Abstract
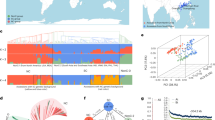
Pigeonpea (Cajanus cajan), a tropical grain legume with low input requirements, is expected to continue to have an important role in supplying food and nutritional security in developing countries in Asia, Africa and the tropical Americas. From whole-genome resequencing of 292 Cajanus accessions encompassing breeding lines, landraces and wild species, we characterize genome-wide variation. On the basis of a scan for selective sweeps, we find several genomic regions that were likely targets of domestication and breeding. Using genome-wide association analysis, we identify associations between several candidate genes and agronomically important traits. Candidate genes for these traits in pigeonpea have sequence similarity to genes functionally characterized in other plants for flowering time control, seed development and pod dehiscence. Our findings will allow acceleration of genetic gains for key traits to improve yield and sustainability in pigeonpea.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Van der Maesen, L.G.J. in The Pigeonpea (eds. Nene, Y.L., Hall, S.D. & Sheilla, V.K.) 15–46 (C.A.B. International, 1990).
Saxena, R.K. et al. Genetic diversity and demographic history of Cajanus spp. illustrated from genome-wide SNPs. PLoS One 9, e88568 (2014).
Vavilov, N.I. The origin, variation, immunity, and breeding of cultivated plants. Chron. Bot. 13, 1–366 (1951).
Varshney, R.K. et al. Draft genome sequence of pigeonpea (Cajanus cajan), an orphan legume crop of resource-poor farmers. Nat. Biotechnol. 30, 83–89 (2011).
Varshney, R.K., Thudi, M., May, G.D. & Jackson, S.A. Legume genomics and breeding. Plant Breed. Rev. 33, 257–304 (2010).
Upadhyaya, H.D. et al. Phenotyping chickpeas and pigeonpeas for adaptation to drought. Front. Physiol. 3, 179 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Clark, R.M. et al. Common sequence polymorphisms shaping genetic diversity in Arabidopsisthaliana. Science 317, 338–342 (2007).
Lam, H.M. et al. Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat. Genet. 42, 1053–1059 (2010).
Pritchard, J.K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Tang, H., Peng, J., Wang, P. & Risch, N.J. Estimation of individual admixture: analytical and study design considerations. Genet. Epidemiol. 28, 289–301 (2005).
Van der Maesen, L.J.G. Cajanus DC and Atylosia W. & A. (Leguminosae) (Agricultural University Wageningen Papers) (Wageningen Universiteit Project, 1986).
Zhou, Z. et al. Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat. Biotechnol. 33, 408–414 (2015).
Wang, Q., Tian, F., Pan, Y., Buckler, E.S. & Zhang, Z. A SUPER powerful method for genome wide association study. PLoS One 9, e107684 (2014).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Gudbjartsson, D.F. et al. Variants conferring risk of atrial fibrillation on chromosome 4q25. Nature 448, 353–357 (2007).
McPherson, R. et al. A common allele on chromosome 9 associated with coronary heart disease. Science 316, 1488–1491 (2007).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007).
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 466, 368–372 (2010).
Stefansson, H. et al. Large recurrent microdeletions associated with schizophrenia. Nature 455, 232–236 (2008).
McCarthy, S.E. et al. Microduplications of 16p11.2 are associated with schizophrenia. Nat. Genet. 41, 1223–1227 (2009).
Diskin, S.J. et al. Copy number variation at 1q21.1 associated with neuroblastoma. Nature 459, 987–991 (2009).
Horiguchi, G., Gonzalez, N., Beemster, G.T., Inzé, D. & Tsukaya, H. Impact of segmental chromosomal duplications on leaf size in the grandifolia-D mutants of Arabidopsisthaliana. Plant J. 60, 122–133 (2009).
Xiao, H., Jiang, N., Schaffner, E., Stockinger, E.J. & van der Knaap, E. A retrotransposon-mediated gene duplication underlies morphological variation of tomato fruit. Science 319, 1527–1530 (2008).
Maron, L.G. et al. Aluminum tolerance in maize is associated with higher MATE1 gene copy number. Proc. Natl. Acad. Sci. USA 110, 5241–5246 (2013).
Chia, J.M. et al. Maize HapMap2 identifies extant variation from a genome in flux. Nat. Genet. 44, 803–807 (2012).
Saxena, R.K., Edwards, D. & Varshney, R.K. Structural variations in plant genomes. Brief. Funct. Genomics 13, 296–307 (2014).
Upadhyaya, H.D. et al. Pigeonpea composite collection and identification of germplasm for use in crop improvement programmes. Plant Genet. Resour. 9, 97–108 (2011).
Meyer, R.S. & Purugganan, M.D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Weller, J.L. et al. A conserved molecular basis for photoperiod adaptation in two temperate legumes. Proc. Natl. Acad. Sci. USA 109, 21158–21163 (2012).
Tajima, F. Evolutionary relationship of DNA sequences in finite populations. Genetics 105, 437–460 (1983).
Weir, B.S. & Cockerham, C.C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Upadhyaya, H.D. & Gowda, C.L.L. Managing and Enhancing the Use of Germplasm—Strategies and Methodologies (International Crops Research Institute for the Semi-Arid Tropics, 2009).
Bradbury, P.J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635 (2007).
