Abstract
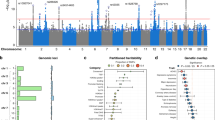
Intelligence is associated with important economic and health-related life outcomes1. Despite intelligence having substantial heritability2 (0.54) and a confirmed polygenic nature, initial genetic studies were mostly underpowered3,4,5. Here we report a meta-analysis for intelligence of 78,308 individuals. We identify 336 associated SNPs (METAL P < 5 × 10−8) in 18 genomic loci, of which 15 are new. Around half of the SNPs are located inside a gene, implicating 22 genes, of which 11 are new findings. Gene-based analyses identified an additional 30 genes (MAGMA P < 2.73 × 10−6), of which all but one had not been implicated previously. We show that the identified genes are predominantly expressed in brain tissue, and pathway analysis indicates the involvement of genes regulating cell development (MAGMA competitive P = 3.5 × 10−6). Despite the well-known difference in twin-based heritability2 for intelligence in childhood (0.45) and adulthood (0.80), we show substantial genetic correlation (rg = 0.89, LD score regression P = 5.4 × 10−29). These findings provide new insight into the genetic architecture of intelligence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
31 May 2017
In the version of this article initially published online, heritability was misspelled in the penultimate sentence of the abstract. The error has been corrected in the print, PDF and HTML versions of this article.
References
Deary, I.J. Intelligence. Annu. Rev. Psychol. 63, 453–482 (2012).
Polderman, T.J.C. et al. Meta-analysis of the heritability of human traits based on fifty years of twin studies. Nat. Genet. 47, 702–709 (2015).
Chabris, C.F. et al. Most reported genetic associations with general intelligence are probably false positives. Psychol. Sci. 23, 1314–1323 (2012).
Davies, G. et al. Genome-wide association studies establish that human intelligence is highly heritable and polygenic. Mol. Psychiatry 16, 996–1005 (2011).
Benyamin, B. et al. Childhood intelligence is heritable, highly polygenic and associated with FNBP1L. Mol. Psychiatry 19, 253–258 (2014).
Davies, G. et al. Genome-wide association study of cognitive functions and educational attainment in UK Biobank (N =112151). Mol. Psychiatry 21, 758–767 (2016).
Rietveld, C.A. et al. Common genetic variants associated with cognitive performance identified using the proxy-phenotype method. Proc. Natl. Acad. Sci. USA 111, 13790–13794 (2014).
Deary, I.J., Penke, L. & Johnson, W. The neuroscience of human intelligence differences. Nat. Rev. Neurosci. 11, 201–211 (2010).
Johnson, W., Bouchard, T.J., Krueger, R.F., McGue, M. & Gottesman, I.I. Just one g: consistent results from three test batteries. Intelligence 32, 95–107 (2004).
Ree, M.J. & Earles, J.A. The stability of g across different methods of estimation. Intelligence 15, 271–278 (1991).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Bulik-Sullivan, B.K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Yang, J. et al. Genomic inflation factors under polygenic inheritance. Eur. J. Hum. Genet. 19, 807–812 (2011).
Davies, G. et al. Genetic contributions to variation in general cognitive function: a meta-analysis of genome-wide association studies in the CHARGE consortium (N=53949). Mol. Psychiatry 20, 183–192 (2015).
Boyle, A.P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Vilhjálmsson, B.J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
de Leeuw, C.A., Mooij, J.M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Fabregat, A. et al. The Reactome pathway knowledgebase. Nucleic Acids Res. 44, D481–D487 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Newman, A.B. & Murabito, J.M. The epidemiology of longevity and exceptional survival. Epidemiol. Rev. 35, 181–197 (2013).
Willcox, B.J. et al. FOXO3A genotype is strongly associated with human longevity. Proc. Natl. Acad. Sci. USA 105, 13987–13992 (2008).
Flachsbart, F. et al. Association of FOXO3A variation with human longevity confirmed in German centenarians. Proc. Natl. Acad. Sci. USA 106, 2700–2705 (2009).
Behrens, P., Brinkmann, U. & Wellmann, A. CSE1L/CAS: its role in proliferation and apoptosis. Apoptosis 8, 39–44 (2003).
Velez Edwards, D.R. et al. Gene–environment interactions and obesity traits among postmenopausal African-American and Hispanic women in the Women's Health Initiative SHARe Study. Hum. Genet. 132, 323–336 (2013).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Willer, C.J. et al. Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 41, 25–34 (2009).
Locke, A.E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Comuzzie, A.G. et al. Novel genetic loci identified for the pathophysiology of childhood obesity in the Hispanic population. PLoS One 7, e51954 (2012).
Berndt, S.I. et al. Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat. Genet. 45, 501–512 (2013).
