Abstract
Spider silks are the toughest known biological materials, yet are lightweight and virtually invisible to the human immune system, and they thus have revolutionary potential for medicine and industry. Spider silks are largely composed of spidroins, a unique family of structural proteins. To investigate spidroin genes systematically, we constructed the first genome of an orb-weaving spider: the golden orb-weaver (Nephila clavipes), which builds large webs using an extensive repertoire of silks with diverse physical properties. We cataloged 28 Nephila spidroins, representing all known orb-weaver spidroin types, and identified 394 repeated coding motif variants and higher-order repetitive cassette structures unique to specific spidroins. Characterization of spidroin expression in distinct silk gland types indicates that glands can express multiple spidroin types. We find evidence of an alternatively spliced spidroin, a spidroin expressed only in venom glands, evolutionary mechanisms for spidroin diversification, and non-spidroin genes with expression patterns that suggest roles in silk production.
Similar content being viewed by others
Main
More than 380 million years of evolution have produced >46,000 extant spider species, exhibiting an incredible diversity of silks used for prey capture and reproduction1,2,3. Spider silks can be stronger than steel and tougher than Kevlar, yet are much lighter weight than these manmade materials4. Silks vary in extensibility5, are temperature resilient6, can enable electrical conduction7, and can inhibit bacterial growth while being nearly invisible to the human immune system8. Thus, novel materials derived from spider silks offer tremendous potential for medical and industrial innovation. To take advantage of their desirable properties, we must learn more about spider silk genetic structure, functional diversity, and production.
A female orb-weaving spider can have up to seven morphologically differentiated types of silk glands, each believed to extrude a distinct class of silk with biophysical characteristics resulting from the expression of a unique combination of silk genes in that gland9,10. The silk classes of a typical 'gluey silk' orb-weaver (Araneoidea) female include (i) major ampullate silk, which exhibits great tensile strength and is employed in draglines, bridgelines, and web radii11,12; (ii) minor ampullate silk, used for inelastic temporary spirals during web building11,12; (iii) cement-like piriform silk that bonds fibers together and to other substrates13,14; (iv) strong, yet flexible aciniform silk used for prey wrapping and egg case insulation15; (v) tubuliform and cylindriform silk that constitutes the tough outer layer of egg cases16,17; (vi) flagelliform silk that exhibits unparalleled extensibility and is used in the capture spiral18,19; and (vii) the viscous and sticky aggregate silk that aids in prey capture20,21,22,23,24. Many spider species produce just a subset of these silk classes, and some produce yet other silk types, including cribellate silk25. Each species possesses an assortment of specialized gland types that are thought to produce distinct classes of silks to fit specific needs9,26,27.
Spider silks are composed primarily of spidroin proteins (where a 'spidroin' is a spider fibroin28,29,30,31) that, by convention, have been named and classified according to the specific silk gland in which they were first discovered. Spidroin proteins have conserved N- and C-terminal domains that flank long runs of repeated motifs32,33,34, the composition and number of which confer specific physical properties to silks27. Yet, despite decades of research on orb-weaver silks, there is incomplete knowledge of all the spidroins within an orb-weaver species.
Adding to the sampling of sequences obtained from targeted investigations, the assembly of the velvet spider (Stegodyphus mimosarum) genome yielded 19 spidroins, the largest collection from any single species27. Owing to the challenges of assembling arrays of repeats, several of the S. mimosarum spidroin sequences are incomplete, without the sequences encoding N- and C-terminal domains anchored on a single scaffold10,11,12. Furthermore, this cribellate-sheetweb-building spider lacks the flagelliform and aggregate silks found in orb webs, limiting the diversity of spidroin sequences cataloged from a single spider species. In contrast, female golden orb-weaver spiders (N. clavipes) use silks from all seven of the araneoid silk gland types35 (Supplementary Fig. 1a–c). The first spidroins were characterized from N. clavipes28,29,30,31, and this species has continued to be useful for investigating silk genes, their diversity and structure, and their evolutionary history as a gene family29. Surprisingly, the full genome of this extensively studied species, the “ubiquitous workhorse of spider research” (ref. 2), has not been reported.
We present the first annotated genome of an orb-weaving spider, cataloging 28 N. clavipes spidroins, including 8 that were previously unreported. Characterization of the repetitive sequences found in these genes has yielded numerous novel motifs and new variants of previously reported motifs9,10. Many of these motifs occur in iterated groups, and we catalog as many as 506 unique 'cassettes' that feature two to four contiguous motifs and that are themselves organized into larger repetitive units (∼200 amino acid residues) known as ensemble repeats30,36. The N. clavipes genome provides evidence for evolutionary mechanisms like tandem duplication that may underlie spidroin diversification, and our data support estimates that rapid silk evolution accompanied the emergence of the orb web ∼213 million years ago37.
