Abstract
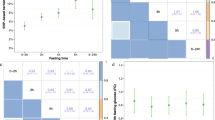
Sequence variants that affect mean fasting glucose levels do not necessarily affect risk for type 2 diabetes (T2D). We assessed the effects of 36 reported glucose-associated sequence variants1 on between- and within-subject variance in fasting glucose levels in 69,142 Icelanders. The variant in TCF7L2 that increases fasting glucose levels increases between-subject variance (5.7% per allele, P = 4.2 × 10−10), whereas variants in GCK and G6PC2 that increase fasting glucose levels decrease between-subject variance (7.5% per allele, P = 4.9 × 10−11 and 7.3% per allele, P = 7.5 × 10−18, respectively). Variants that increase mean and between-subject variance in fasting glucose levels tend to increase T2D risk, whereas those that increase the mean but reduce variance do not (r2 = 0.61). The variants that increase between-subject variance increase fasting glucose heritability estimates. Intuitively, our results show that increasing the mean and variance of glucose levels is more likely to cause pathologically high glucose levels than increase in the mean offset by a decrease in variance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Scott, R.A. et al. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat. Genet. 44, 991–1005 (2012).
Morris, A.P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Mahajan, A. et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat. Genet. 46, 234–244 (2014).
Manning, A.K. et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Voight, B.F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 42, 579–589 (2010).
Gaulton, K.J. et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat. Genet. 47, 1415–1425 (2015).
Rönnegård, L. & Valdar, W. Recent developments in statistical methods for detecting genetic loci affecting phenotypic variability. BMC Genet. 13, 63 (2012).
Wei, W.-H. et al. Major histocompatibility complex harbors widespread genotypic variability of non-additive risk of rheumatoid arthritis including epistasis. Sci. Rep. 6, 25014 (2016).
Yang, J. et al. FTO genotype is associated with phenotypic variability of body mass index. Nature 490, 267–272 (2012).
Topless, R.K. et al. Association of SLC2A9 genotype with phenotypic variability of serum urate in pre-menopausal women. Front. Genet. 6, 313 (2015).
Paré, G., Cook, N.R., Ridker, P.M. & Chasman, D.I. On the use of variance per genotype as a tool to identify quantitative trait interaction effects: a report from the Women's Genome Health Study. PLoS Genet. 6, e1000981 (2010).
Perry, G.M.L. et al. Sex modifies genetic effects on residual variance in urinary calcium excretion in rat (Rattus norvegicus). Genetics 191, 1003–1013 (2012).
Mackay, T.F. & Lyman, R.F. Drosophila bristles and the nature of quantitative genetic variation. Phil. Trans. R. Soc. Lond. B 360, 1513–1527 (2005).
Shen, X., Pettersson, M., Rönnegård, L. & Carlborg, Ö. Inheritance beyond plain heritability: variance-controlling genes in Arabidopsisthaliana. PLoS Genet. 8, e1002839 (2012).
Steinthorsdottir, V. et al. Identification of low-frequency and rare sequence variants associated with elevated or reduced risk of type 2 diabetes. Nat. Genet. 46, 294–298 (2014).
Grant, S.F. et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat. Genet. 38, 320–323 (2006).
Perry, J.R. et al. Stratifying type 2 diabetes cases by BMI identifies genetic risk variants in LAMA1 and enrichment for risk variants in lean compared to obese cases. PLoS Genet. 8, e1002741 (2012).
Cauchi, S. et al. The genetic susceptibility to type 2 diabetes may be modulated by obesity status: implications for association studies. BMC Med. Genet. 9, 45 (2008).
Azizi, F. et al. Prevention of non-communicable disease in a population in nutrition transition: Tehran Lipid and Glucose Study phase II. Trials 10, 5 (2009).
Soranzo, N. et al. Common variants at 10 genomic loci influence hemoglobin A1C levels via glycemic and nonglycemic pathways. Diabetes 59, 3229–3239 (2010).
Falconer, D.S. & Mackay, T.F.C. in Introduction to Quantitative Genetics 4th edn, Ch. 10 (Pearson, 1996).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Mitchell, R.K. et al. Selective disruption of Tcf7l2 in the pancreatic β cell impairs secretory function and lowers β cell mass. Hum. Mol. Genet. 24, 1390–1399 (2015).
