Abstract
The mammalian TET enzymes catalyze DNA demethylation. While they have been intensely studied as major epigenetic regulators, little is known about their physiological roles and the extent of functional redundancy following embryo implantation. Here we define non-redundant roles for TET1 at an early postimplantation stage of the mouse embryo, when its paralogs Tet2 and Tet3 are not detectably expressed. TET1 regulates numerous genes defining differentiation programs in the epiblast and extraembryonic ectoderm. In epiblast cells, TET1 demethylates gene promoters via hydroxymethylation and maintains telomere stability. Surprisingly, TET1 represses a majority of epiblast target genes independently of methylation changes, in part through regulation of the gene encoding the transcriptional repressor JMJD8. Dysregulated gene expression in the absence of TET1 causes embryonic defects, which are partially penetrant in an inbred strain but fully lethal in non-inbred mice. Collectively, our study highlights an interplay between the catalytic and non-catalytic activities of TET1 that is essential for normal development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Gene Expression Omnibus
References
Cantone, I. & Fisher, A.G. Epigenetic programming and reprogramming during development. Nat. Struct. Mol. Biol. 20, 282–289 (2013).
Smith, Z.D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012).
Bird, A.P. & Wolffe, A.P. Methylation-induced repression—belts, braces, and chromatin. Cell 99, 451–454 (1999).
Smith, Z.D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Rossant, J. Stem cells and early lineage development. Cell 132, 527–531 (2008).
Ying, Q.L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Ficz, G. et al. FGF signaling inhibition in ESCs drives rapid genome-wide demethylation to the epigenetic ground state of pluripotency. Cell Stem Cell 13, 351–359 (2013).
Habibi, E. et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13, 360–369 (2013).
Hayashi, K., Ohta, H., Kurimoto, K., Aramaki, S. & Saitou, M. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells. Cell 146, 519–532 (2011).
Brons, I.G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Tesar, P.J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Kojima, Y. et al. The transcriptional and functional properties of mouse epiblast stem cells resemble the anterior primitive streak. Cell Stem Cell 14, 107–120 (2014).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Koh, K.P. et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8, 200–213 (2011).
Ficz, G. et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473, 398–402 (2011).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Wu, H. & Zhang, Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 156, 45–68 (2014).
Hackett, J.A. et al. Synergistic mechanisms of DNA demethylation during transition to ground-state pluripotency. Stem Cell Rep. 1, 518–531 (2013).
Sohni, A. et al. Dynamic switching of active promoter and enhancer domains regulates Tet1 and Tet2 expression during cell state transitions between pluripotency and differentiation. Mol. Cell. Biol. 35, 1026–1042 (2015).
Dawlaty, M.M. et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9, 166–175 (2011).
Yamaguchi, S. et al. Tet1 controls meiosis by regulating meiotic gene expression. Nature 492, 443–447 (2012).
Zhang, R.R. et al. Tet1 regulates adult hippocampal neurogenesis and cognition. Cell Stem Cell 13, 237–245 (2013).
Kang, J. et al. Simultaneous deletion of the methylcytosine oxidases Tet1 and Tet3 increases transcriptome variability in early embryogenesis. Proc. Natl. Acad. Sci. USA 112, E4236–E4245 (2015).
Dawlaty, M.M. et al. Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev. Cell 24, 310–323 (2013).
Dai, H.Q. et al. TET-mediated DNA demethylation controls gastrulation by regulating Lefty–Nodal signalling. Nature 538, 528–532 (2016).
Li, J. & Zhou, B.P. Activation of β-catenin and Akt pathways by Twist are critical for the maintenance of EMT associated cancer stem cell–like characters. BMC Cancer 11, 49 (2011).
Lin, H.H. et al. Neuronatin promotes neural lineage in ESCs via Ca2+ signaling. Stem Cells 28, 1950–1960 (2010).
Shen, M.M. Nodal signaling: developmental roles and regulation. Development 134, 1023–1034 (2007).
Wu, H. et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473, 389–393 (2011).
Zhou, W. et al. HIF1α induced switch from bivalent to exclusively glycolytic metabolism during ESC-to-EpiSC/hESC transition. EMBO J. 31, 2103–2116 (2012).
Tanaka, S., Kunath, T., Hadjantonakis, A.K., Nagy, A. & Rossant, J. Promotion of trophoblast stem cell proliferation by FGF4. Science 282, 2072–2075 (1998).