We used the results of our genome-wide approach to profile transcripts from all loci in tissues isolated from N. clavipes females. Using quantitative expression analysis of the 28 N. clavipes spidroins in this spider's morphologically distinct silk glands, we have examined the idea that spiders have evolved multiple types of silk glands that produce unique combinations of silk proteins, usually with one or two spidroins dominating38,39,40,41 (Supplementary Fig. 1b). Our complete expression profile of N. clavipes spidroin transcripts across the set of silk glands reveals the fuller extent of this phenomenon. We demonstrate that a novel N. clavipes spidroin is expressed exclusively in venom glands rather than silk glands, a radical change in gene regulation. We detect alternative splicing of a spidroin transcript, a mechanism conjectured for spidroins31 but not, to our knowledge, previously shown. We also identify non-spidroin genes that are highly expressed in silk glands, suggesting these genes as candidates for further study of spider silk production.
Results
An annotated genome for N. clavipes
We sequenced genomic DNA isolated from field-collected N. clavipes females and used a combination of strategies to de novo assemble 2.44 Gb of genome (Table 1, Supplementary Tables 1–4, and Supplementary Note). The predicted size of the entire N. clavipes genome is 3.45 Gb, with 55% estimated as repetitive sequence. Our annotated meta-assembly consists of 180,236 scaffolds (N50 scaffold size, 62.9 kb; N50 contig size, 8.1 kb), with 98.5× coverage from re-mapping of over 2.48 billion 100-bp unique reads (48.9× from unique pairs; Supplementary Table 5).
To determine gene locations within the N. clavipes genome, we sequenced RNA from 16 different tissue isolates (for example, whole body, brain, and individual silk and venom glands) collected from four female individuals and then de novo assembled the transcriptome for each isolate using strand-specific 100-bp paired-end reads (Supplementary Note). We also assembled a transcriptome representing the union of all isolates (1.53 billion reads; Table 1 and Supplementary Tables 6 and 7). To quantify the completeness of the protein-coding genome, we searched our draft assemblies for homology to >2,000 curated arachnid sequences42, and we estimate that our draft genome is 94% complete and our all-isolate transcriptome assembly is 99% complete (Supplementary Tables 5 and 7).
In total, >32 million features are annotated on the N. clavipes draft genome (Supplementary Table 8), which was achieved using (i) our transcriptome from all isolates, (ii) results from two gene prediction algorithms, (iii) libraries of transposable elements and repeated motifs, and (iv) coding sequences from related species18,19 (Supplementary Table 8). Using gene modeling, we conservatively predict 14,025 genes present in the N. clavipes genome; 2,023 gene models transcribe >1 alternative spliceoform, resulting in 3,937 additional transcripts for a total of 17,962 mRNAs in the final gene set (Supplementary Table 9).
A first-generation araneoid spidroin catalog
To identify N. clavipes spidroin genes, we searched the assembled genome, transcriptomes, and annotated gene models for sequences similar to published spidroins (Supplementary Table 10), finding 28 candidates. We used long-range PCR followed by single-molecule real-time sequencing to reconstruct and validate each assembled locus at >100× coverage (Supplementary Table 11 and Supplementary Note). For 27 of the 28 spidroins, the sequences encoding the N- and C-terminal domains were connected on a single scaffold (Fig. 1). We obtained 20 complete full-length spidroin sequences, and, while gaps persist in the remaining spidroins, substantial portions of their repeated motif structures are described (Fig. 1). We note three partial spidroin-like sequences that could not be assembled, suggesting that there are additional N. clavipes spidroins yet to be characterized (Supplementary Table 12).
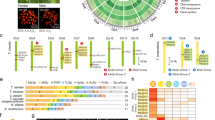
Phylogenetic tree showing the evolutionary relationship among the assembled N. clavipes spidroins using N-terminal sequences (∼130 residues) from each putative gene product (bootstrap values provided in Supplementary Fig. 3a). Circular symbols denote the silk class—with putative functional application and presumed gland of origin (Supplementary Fig. 1b,c)—of each spidroin, determined by alignment to known spidroin sequences (spidroins that did not cluster are designated “unknown”). Genic structures are drawn to scale. N-terminal domains are colored green, and C-terminal domains are colored red. The illustration of the arrays of repeated motifs found within the internal coding regions is simplified, symbolized by alternating light and dark gray bars. Non-repetitive coding 'spacer' sequences (pink), scaffold gaps (black bars), 'linked' scaffolds validated by long-range PCR (string of “NNNNN”), and flanking noncoding or intronic sequences (thin lines and arrows) are also shown.
