Abstract
Recent work has hinted at the linkage disequilibrium (LD)-dependent architecture of human complex traits, where SNPs with low levels of LD (LLD) have larger per-SNP heritability. Here we analyzed summary statistics from 56 complex traits (average N = 101,401) by extending stratified LD score regression to continuous annotations. We determined that SNPs with low LLD have significantly larger per-SNP heritability and that roughly half of this effect can be explained by functional annotations negatively correlated with LLD, such as DNase I hypersensitivity sites (DHSs). The remaining signal is largely driven by our finding that more recent common variants tend to have lower LLD and to explain more heritability (P = 2.38 × 10−104); the youngest 20% of common SNPs explain 3.9 times more heritability than the oldest 20%, consistent with the action of negative selection. We also inferred jointly significant effects of other LD-related annotations and confirmed via forward simulations that they jointly predict deleterious effects.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Yang, J., Lee, S.H., Goddard, M.E. & Visscher, P.M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Yang, J. et al. Genome partitioning of genetic variation for complex traits using common SNPs. Nat. Genet. 43, 519–525 (2011).
Shi, H., Kichaev, G. & Pasaniuc, B. Contrasting the genetic architecture of 30 complex traits from summary association data. Am. J. Hum. Genet. 99, 139–153 (2016).
Lee, S.H. et al. Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat. Genet. 44, 247–250 (2012).
Davis, L.K. et al. Partitioning the heritability of Tourette syndrome and obsessive compulsive disorder reveals differences in genetic architecture. PLoS Genet. 9, e1003864 (2013).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Finucane, H.K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Loh, P.-R. et al. Contrasting genetic architectures of schizophrenia and other complex diseases using fast variance-components analysis. Nat. Genet. 47, 1385–1392 (2015).
Won, H.-H. et al. Disproportionate contributions of select genomic compartments and cell types to genetic risk for coronary artery disease. PLoS Genet. 11, e1005622 (2015).
Speed, D., Hemani, G., Johnson, M.R. & Balding, D.J. Improved heritability estimation from genome-wide SNPs. Am. J. Hum. Genet. 91, 1011–1021 (2012).
Gusev, A. et al. Quantifying missing heritability at known GWAS loci. PLoS Genet. 9, e1003993 (2013).
Yang, J. et al. Genetic variance estimation with imputed variants finds negligible missing heritability for human height and body mass index. Nat. Genet. 47, 1114–1120 (2015).
Koch, C.M. et al. The landscape of histone modifications across 1% of the human genome in five human cell lines. Genome Res. 17, 691–707 (2007).
Smith, A.V., Thomas, D.J., Munro, H.M. & Abecasis, G.R. Sequence features in regions of weak and strong linkage disequilibrium. Genome Res. 15, 1519–1534 (2005).
Hussin, J.G. et al. Recombination affects accumulation of damaging and disease-associated mutations in human populations. Nat. Genet. 47, 400–404 (2015).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Bulik-Sullivan, B.K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Liu, X. et al. Functional architectures of local and distal regulation of gene expression in multiple human tissues. Am. J. Hum. Genet. 100, 605–616 (2017).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Rasmussen, M.D., Hubisz, M.J., Gronau, I. & Siepel, A. Genome-wide inference of ancestral recombination graphs. PLoS Genet. 10, e1004342 (2014).
McVean, G.A.T. et al. The fine-scale structure of recombination rate variation in the human genome. Science 304, 581–584 (2004).
Myers, S., Bottolo, L., Freeman, C., McVean, G. & Donnelly, P. A fine-scale map of recombination rates and hotspots across the human genome. Science 310, 321–324 (2005).
McVicker, G., Gordon, D., Davis, C. & Green, P. Widespread genomic signatures of natural selection in hominid evolution. PLoS Genet. 5, e1000471 (2009).
Koren, A. et al. Differential relationship of DNA replication timing to different forms of human mutation and variation. Am. J. Hum. Genet. 91, 1033–1040 (2012).
Palamara, P.F. ARGON: fast, whole-genome simulation of the discrete time Wright–Fisher process. Bioinformatics 32, 3032–3034 (2016).