Rugg-Gunn, P.J., Cox, B.J., Ralston, A. & Rossant, J. Distinct histone modifications in stem cell lines and tissue lineages from the early mouse embryo. Proc. Natl. Acad. Sci. USA 107, 10783–10790 (2010).
Buecker, C. et al. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 14, 838–853 (2014).
Booth, M.J. et al. Quantitative sequencing of 5-methylcytosine and 5-hydroxymethylcytosine at single-base resolution. Science 336, 934–937 (2012).
Stadler, M.B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Shirane, K. et al. Global landscape and regulatory principles of DNA methylation reprogramming for germ cell specification by mouse pluripotent stem cells. Dev. Cell 39, 87–103 (2016).
Yu, M. et al. Base-resolution analysis of 5-hydroxymethylcytosine in the mammalian genome. Cell 149, 1368–1380 (2012).
Zalzman, M. et al. Zscan4 regulates telomere elongation and genomic stability in ES cells. Nature 464, 858–863 (2010).
Nakai-Futatsugi, Y. & Niwa, H. Zscan4 is activated after telomere shortening in mouse embryonic stem cells. Stem Cell Rep. 6, 483–495 (2016).
Lu, F., Liu, Y., Jiang, L., Yamaguchi, S. & Zhang, Y. Role of Tet proteins in enhancer activity and telomere elongation. Genes Dev. 28, 2103–2119 (2014).
Yang, J. et al. Tet enzymes regulate telomere maintenance and chromosomal stability of mouse ESCs. Cell Rep. 15, 1809–1821 (2016).
Johansson, C. et al. The roles of Jumonji-type oxygenases in human disease. Epigenomics 6, 89–120 (2014).
Kim, T.G., Kraus, J.C., Chen, J. & Lee, Y. JUMONJI, a critical factor for cardiac development, functions as a transcriptional repressor. J. Biol. Chem. 278, 42247–42255 (2003).
Rudenko, A. et al. Tet1 is critical for neuronal activity-regulated gene expression and memory extinction. Neuron 79, 1109–1122 (2013).
Liao, J. et al. Targeted disruption of DNMT1, DNMT3A and DNMT3B in human embryonic stem cells. Nat. Genet. 47, 469–478 (2015).
Cimmino, L. et al. TET1 is a tumor suppressor of hematopoietic malignancy. Nat. Immunol. 16, 653–662 (2015).
Vermeire, L. et al. Essential validation of gene trap mouse ES cell lines: a test case with the gene Ttrap. Int. J. Dev. Biol. 53, 1045–1051 (2009).
de Vree, P.J. et al. Targeted sequencing by proximity ligation for comprehensive variant detection and local haplotyping. Nat. Biotechnol. 32, 1019–1025 (2014).
Koentgen, F. et al. Exclusive transmission of the embryonic stem cell–derived genome through the mouse germline. Genesis 54, 326–333 (2016).
Meno, C. et al. Two closely-related left–right asymmetrically expressed genes, lefty-1 and lefty-2: their distinct expression domains, chromosomal linkage and direct neuralizing activity in Xenopus embryos. Genes Cells 2, 513–524 (1997).
Bachman, M. et al. 5-Hydroxymethylcytosine is a predominantly stable DNA modification. Nat. Chem. 6, 1049–1055 (2014).
Czechanski, A. et al. Derivation and characterization of mouse embryonic stem cells from permissive and nonpermissive strains. Nat. Protoc. 9, 559–574 (2014).
Taiwo, O. et al. Methylome analysis using MeDIP–seq with low DNA concentrations. Nat. Protoc. 7, 617–636 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Thomson, J.P. et al. Comparative analysis of affinity-based 5-hydroxymethylation enrichment techniques. Nucleic Acids Res. 41, e206 (2013).
Matarese, F., Carrillo-de Santa Pau, E. & Stunnenberg, H.G. 5-Hydroxymethylcytosine: a new kid on the epigenetic block? Mol. Syst. Biol. 7, 562 (2011).
Robinson, M.D. et al. Evaluation of affinity-based genome-wide DNA methylation data: effects of CpG density, amplification bias, and copy number variation. Genome Res. 20, 1719–1729 (2010).
Masser, D.R., Stanford, D.R. & Freeman, W.M. Targeted DNA methylation analysis by next-generation sequencing. J. Vis. Exp. 96, e52488 (2015).