Gudbjartsson, D.F. et al. Large-scale whole-genome sequencing of the Icelandic population. Nat. Genet. 47, 435–444 (2015).
Hastie, T. & Tibshirani, R. Generalized additive models. Stat. Sci. 1, 297–310 (1986).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Acknowledgements
The authors thank the subjects of the Icelandic deCODE study and the Iranian study for their participation. We also thank the staff at deCODE Genetics core facilities and all our colleagues for their contributions to this work. This research project has been supported by grant no. 121 NRCI Research Project and with the support of the National Research Council of the Islamic Republic of Iran.
Author information
Authors and Affiliations
Contributions
E.V.I., V.S., P.S., H.H., U.T., D.F.G. and K.S. designed the study and interpreted the results. E.V.I. and D.F.G. performed statistical analysis. M.S.D., G.T., S.S., A.B.H., G.S., R. Bjarnason, A.V.T., R. Benediktsson, G.E., O.S., I.O., S.Z. and F.A. performed recruitment and phenotyping. The manuscript was drafted by E.V.I., V.S., D.F.G. and K.S. All authors contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
E.V.I., V.S., G.T., P.S., H.H., S.S., U.T., D.F.G. and K.S. are employees of deCODE Genetics/Amgen, Inc.
Integrated supplementary information
Supplementary Figure 1 Mean and variance of glucose levels against age.
(a) Average of fasting and non-fasting glucose levels (mM) for all measurements on subjects belonging to a 5-year age interval (age = 20 for individuals between 16 and 20 years) against age, for both sexes. (b) Variance of fasting and non-fasting glucose levels against age, for both sexes.
Supplementary Figure 2 BMI interaction effect on fasting glucose.
The BMI interaction effect of 36 published fasting-glucose-associated variants on fasting glucose against their effect on fasting glucose between-subject variance.
Supplementary Figure 3 Effect of glucose-associated variants on mean and between-subject variance in HbA1c levels.
(a) Effect of 36 published fasting-glucose-associated variants on HbA1c and between-subject variance in HbA1c. Variants are colored blue if they significantly decrease the variance and red if they significantly increase it. (b) Effect of 36 published fasting-glucose-associated variants on HbA1c between-subject variance against fasting glucose between-subject variance.
Supplementary Figure 4 Glucose-associated variants and their effect on between-subject variance in HbA1c levels.
(a) Effect of 36 published fasting-glucose-associated variants on HbA1c between-subject variance (I) against the effect on HbA1c between-subject variance after removing all subjects with levels below 4.5 and above 6.5 (II). (b) Effect of 36 published fasting-glucose-associated variants on HbA1c between-subject variance (I) against the effect on HbA1c between-subject variance after setting values above 6.5 and below 4.5 to the respective boundary values (III).
Supplementary Figure 5 HbA1c variants and their effect on between-subject variance in HbA1c levels.
(a) Effect of eight published HbA1c-associated variants on HbA1c between-subject variance (I) against the effect on HbA1c between-subject variance after removing all subjects with levels below 4.5 and above 6.5 (II). (b) Effect of eight published HbA1c-associated variants on HbA1c between-subject variance (I) against the effect on HbA1c between-subject variance after setting values above 6.5 and below 4.5 to the respective boundary values (III).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–15 and Supplementary Note (PDF 1990 kb)
Supplementary Code
Code to detect variance effects. (TXT 3 kb)
Rights and permissions
About this article
Cite this article
Ivarsdottir, E., Steinthorsdottir, V., Daneshpour, M. et al. Effect of sequence variants on variance in glucose levels predicts type 2 diabetes risk and accounts for heritability. Nat Genet 49, 1398–1402 (2017). https://doi.org/10.1038/ng.3928
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3928
This article is cited by
-
A longitudinal genome-wide association study of bone mineral density mean and variability in the UK Biobank
Osteoporosis International (2023)
-
Risk of type 2 diabetes and KCNJ11 gene polymorphisms: a nested case–control study and meta-analysis
Scientific Reports (2022)
-
Heritability and genome-wide association analyses of fasting plasma glucose in Chinese adult twins
BMC Genomics (2020)
-
Dietary Fat and the Genetic Risk of Type 2 Diabetes
Current Diabetes Reports (2019)
-
Genome-wide Association Study of Change in Fasting Glucose over time in 13,807 non-diabetic European Ancestry Individuals
Scientific Reports (2019)