To assign correspondence between our N. clavipes spidroins and those previously described, we performed alignments of conserved N- and C-terminal sequences and internal motifs reported to be specific to a particular spidroin class43,44,45. Most of our N. clavipes spidroins clustered with one of the seven documented classes (Fig. 1 and Supplementary Figs. 2 and 3). We found novel members, expanding the minor ampullate, flagelliform, and aggregate classes of N. clavipes in comparison to other spider species (Supplementary Table 10). Surprisingly, seven N. clavipes spidroins (for example, Sp-907, Sp-1339, Sp-5803, Sp-8175, Sp-14910-A, Sp-14910-B, and Sp-74867) eluded assignment to the known classes on the basis of these alignments, suggesting the existence of additional spidroin classes or that class boundaries are less defined by sequence than previously assumed (Fig. 1 and Supplementary Figs. 2 and 3).
The coding lengths of the 20 full-length N. clavipes spidroins varied greatly, from 407 (MaSp-b) to 5,939 (AgSp-d) encoded amino acids (Fig. 1). A previous study reported two MiSp transcripts (∼7.5 kb and ∼9.5 kb) larger than the MiSp genes cataloged here46. We saw larger bands in our long-range PCR amplification of MiSp-c and MiSp-d (Supplementary Fig. 4), so the previously reported transcripts could represent length polymorphisms or additional unassembled MiSp genes. Our assemblies showed multiexon splicing at the two flagelliform-type loci (FLAG-a and FLAG-b), consistent with previous reports27 (Fig. 1). N. clavipes spidroins were outliers in their amino acid frequencies relative to all other predicted genes in the N. clavipes genome, being notably enriched for glycine (20.1%, Wilcoxon rank-sum test, P = 2.8 × 10−15), alanine (14.6%, P = 2.6 × 10−8), and serine (11.6%, P = 8.5 × 10−4) residues found in known repeated motifs: (GA)n, (A)n (polyalanine), and GGX (where X = A, S, or Y)44 (Supplementary Figs. 5 and 6).
To catalog repetitive elements found within N. clavipes spidroins, we performed computational motif discovery and labeling43, followed by searches for larger repetitive structures (Supplementary Note). We observed 394 unique motif variants, ranging from 4 to 34 amino acids in length (Fig. 2a). In addition to previously reported motifs, hundreds of the N. clavipes motif variants were completely novel and others were new variants of previously documented motifs (Fig. 2a and Supplementary Table 13). Arrays of repeated motifs spanned 50–96% (median 81%) of the internal coding lengths of the 20 complete spidroins (Fig. 2b). The overall diversity and complexity of these repetitive structures were greater than expected, considering previous reports29.
(a) Summary of the 394 distinct repetitive motifs found in N. clavipes spidroins, grouped by amino acid sequence. The most frequently observed motifs (≥30 occurrences) are listed here; “X” indicates a variable amino acid position. Our motif catalog includes known motifs (dark gray), new variants of known groups (light blue), and novel motifs not previously described in the literature (gold). Motifs that are repeated less frequently (<30 occurrences) or cannot be informatively grouped are designated “additional” and “unassigned” (purple), respectively. The asterisks indicate a diverse group that cannot be informatively exemplified. The complete list of N. clavipes motifs is provided in Supplementary Table 13. Circular symbols indicate the spidroin classes in which each motif group is observed. (b) Bar graph showing the extent of repetitive motif coverage in the structure of each spidroin. Beside each bar, we provide the percentage of the internal coding region composed of repetitive motifs calculated for fully assembled spidroins (NC, not calculated).
To better understand their diversity, we organized the unique motif variants into 49 motif groups on the basis of homology comparisons. One motif group consisted of GXGGX-containing motif variants, including the well-known motif variant GPGGY18,29 (Fig. 2a). Polyalanine motif variants29, four to ten residues in length, were grouped with novel polyalanine-like motif variants that also contained other residues (Supplementary Table 13). Meanwhile, one of the most frequently occurring motif groups was the novel DTXSYXTGEY group. Confined to two aggregate and one unclassified spidroin, three variants of this motif cumulatively occurred 554 times. Other frequently occurring novel motif groups included (i) GPGTTPGTI, (ii) multi TTX, (iii) multi GL, (iv) multi SQ/XQQ, and (v) non-alanine homopolymer runs. MaSp-g contained 73 different unique motif variants from 20 motif groups, the largest number observed in N. clavipes (Fig. 2b). AgSp-d contained the longest array (n = 546) of repeated motif occurrences (Supplementary Fig. 7). MaSp-f, MaSp-g, and MiSp-c displayed the greatest diversity, with motifs representing 20 of the 49 motif groups (Fig. 2b and Supplementary Table 13).