Kumaki, Y., Oda, M. & Okano, M. QUMA: quantification tool for methylation analysis. Nucleic Acids Res. 36, W170–W175 (2008).
Krueger, F. & Andrews, S.R. Bismark: a flexible aligner and methylation caller for Bisulfite–Seq applications. Bioinformatics 27, 1571–1572 (2011).
Smallwood, S.A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Wang, Z. et al. swDMR: a sliding window approach to identify differentially methylated regions based on whole genome bisulfite sequencing. PLoS One 10, e0132866 (2015).
Yu, G., Wang, L.G. & He, Q.Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Acknowledgements
We thank G. Daley and A. Yabuuchi (Harvard Medical School) for generating the Tet1 GT mouse. SMARTer RNA–seq and amplicon sequencing were performed by K. Coeck at the VIB Nucleomics Core (KU Leuven) under the expert guidance of P. Verhasselt, R. Janky, W. Van Delm and S. Derveaux. WGBS and oxWGBS were performed at Novogene, and data were analyzed by Y.S. Li. We also thank L. Vermeire and L. Umans for technical guidance in mouse embryo manipulation and protocols for WISH, and H. Zhao for help with bioinformatic analyses. We are grateful for suggestions from M. Wilkinson and C. Verfaillie in the critique of this manuscript. This work was supported by Fonds voor Wetenschappelijk Onderzoek (FWO) Research Foundation–Flanders grants G.0C56.13N and G.0632.13N, Ministerie van de Vlaamse Gemeenschap and Marie Curie Career Integration grant PCIG-GA-2012-321658 (K.P.K.), US NIH grant R35 CA210043 (A.R.) and ERC Consolidator Grant award CHAMELEON 617595 (D.L.).
Author information
Authors and Affiliations
Contributions
R.K. performed all experiments involving mouse embryo dissection and manipulation, derivation of ESC lines, cell culture, cellular flux assays and amplicon sequencing analysis. A.S. performed RNA–seq analysis, Flow-FISH and luciferase reporter assays and generated TET1 and JMJD8 rescue cell lines. B.T. performed MeDIP, hMeDIP, mass spectrometry and integrative ChIP–seq with WGBS analysis. X.L. performed TET1 and ΔN-JMJD8 ChIP–qPCR and ChIP–seq analysis. J.V.V. performed dot blot analysis, ELISA and confocal microscopy. M.B. contributed X-gal staining of embryos and assisted in derivation of ESC lines. B.B. contributed IPA analysis. A.Z. contributed expertise on mouse dissection and WISH assays. A.R. contributed the Tet1 GT mouse strain. D.L. provided next-generation sequencing expertise and support. R.K. and A.S. wrote the methods and prepared all figures. K.P.K. conceived the study, directed the research and wrote the manuscript with assistance from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Temporal patterns of 5hmC and mapping of the insertion cassette in the Tet1 gene-trap allele.
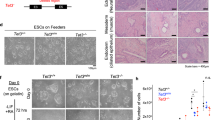
(a) 5hmC immunohistochemistry on longitudinal sections of wild-type embryos at E7.5–8.5. Scale bars, 100 μm. Epi, epiblast or embryonic ectoderm; ExE, extraembryonic tissues; VE, visceral endoderm; M, mesoderm; HF, head fold; Ch, chorion; Am, amnion. (b) Schematic representation of the Tet1 gene from exons 2 to 13 (E2 to E13; white boxes) and the position of the gene-trap cassette insertion. En2 intr1, engrailed-2 intron 1; SA, splice acceptor; β-geo, beta-galactosidase–neomycin resistance fusion gene; pA, poly(A). (c) Genome-wide coverage plot of targeted locus amplification (TLA) sequencing results confirms a single gene-trap event at the Tet1 locus on chromosome 10. (d) UCSC Genome Browser view of TLA read coverage over the Tet1 intron, between exon 2 and exon 3, at Chr10:62330560–62338166 (NCBI37/mm9). Note that the reads are mapped to the minus strand. Informative reads (black boxes) flanking regions with no coverage (deletions) or indicating fusion events with the gene-trap cassette are shown below, aligned to the genomic location. A read sequence fragmented into two parts of divergent sequence directionality (red arrows) is shown joined by a curve to indicate an inversion event. Genomic sections corresponding to all read fragments are labeled by lettered green bars shown above the track. (e) Southern blot analysis of genomic DNA harvested from wild-type (no bands detected) and RRG140 ESCs, using a probe for the β-geo sequence (position marked by the red line in h), suggesting integration of three copies of the gene-trap cassette. (f) Quantitative gene copy dosage measurement of the lacZ gene in genomic DNA harvested from the RRG140 cell line and from Tet1GT/wt mice (generation N2–3), indicating three copies of the gene-trap cassette per haploid genome. Similar results were obtained with Tet1GT/wt mice of further backcross generations N6–8 (data not shown). Error bars, s.e.m. of technical replicates of the cell line and biological replicates of mice (n = 8). (g) Sequences of fusion regions determined by Sanger sequencing of a long-range PCR product amplified using a forward primer in region A and a reverse primer in the gene-trap En2 region, confirming the fusion event at the site of insertion and a deletion between regions A and D. (h) Schematic reconstruction of the genome rearrangements at the Tet1 gene-trap insertion site inferred from the results of c–g. Arrows indicate the direction of transcription. RV, EcoRV restriction site. An asterisk refers to a possibly truncated third tandem copy of the integrated cassette of <4 kb.