Gravel, S. et al. Demographic history and rare allele sharing among human populations. Proc. Natl. Acad. Sci. USA 108, 11983–11988 (2011).
Hill, W.G. & Robertson, A. The effect of linkage on limits to artificial selection. Genet. Res. 8, 269–294 (1966).
UK10K Consortium. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015).
International Schizophrenia Consortium. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Karlsson, E.K., Kwiatkowski, D.P. & Sabeti, P.C. Natural selection and infectious disease in human populations. Nat. Rev. Genet. 15, 379–393 (2014).
Messer, P.W. SLiM: simulating evolution with selection and linkage. Genetics 194, 1037–1039 (2013).
Maruyama, T. The age of a rare mutant gene in a large population. Am. J. Hum. Genet. 26, 669–673 (1974).
Charlesworth, B., Morgan, M.T. & Charlesworth, D. The effect of deleterious mutations on neutral molecular variation. Genetics 134, 1289–1303 (1993).
Sabeti, P.C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Fu, W. et al. Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature 493, 216–220 (2013).
Kiezun, A. et al. Deleterious alleles in the human genome are on average younger than neutral alleles of the same frequency. PLoS Genet. 9, e1003301 (2013).
Eyre-Walker, A. Genetic architecture of a complex trait and its implications for fitness and genome-wide association studies. Proc. Natl. Acad. Sci. USA 107 (Suppl. 1), 1752–1756 (2010).
Agarwala, V., Flannick, J., Sunyaev, S., GoT2D Consortium & Altshuler, D. Evaluating empirical bounds on complex disease genetic architecture. Nat. Genet. 45, 1418–1427 (2013).
Zuk, O. et al. Searching for missing heritability: designing rare variant association studies. Proc. Natl. Acad. Sci. USA 111, E455–E464 (2014).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Shi, J. et al. Winner's curse correction and variable thresholding improve performance of polygenic risk modeling based on genome-wide association study summary-level data. PLoS Genet. 12, e1006493 (2016).
Pickrell, J.K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Sveinbjornsson, G. et al. Weighting sequence variants based on their annotation increases power of whole-genome association studies. Nat. Genet. 48, 314–317 (2016).
Faye, L.L., Machiela, M.J., Kraft, P., Bull, S.B. & Sun, L. Re-ranking sequencing variants in the post-GWAS era for accurate causal variant identification. PLoS Genet. 9, e1003609 (2013).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 10, e1004722 (2014).
Lee, S., Abecasis, G.R., Boehnke, M. & Lin, X. Rare-variant association analysis: study designs and statistical tests. Am. J. Hum. Genet. 95, 5–23 (2014).
Johnson, T. & Barton, N. Theoretical models of selection and mutation on quantitative traits. Philos. Trans. R. Soc. B Biol. 360, 1411–1425 (2005).
Hernandez, R.D. et al. Classic selective sweeps were rare in recent human evolution. Science 331, 920–924 (2011).
Enard, D., Messer, P.W. & Petrov, D.A. Genome-wide signals of positive selection in human evolution. Genome Res. 24, 885–895 (2014).
Charlesworth, D. Balancing selection and its effects on sequences in nearby genome regions. PLoS Genet. 2, e64 (2006).
Francioli, L.C. et al. Genome-wide patterns and properties of de novo mutations in humans. Nat. Genet. 47, 822–826 (2015).
Vahedi, G. et al. Super-enhancers delineate disease-associated regulatory nodes in T cells. Nature 520, 558–562 (2015).
Davydov, E.V. et al. Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLoS Comput. Biol. 6, e1001025 (2010).
Lee, S.H. et al. Estimation of SNP heritability from dense genotype data. Am. J. Hum. Genet. 93, 1151–1155 (2013).
Chang, C.C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Hinch, A.G. et al. The landscape of recombination in African Americans. Nature 476, 170–175 (2011).
Kong, A. et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 467, 1099–1103 (2010).
Gudbjartsson, D.F. et al. Large-scale whole-genome sequencing of the Icelandic population. Nat. Genet. 47, 435–444 (2015).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Dubois, P.C.A. et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 42, 295–302 (2010).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Teslovich, T.M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Tobacco and Genetics Consortium. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat. Genet. 42, 441–447 (2010).