Xu, X. et al. Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat. Biotechnol. 30, 105–111 (2011).
Lipka, A.E. et al. GAPIT: genome association and prediction integrated tool. Bioinformatics 28, 2397–2399 (2012).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Acknowledgements
The authors are thankful to the US Agency for International Development (USAID) for providing financial support to R.K.V. The authors would like to thank A. Gafoor, B. Poornima and P. Bajaj for their support in this work. This work has been undertaken as part of the CGIAR Research Program on Grain Legumes. ICRISAT is a member of the CGIAR Consortium.
Author information
Authors and Affiliations
Contributions
R.K.V., R.K.S., Y.Y., C.K., D.K., J.K., S.A., V.K., J.-S.K. and W.Z. contributed to generation of whole-genome resequencing data. H.D.U. and R.K.V. contributed genetic material. H.D.U., G.A., K.N.Y. and S.M. performed phenotyping. R.K.V., R.K.S., H.D.U., A.W.K., C.K., A.R., D.K., J.K., S.A., J.-S.K., R.V.P., E.v.W. and S.K.D. worked on different analyses. R.K.V. and R.K.S. together with C.K., A.R., J.-S.K., R.V.P. and E.v.W. wrote and finalized the manuscript. R.K.V. and R.K.S. directed the project, and R.K.V. conceived and designed the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 10–12 and 17, and Supplementary Note (PDF 16895 kb)
Supplementary Table 1
Details on 300 Cajanus accessions (breeding lines, landraces and wild species accessions) including biological status, species, geographical region, country and state. (XLSX 20 kb)
Supplementary Table 2
Details on raw sequencing data generated on 292 Cajanus accessions. (XLSX 30 kb)
Supplementary Table 3
Identification and distribution of molecular variation (SNPs and indels) among 11 pseudomolecules CcLG01 to CcLG11 and unanchored genome sequence as CcLG0. (XLSX 21 kb)
Supplementary Table 4
Nonsynonymous-to-synonymous ratio in breeding lines, landraces and wild species in 1-Mb non-overlapping windows. (XLSX 17 kb)
Supplementary Table 5
Non-synonymous to synonymous ratio in breeding lines, landraces and wild species in 10 Kb non-overlapping windows (XLSX 1172 kb)
Supplementary Table 6
19 genomic regions (R1 to R19) showing high (>2.5) nonsynonymous-to-synonymous ratio in breeding lines, landraces and wild species in 1-Mb non-overlapping windows. (XLS 1068 kb)
Supplementary Table 7
Structural variations (CNVs and PAVs) identified in breeding lines as compared to the reference genome. (XLSX 29 kb)
Supplementary Table 8
Structural variations (CNVs and PAVs) identified in landraces as compared to the reference genome. (XLSX 24 kb)
Supplementary Table 9
Structural variations (CNVs and PAVs) identified in wild species accessions as compared to the reference genome. (XLSX 24 kb)
Supplementary Table 13
ROD values calculated during domestication (wild species versus landraces) and breeding (landraces versus breeding lines) at 10-kb non-overlapping windows. (XLSX 582 kb)
Supplementary Table 14
FST values for ROD regions with maximum values calculated in a pairwise manner for landraces versus breeding lines and wild species versus landraces. (XLSX 119 kb)
Supplementary Table 15
Genes that played an important role in domestication of crop species and their homologs in Cajanus. (XLSX 23 kb)
Supplementary Table 16
Trait phenotyping data used for GWAS. (XLSX 59 kb)
Supplementary Table 18
MTAs identified for target traits with P values. (XLSX 43 kb)
Supplementary Table 19
Number of favorable alleles identified in Cajanus accessions for detected MTAs in each trait. (XLSX 31 kb)
Supplementary Table 20
The distribution of favorable alleles for 17 MTAs detected for 100-seed weight in Cajanus accessions. (XLSX 25 kb)
Supplementary Table 21
MTAs identified in GWAS for different target traits and their corresponding structural variations (CNVs and PAVs) in breeding lines, landraces and wild species accessions. (XLSX 61 kb)
Rights and permissions
About this article
Cite this article
Varshney, R., Saxena, R., Upadhyaya, H. et al. Whole-genome resequencing of 292 pigeonpea accessions identifies genomic regions associated with domestication and agronomic traits. Nat Genet 49, 1082–1088 (2017). https://doi.org/10.1038/ng.3872
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3872
This article is cited by
-
Identification of superior haplotypes for seed protein content in pigeonpea (Cajanus cajan L.)
Journal of Plant Biochemistry and Biotechnology (2024)
-
Genetic diversity, population structure, and genome-wide association study for the flowering trait in a diverse panel of 428 moth bean (Vigna aconitifolia) accessions using genotyping by sequencing
BMC Plant Biology (2023)
-
Identification and expression analysis of SQUAMOSA promoter-binding protein (SBP) genes in mungbean
Plant Biotechnology Reports (2023)
-
Genome-wise association study identified genomic regions associated with drought tolerance in mungbean (Vigna radiata (L.) R. Wilczek)
Theoretical and Applied Genetics (2023)
-
Whole chloroplast genome-specific non-synonymous SNPs reveal the presence of substantial diversity in the pigeonpea mini-core collection
3 Biotech (2023)