Supplementary Figure 2 Expression of Tet1 in Tet1GT/wt embryos and phenotypic analyses of Tet1-deficient mouse strains.
(a) X-gal staining (blue) of preimplantation Tet1GT/wt embryos. (b) Whole-mount X-gal staining of TetGT/wt postimplantation E5.5–10.5 embryos. (c) Whole-mount X-gal staining of E5.5–10.5 Tet1GT/GT embryos from early (N ≤ 3) backcross generations. (d) Sagittal sections of stained Tet1GT/GT embryos from early (N ≤ 3) backcross generations. (e) Whole-mount X-gal staining of E8.0–9.5 Tet1GT/GT embryos from late (N ≥ 6) backcross generations. (f) Sagittal sections of stained Tet1GT/GT embryos from late (N ≥ 6) backcross generations. (g) Schematic representation of the Tet1 targeted mutation in the Tet1tm1koh line, showing the insertion of a floxed reporter cassette at the ATG start codon of exon 2 (E2). NLS, nuclear localization signal; BGHpA, bovine growth hormone polyadenylation; pA, polyadenylation. (h) Whole-mount X-gal staining of an E6.5 Tet1lacZ/wt embryo obtained from crossing wt with Tet1lacZ/wt mice. Epi, epiblast or embryonic ectoderm; ExE, extraembryonic tissues; EPC, ectoplancental cone; M, mesoderm; HF, head fold; FB, future brain; NT, neural tube; Ch, chorion; Am, amnion; Al, allantois; NE, neuroepithelium.
Supplementary Figure 3 Quality control of low-input RNA–seq and analysis of differentially expressed genes in the epiblast.
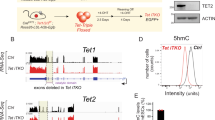
(a) Unsupervised clustering on a Spearman correlation matrix using normalized counts of all sample libraries shows segregation of epiblast (Epi) and extraembryonic ectoderm (ExE) samples. (b) RPKM of markers for epiblast (Oct4, Nanog, Otx2 and Fgf5), ExE (Tead4 and Elf5) and visceral endoderm (VE) (Afp). In Epi samples, epiblast markers (Oct4, Nanog, Otx2 and Fgf5) were detected with high read counts and trophoblast markers (Elf5 and Tead4) were absent. In ExE samples, epiblast markers were absent and trophoblast markers were highly expressed. Reads for the VE marker AFP were only detected at low levels in 4 of the 23 samples, indicating that the dissection had effectively eliminated VE cells within experimental limits. (c) RPKM of Tet1 and RPM of the bacterial β-galactosidase gene (lacZ). The dotted line represents the hypothetical threshold level for Tet1 WISH detection and for lacZ X-gal staining detection. (d) Venn diagram of differentially expressed (DE) genes in the Epi using P < 0.001. Gene names highlighted in red are those in DE1 or DE2 at FDR < 0.05 (Fig. 2b). (e,f) Methylation analysis by clonal Sanger sequencing of bisulfite conversions at Lefty2 (e) and Tet1 (f) in E6.25 wt and Tet1GT/GT Epi. The boxed regions indicate the subset further analyzed by high-throughput amplicon sequencing in Figure 2h,i. Independent biological replicate pools were used to generate Sanger (shown here) and amplicon (Fig. 2i) sequencing data at Tet1 loci. Open and filled circles represent unmethylated and methylated (5mC + 5hmC) CpGs, respectively. The percentages below indicate average methylation levels of all CpGs over the sequences analyzed. TET1 ChIP–seq tracks are binding peaks profiled in mouse ESCs18; the y-axis denotes Tet1 ChIP-seq peaks detected using an antibody generated against a C-terminal epitope normalized to IgG ChIP-seq control. Paired Student’s t test, *P < 0.05.