Psychiatric GWAS Consortium Bipolar Disorder Working Group. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat. Genet. 43, 977–983 (2011).
Schunkert, H. et al. Large-scale association analysis identifies 13 new susceptibility loci for coronary artery disease. Nat. Genet. 43, 333–338 (2011).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Manning, A.K. et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Morris, A.P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Rietveld, C.A. et al. GWAS of 126,559 individuals identifies genetic variants associated with educational attainment. Science 340, 1467–1471 (2013).
Boraska, V. et al. A genome-wide association study of anorexia nervosa. Mol. Psychiatry 19, 1085–1094 (2014).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Perry, J.R.B. et al. Parent-of-origin-specific allelic associations among 106 genomic loci for age at menarche. Nature 514, 92–97 (2014).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Bentham, J. et al. Genetic association analyses implicate aberrant regulation of innate and adaptive immunity genes in the pathogenesis of systemic lupus erythematosus. Nat. Genet. 47, 1457–1464 (2015).
Cordell, H.J. et al. International genome-wide meta-analysis identifies new primary biliary cirrhosis risk loci and targetable pathogenic pathways. Nat. Commun. 6, 8019 (2015).
Day, F.R. et al. Large-scale genomic analyses link reproductive aging to hypothalamic signaling, breast cancer susceptibility and BRCA1-mediated DNA repair. Nat. Genet. 47, 1294–1303 (2015).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Okbay, A. et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat. Genet. 48, 624–633 (2016).
Palamara, P.F. et al. Leveraging distant relatedness to quantify human mutation and gene-conversion rates. Am. J. Hum. Genet. 97, 775–789 (2015).
Acknowledgements
We thank the research participants and employees of 23andMe for making this work possible. We thank S. Sunyaev, Y. Reshef, G. Kichaev, D. Speed, and F. Day for helpful discussions. This research has been conducted using the UK Biobank Resource (application number 16549). This research was funded by NIH grants R01 MH101244, R01 MH107649, and U01 HG009088.
Author information
Authors and Affiliations
Contributions
S.G. and A.L.P. designed experiments. S.G. performed experiments. S.G., H.K.F., N.A.F., and P.-R.L. analyzed data. S.G. and A.L.P. wrote the manuscript with assistance from H.K.F., N.A.F., P.-R.L., P.F.P., X.L., A.S., B.B.-S., B.M.N., and A.G.
Corresponding authors
Ethics declarations
Competing interests
N.A.F. is an employee of 23andMe, Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and 10–16, Supplementary Tables 5, 6, 8a, 10–16, 18–22, 24 and 25, and Supplementary Note (PDF 5807 kb)
Supplementary Tables 1–4, 7, 8b,c, 17 and 23
Supplementary Tables 1–4, 7, 8b,c, 17 and 23 (XLSX 150 kb)
Supplementary Figure 9
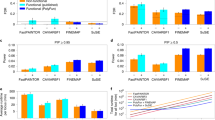
Proportion of heritability explained by the quintiles of each LD-related annotation of the baseline-LD model for each of the 62 data sets analyzed. (PDF 181 kb)
Supplementary Table 9
Effect size and enrichment of each annotation of the baseline-LD model in each of the 62 data sets analyzed. (XLSX 342 kb)
Rights and permissions
About this article
Cite this article
Gazal, S., Finucane, H., Furlotte, N. et al. Linkage disequilibrium–dependent architecture of human complex traits shows action of negative selection. Nat Genet 49, 1421–1427 (2017). https://doi.org/10.1038/ng.3954
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3954
This article is cited by
-
A distinct class of pan-cancer susceptibility genes revealed by an alternative polyadenylation transcriptome-wide association study
Nature Communications (2024)
-
Epigenetic variation impacts individual differences in the transcriptional response to influenza infection
Nature Genetics (2024)
-
Single-cell multi-ome regression models identify functional and disease-associated enhancers and enable chromatin potential analysis
Nature Genetics (2024)
-
Leveraging single-cell ATAC-seq and RNA-seq to identify disease-critical fetal and adult brain cell types
Nature Communications (2024)
-
Pleiotropy, epistasis and the genetic architecture of quantitative traits
Nature Reviews Genetics (2024)