We acknowledge use of DNA from the 1958 Birth Cohort collection, funded by the MRC and the Wellcome Trust, which was genotyped by the Wellcome Trust Case Control Consortium and the Type-1 Diabetes Genetics Consortium, sponsored by the US National Institute of Diabetes and Digestive and Kidney Diseases, National Institute of Allergy and Infectious Diseases, National Human Genome Research Institute, National Institute of Child Health and Human Development and Juvenile Diabetes Research Foundation International. The Antwerp site was supported by the VIB Genetic Service Facility, the Biobank of the Institute Born-Bunge, the Special Research Fund of the University of Antwerp, the Fund for Scientific Research-Flanders, the Foundation for Alzheimer Research and the Interuniversity Attraction Poles program P6/43 of the Belgian Federal Science Policy Office. K.S. is a postdoctoral fellow and K.B. a PhD fellow (Fund for Scientific Research-Flanders). We thank R. Brown, J. Landers, D. Warden, D. Lehmann, N. Leigh, J. Uphill, J. Beck, T. Campbell, S. Klier, G. Adamson, J. Wyatt, M.L. Perez, T. Meitinger, P. Lichtner, G. Eckstein, N. Graff-Radford, R. Petersen, D. Dickson, G. Fischer, H. Bickel, H. Jahn, H. Kaduszkiewicz, C. Luckhaus, S. Riedel-Heller, S. Wolf, S. Weyerer, the Helmholtz Zentrum München genotyping staff, E. Reiman, TGEN and the NIMH AD Genetics Initiative. We thank Advanced Research Computing @Cardiff (ARCCA), which facilitated data analysis.
Author information
Authors and Affiliations
Contributions
J. Williams, M.J.O. and M.O. directed this study, assisted by R.A., P.H. and P.A.H. D.H. and J. Williams took primary responsibility for drafting the manuscript assisted by R.A., P.H., R.S., A.G., M.O. and M.J.O. J. Williams, R.A., P.H., R.S., A.G., K.D., A.W., N.J., C.T., A.S., A.R.M., S. Lovestone, J.P., P.P., M.K.L., C.B., D.C.R., M.G., B.L., A.L., K. Morgan, K.S.B., P.A.P., D.C., B.M., S.T., C.H., D.M., A.D.S., S. Love, P.G.K., J.H., S. Mead, N.F., M.R., J.C., W.M., F.J., B.S., R.H., H.v.d.B., I.H., J.K., J. Wiltfang, M.D., L.F., A.M.G., J.S.K.K., C.C., P.N., J.C.M., K. Mayo, G.L., N.J.B., H.G. and A.M. contributed towards clinical sample collection, ascertainment, diagnosis and preparation of samples from the '610 group' from the stage 1 'discovery sample' and in some cases also provided stage 2 'follow up' samples. R.A. and P.H. were responsible for the coordination, collection, transit and selection of samples for genotyping from the '610 group'. R. Gwilliam and P.D. were responsible for procedures related to genotyping the 610 group on the Illumina platform. A.A.-C., C.E.S., A.B.S., R. Guerreiro, T.W.M., M.M.N., S.M., K.-H.J., N.K., H.-E.W., M.M.C., V.S.P., S.G.Y., H.H., D.R. and M.H. were involved in clinical sample collection, ascertainment, diagnosis, preparation of samples and genotyping of 'collaborative samples' included in Stage 1 (i.e. samples other than the '610 group'). K.S., K.B., S.E., P.P.D.D., C.v.B. and M.T. contributed towards sample collection, diagnosis and preparation of case-control material for the stage 2 'replication sample'. Replication genotyping was coordinated and performed by R.A., assisted by R.S. and A.G. and J.S.P. developed the database for the GWA project in which the data were stored, under the supervision of J. Williams. D.H. completed statistical quality control and produced association statistics, under the supervision of J. Williams, M.L.H., V.M. and P.A.H. All authors discussed the results and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have applied for a patent based on the results of this research.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1,3 and 4, Supplementary Note (PDF 544 kb)
Supplementary Table 2
SNPs showing association with AD (P ≤ 1×10−3) in the GWAS. (XLS 258 kb)
Supplementary Table 5
Results for SNPs highlighted by previous GWA studies in our sample (XLS 37 kb)
Supplementary Table 6
SNPs showing association with AD (P ≤ 1×10−3) in the APOE-ε4 positive sample (XLS 76 kb)
Supplementary Table 7
SNPs showing association with AD (P ≤ 1×10−3) in the APOE-ε4 negative sample (XLS 76 kb)
Rights and permissions
About this article
Cite this article
Harold, D., Abraham, R., Hollingworth, P. et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer's disease. Nat Genet 41, 1088–1093 (2009). https://doi.org/10.1038/ng.440
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.440
This article is cited by
-
Revealing third-order interactions through the integration of machine learning and entropy methods in genomic studies
BioData Mining (2024)
-
A mediation analysis of the role of total free fatty acids on pertinence of gut microbiota composition and cognitive function in late life depression
Lipids in Health and Disease (2024)
-
Trends in the application of “omics” to Alzheimer’s disease: a bibliometric and visualized study
Neurological Sciences (2024)
-
Simple model systems reveal conserved mechanisms of Alzheimer’s disease and related tauopathies
Molecular Neurodegeneration (2023)
-
Alzheimer’s disease-associated complement gene variants influence plasma complement protein levels
Journal of Neuroinflammation (2023)