Supplementary Figure 4 Analysis of differentially expressed genes in the ExE.
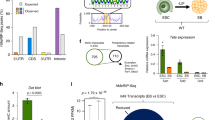
(a) Venn diagram of DE genes in the ExE using P < 0.001. Gene names highlighted in red are those in DE1 or DE2 defined using FDR < 0.05 (Fig. 3c). (b) RPKM values of selected ExE DE genes. ANOVA, *P <0.05, **P < 0.01. (c) Results of IPA analysis of DE genes between Tet1GT/GT and wt (DE1, P < 0.05), showing enrichment in the oxidative phosphorylation pathway. Genes upregulated significantly in Tet1GT/GT relative to wt are shown in shades of pink through red (higher color intensity with increasing fold change). A double border in red indicates a group or complex that shows significant collective dysregulation of constituent genes. Genes in gray show fold changes with P > 0.05. Genes in white are part of the pathway but had no fold change (P =1) in DESeq2 analysis. (d) qRT–PCR of Tet transcript copies normalized to Gapdh copies. Error bars, s.e.m. of biological replicates of wt ExE (n = 4; from Fig. 3b) and of independent culture passages of TSCs (n = 3). The dotted line indicates background levels. (e) Methylation analysis by clonal Sanger sequencing of bisulfite conversions at the Xaf1 locus in E6.25 wt and Tet1GT/GT ExE. The same bisulfite-treated DNA pool was used for high-throughput amplicon sequencing analysis of the boxed region to generate the data shown in Figure 3i. Results are presented as described in Supplementary Figure 3e,f. (f–h) Methylation analysis by amplicon sequencing of bisulfite conversions at ExE DE gene loci, indicated by bold horizontal bars below gene structure schematic. Percentages for average methylation levels of all CpGs over each sequence analyzed are shown for wt (in blue) and Tet1GT/GT (in black). Paired Student’s t test, *P < 0.05.
Supplementary Figure 5 Methylation changes in Tet1-deficient primed epiblast-like cells examined by dot blot and DNA immunoprecipitation.
(a) qRT–PCR for Tet1 in wt, het and KO ESCs detected using primers spanning exons 12–13 (left) or exons 3–4 (right). Error bars, s.e.m. of biological replicates (n = 3 cell lines). (b) Western blot analysis using a TET1 antibody. (c,d) 5mC dot blot quantification (c) and mass spectrometry measurements (d) of genomic DNA from wt, het and KO ESCs cultured in serum + LIF (SL), in ground state (2iL) and converted to EpiLCs. The red circles in c denote KO male line 12 showing higher levels of 5mC as compared to the other two KO female lines analyzed in parallel. Error bars, s.e.m. (n = 3) of biological replicate lines. (e) Unsupervised hierarchical heat map clustering of the 1,000 most variant peaks from the hMeDIP (left) and MeDIP (right) data sets of EpiLC lines. Red boxes denote KO samples. The sex of each line is indicated below the sample name. (f,g) MA plots of differentially hydroxymethylated regions (DHMRs) (f) and differentially methylated regions (DMRs) (g) in Tet1 KO EpiLCs as compared to wt or het. FDR < 0.05. (h) MA plot of DMRs at CpG islands. FDR < 0.1. (i) Normalized MeDIP–seq read counts in wt, het and KO EpiLCs (left) and distribution of MeDIP peaks gained in KO relative to all peaks (right) among high- (HCP), intermediate- (ICP) and low- (LCP) density CpG promoters. (j) MA plots of 18,762 5hmC peaks on collated hMeDIP data (left) and 1,355 5mC peaks on collated MeDIP data (right) that were significantly gained (red) and reduced (green) in KO 2iL cells compared to wt or het using FDR < 0.05. (k) MA plot of DHMRs with loss of 5hmC in KO EpiLCs superimposed over DHMRs in 2iL ESCs. (l) MA plot of DMRs with gain of 5mC in KO EpiLCs superimposed over DMRs in 2iL ESCs. het, Tet1GT/wt; KO, Tet1GT/GT.