Wheeler, E. et al. Genome-wide SNP and CNV analysis identifies common and low-frequency variants associated with severe early-onset obesity. Nat. Genet. 45, 513–517 (2013).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Allen, N.E., Sudlow, C., Peakman, T. & Collins, R. UK Biobank data: come and get it. Sci. Transl. Med. 6, 224ed4 (2014).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Bonder, M.J. et al. Disease variants alter transcription factor levels and methylation of their binding sites. Nat. Genet. 49, 131–138 (2016).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Taskesen, E. & Reinders, M.J.T. 2D representation of transcriptomes by t-SNE exposes relatedness between human tissues. PLoS One 11, e0149853 (2016).
Acknowledgements
This work was funded by the Netherlands Organization for Scientific Research (NWO VICI 453-14-005). The analyses were carried out on the Genetic Cluster Computer, which is financed by the Netherlands Scientific Organization (NWO: 480-05-003), by VU University, Amsterdam, the Netherlands, and by the Dutch Brain Foundation and is hosted by the Dutch National Computing and Networking Services SurfSARA. This research has been conducted using the UK Biobank resource under application number 16406. We thank the participants and researchers who collected and contributed to the data.
Summary statistics have been made available for download from http://ctg.cncr.nl/software/summary_statistics.
Author information
Authors and Affiliations
Contributions
S. Sniekers performed the analyses. D.P. conceived the study. S. Stringer performed quality control on the UK Biobank data. K.W. and E.T. conducted in silico follow-up analyses. P.R.J., E.K. and J.R.I.C. conducted polygenic risk score analyses. P.K., C.A.R., D.Z., H.T., C.M.v.D., N.A., P.M., D.C., M.J., M.M., M.B.M., W.G.I., J.J.L., G.B., R.P., N.P., A.P., W.E.R.O., M.A.I. and C.F.C. contributed data. A.R.H. provided scripts for the pathway analyses. A.O. performed the educational attainment meta-analysis. S. Sniekers and D.P. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Quantile–quantile plots.
Quantile–quantile plot for SNP-based P values (top) and gene-based P values (bottom).
Supplementary Figure 2 Regional chromatin state plots for SNPs with P < 5 × 10−8 in four genomic loci.
(a–d) Chromatin state plots are included for 4 of the 18 genome-wide significant loci. The 1p31.1 and 20q13.13 loci are not included because the lead SNPs in these regions (rs66495454 and rs113315451) are indels. In each picture, the top panel shows the lead SNP (purple) and all other SNPs reaching genome-wide significance in the region. The colors represent r2 with the lead SNP. The bottom panel shows chromatin states for 127 tissue types (y axis) across the whole region. Different colors represent the different states, varying from “active TSS” (state 1) to “quiescent/low” (state 15). This information can be used to determine which SNPs to study in a functional follow-up.
Supplementary Figure 3 Regional chromatin state plots for SNPs with P < 5 × 10−8 in six genomic loci.
(a–f) Chromatin state plots are included for 6 of the 18 genome-wide significant loci.
Supplementary Figure 4 Regional chromatin state plots for SNPs with P < 5 × 10−8 in six genomic loci.
(a–f) Chromatin state plots are included for 6 of the 18 genome-wide significant loci.
Supplementary Figure 5 Predictive power (R2) of the polygenic score based on different intelligence discovery GWAS studies in four independent hold-out samples.
Comparisons of the explained variance (R2) in cognitive ability between polygenic scores based on the current meta-analysis and previous GWAS studies. The error bars represent the standard error. Cohorts: HIQ: High IQ sample; RS: Rotterdam Study; TEDS: Twins Early Development Study; ACPRC: Age and Cognitive Performance Research Centre; Discovery GWAS: Benyamin et al. 2014: childhood IQ; Davies et al. 2016: UK Biobank cognitive test (touchscreen). The R2 for HIQ is reported on the liability scale (assuming a population prevalence of 3x10-4).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Note. (PDF 2684 kb)
Supplementary Tables 1–18
Supplementary Tables 1–18. (XLSX 2999 kb)
Rights and permissions
About this article
Cite this article
Sniekers, S., Stringer, S., Watanabe, K. et al. Genome-wide association meta-analysis of 78,308 individuals identifies new loci and genes influencing human intelligence. Nat Genet 49, 1107–1112 (2017). https://doi.org/10.1038/ng.3869
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3869
This article is cited by
-
Prioritization of potential causative genes for schizophrenia in placenta
Nature Communications (2023)
-
Genome-wide association study of population-standardised cognitive performance phenotypes in a rural South African community
Communications Biology (2023)
-
Genomic patterns linked to gray matter alterations underlying working memory deficits in adults and adolescents with attention-deficit/hyperactivity disorder
Translational Psychiatry (2023)
-
Comment on “Cognitive performance protects against Alzheimer’s disease independently of educational attainment and intelligence” by Hu et al.
Molecular Psychiatry (2023)
-
Addressing the ethical and societal challenges posed by genome-wide association studies of behavioral and brain-related traits
Nature Neuroscience (2023)