Supplementary Figure 6 Methylation and hydroxymethylation changes in Tet1-deficient primed epiblast-like cells examined by whole-genome bisulfite and oxidative bisulfite sequencing.
(a) Percentage of methylated cytosines (mCs) among all cytosines covered by ≥5 reads in WGBS and oxWGBS. (b) Distribution of mCs in CG, CHH and CHG contexts. (c) Distribution of methylated CpG (mCpG) frequencies by 10% 5mC level bins at all mCpGs with coverage ≥10. HMR, highly methylated region; LMR, lowly methylated region; UMR, unmethylated region. (d) Distribution of methylated CHH or CHG by 5mC level bins at all mCs with coverage ≥10. (e) 5mC levels of CpGs distributed within annotated genomic features in wt (line 19) and KO (line 2) EpiLC samples. (f) Distribution of statistically validated hmCs identified in CG, CHH and CHG contexts. (g) Distribution of hmCpG frequencies by hmC level bins at mCpG sites with coverage >10. The total 5hmCpG number of each sample is shown in brackets. (h) Distribution of hmCpG frequencies by 5mC level bins at mCpG sites with coverage ≥5. The total 5hmCpG number of each sample is shown in brackets. (i) hmC levels at mCpGs distributed within annotated genomic features in wt and KO EpiLC samples. Below, a whisker plot representing 5hmC levels at CGIs for all samples. (j) Heat map showing 5mC levels within 100-bp bins spanning ±5-kb regions centred at TET1 ChIP–seq binding peaks.
Supplementary Figure 7 GO and KEGG analyses of RNA–seq, WGBS and oxWGBS data.
(a) Significantly enriched GO terms associated with up- or downregulated genes identified in RNA–seq of KO relative to wt EpiLCs. (b) List of the most significantly enriched GO terms associated with DMR-related genes from both WGBS (top) and oxWGBS (bottom) analyses. (c) Top 20 KEGG pathways associated with DMR-related genes from both WGBS (top) and oxWGBS (bottom) analyses. Dot size is proportional to the number of DMR-related genes in each pathway; colors reflect the range of q values (corrected P values) and the rich factor represents the degree of enrichment.
Supplementary Figure 8 Regulation of Zscan4 expression by TET1.
(a) Expression of Zscan4 in different cell states. Error bars, s.e.m. (n = 3 independent cultures) of each biological replicate. (b) Telomere length measurement by flow-FISH using Cy3-conjugated telomere probe in three wt and three KO ESC lines, cultured in 2iL medium for >5 passages. Percentages of cells with shorter, normal or longer telomeres are indicated with s.d. Two technical replicates are superimposed (light and dark blue lines). The values of subtelomere length fractions are shown below as the mean ± s.e.m. of biological triplicates (n = 3 cell lines). For telomere length data on EpiLCs, refer to Figure 5e,f. KO, Tet1GT/GT.
Supplementary Figure 9 Base-resolution methylation changes at TET1-bound sites and rescue of TET1 expression.
(a) IGV tracks at specific target genes, representing ChIP–seq peaks for TET1 binding and WGBS and oxWGBS methylation levels in EpiLC lines. Green bars indicate the location of CpG islands. Boxes demarcate TET1-bound regions. Regions showing the presence of 5hmC in wt EpiLCs are highlighted in blue. Regions that are lowly methylated in wt but gain methylation in KO are highlighted in red. (b) Western blot for TET1 in HEK293T cells expressing full-length TET1 from the pEF1 or PiggyBac (PB) vector. (c) 5hmC dot blot of genomic DNA extracted from HEK293T cells transfected with a PB vector expressing full-length TET1 or a mutant with base substitutions at the catalytic residues (mHxD). (d) Western blot for TET1 in ESC clones stably transfected with a PB vector expressing full-length TET1 or catalytic mutant TET1 (mHxD). The western blot shows TET1 expression with and without doxycycline induction, with leakiness of the tetO-CMV minimal promoter activity in the absence of doxycycline. HEK293T cells transfected with a PB vector containing full-length TET1, and also wt and KO ESCs, were used as controls. KO, Tet1GT/GT.
Supplementary Figure 10 Analysis of differentially expressed genes in epiblast-like cells on the CD1 background.
Heat map clustering of genes differentially expressed in three wt and three KO EpiLCs of CD1 background. The sex of each line is indicated below the sample name. (b) Overlap of DE genes from RNA–seq of EpiLCs derived on the CD1 background in comparison to data obtained from B6 lines. Gene names highlighted in red are DE genes common in B6 Epi and EpiLCs. (c) GO terms associated with DE genes in KO versus wt EpiLCs on the B6 and CD1 backgrounds. KO, Tet1GT/GT.
Supplementary Figure 11 Subcellular localization and chromatin binding of JMJD8.
(a) Western blot on subcellular fractions of HEK293T cells overexpressing C-terminal V5/His-tagged JMJD8(1–271), JMJD8(27–271) (ΔN-JMJD8) or empty pEF1-V5/His vector (mock). The purity of nuclear (N) and cytosol (Cy) fractions is indicated by lamin A and tubulin, respectively. WCL, whole-cell lysate. (b) Western blot detection of 6×His and V5 tags in ESC clones overexpressing C-terminally V5/His-tagged JMJD8(27–271) (ΔN-JMJD8; two clones) or empty vector (mock) after conversion to EpiLCs. (c) IGV tracks representing ChIP–seq peaks for JMJD8 binding at target genes. ChIP was performed using V5 or His antibodies in clone 2 expressing JMJD8(27–271) in wt EpiLC, using a mock clone as a negative control. (d) Western blot detection of epitope-tagged ΔN-JMJD8 in two independent doxycycline (DOX)-inducible clones.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Note. (PDF 3490 kb)
Supplementary Table 1
Differential expression analysis of epiblast (Epi) samples by DESeq2. (XLSX 4433 kb)
Supplementary Table 2
RPKM values of gene expression in all Epi and ExE samples. (XLSX 7201 kb)
Supplementary Table 3
Differential expression analysis of ExE samples by DESeq2. (XLSX 2772 kb)
Supplementary Table 4
List of differentially expressed genes in Tet1 KO ExE involved in mitochondrial oxidative phosphorylation. (XLSX 12 kb)
Supplementary Table 5
Differentially methylated regions in Tet1 KO EpiLC12 versus wt EpiLC15. (XLSX 2446 kb)
Supplementary Table 6
Differential expression analysis of EpiLC samples by DESeq2. (XLSX 3790 kb)
Supplementary Table 7
RPKM values of gene expression in all EpiLC samples. (XLSX 5006 kb)
Supplementary Table 8
List of primers. (XLSX 15 kb)
Supplementary Data 1
BED file of WGBS DMRs with gain of methylation in EpiLC KO12 compared to EpiLC wt15. (TXT 9 kb)
Supplementary Data 2
BED file of WGBS DMRs with loss of methylation in EpiLC KO12 compared to EpiLC wt15. (TXT 1 kb)
Supplementary Data 3
BED file of oxWGBS DMRs with gain of methylation in EpiLC KO12 compared to EpiLC wt15. (TXT 628 kb)
Supplementary Data 4
BED file of oxWGBS DMRs with loss of methylation in EpiLC KO12 compared to EpiLC wt15. (TXT 0 kb)
Rights and permissions
About this article
Cite this article
Khoueiry, R., Sohni, A., Thienpont, B. et al. Lineage-specific functions of TET1 in the postimplantation mouse embryo. Nat Genet 49, 1061–1072 (2017). https://doi.org/10.1038/ng.3868
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3868
This article is cited by
-
Epigenetic OCT4 regulatory network: stochastic analysis of cellular reprogramming
npj Systems Biology and Applications (2024)
-
Niche Tet maintains germline stem cells independently of dioxygenase activity
The EMBO Journal (2024)
-
Hypoxia switches TET1 from being tumor-suppressive to oncogenic
Oncogene (2023)
-
TET (Ten-eleven translocation) family proteins: structure, biological functions and applications
Signal Transduction and Targeted Therapy (2023)
-
Induced hepatic stem cells maintain self-renewal through the high expression of Myc coregulated by TET1 and CTCF
Cell & Bioscience (2022)