In the N. clavipes catalog of spidroin sequences, 46 of the 49 motif groups (260 of the 394 motif variants; 66%) were found in multiple spidroin genes (Figs. 2b and 3a). Strikingly, 204 of the 260 (78%) shared motifs were found in multiple silk classes. Having motifs in common appears to be a prevalent feature of spidroins (Fig. 3a–e), with MaSp-g containing the largest number of shared motifs (n = 63; Fig. 3a). We noted enrichment of shared novel motifs among the aggregate and unclassified spidroin classes (Fig. 3d). FLAG-a and the new flagelliform, FLAG-b, shared repeated motifs with all spidroins, but, curiously, both displayed less sharing with the aggregate spidroin AgSp-d (Fig. 3f,g). The DTXSYXTGEY motif was found predominately in AgSp-d, suggesting that this motif may confer some of the distinctive properties of this putatively sticky, non-fibrous spidroin.
(a) Bar graph comparing the number of shared (gold) versus private (dark gray) distinct repetitive motifs observed in each spidroin. (b–g) Circos plots illustrate sharing of motif sequences among N. clavipes spidroins, specifically showing sharing of sequences belonging to known motif groups (b), known motif groups with novel N. clavipes variants (c), novel motif groups (d), and unassigned motifs (e). In these plots, genes are arrayed around the circle, and links are drawn to connect similar motif sequences that occur in both genes. (f) Circos plot showing the extensive sharing of motifs between novel FLAG-b (VeSp) and the other N. clavipes spidroins, supporting classification of FLAG-b as a spidroin. (g) The Circos plot of FLAG-a shows a similar distribution of motif sharing as seen for FLAG-b and further highlights the lack of motif sharing of either gene with AgSp-d.
We also observed second-order repetitive organization of repeated motifs43,47. We defined a cassette as the tandem occurrence of unique motif variants repeated two to four times across spidroin sequences, and these cassettes were often organized into larger ensembles32,48. Cassettes were present in all N. clavipes spidroins (Fig. 4a, Supplementary Fig. 8, and Supplementary Table 14). We identified 506 different cassette types and 1,440 occurrences (Fig. 4a), spanning 25–95% of the motif arrays in the 20 full-length spidroins (Fig. 4b and Supplementary Fig. 8). Our catalog of cassettes included documented combinations of motifs such as XGGXGGX + polyalanine, GPG + polyalanine, and GPG + GXGGX9,49. Half of the most frequently occurring cassettes were tandem repetitions of motif variants from the same motif group (for example, tandem SQ: [SQSQQASV]2), while the remaining cassettes were arrays of different motifs (for example, GXGGX + polyalanine: GPGGY + [A]7).
(a) Summary of 506 distinct higher-order repetitive structures (cassettes) found in N. clavipes spidroins. Cassettes are grouped according to the motif types from which they are composed. For cassettes composed of more than one distinct motif, the amino acids belonging to each motif are specified by color (red or black). The asterisk indicates cassettes that are too large to display here or that cannot be informatively exemplified. A list of all N. clavipes cassettes is provided in Supplementary Table 14. Circular symbols indicate the spidroin classes in which each cassette group is observed. (b) Bar graph comparing the number of shared (gold) versus private (dark gray) distinct repetitive cassette types observed in each spidroin. Beside each bar, we provide the percentage of repetitive motifs composed of cassettes calculated for fully assembled spidroins (NC, not calculated).
We examined the extent of cassette sharing across spidroins. Of the 506 distinct cassettes, 480 (95%) were private to individual genes, in striking contrast to the extensive sharing of motifs (Fig. 4a,b). Cassette sharing existed mainly in the major and minor ampullate classes, but these genes still contained substantial numbers of private cassettes. Ten spidroins (MaSp-a, MaSp-d, MaSp-f, TuSp, AcSp, Sp-14910-B, FLAG-b, Sp-8175, AgSp-a, and AgSp-b) contained only private cassettes (Supplementary Fig. 8). These observations support the idea that shared motifs assembled into distinct private cassettes may differentiate spidroin gene functionality, conferring the different physical properties of silks30,36.
Expression profiling of individual spidroins across multiple tissues
Previous experimental findings have suggested that spidroin expression is not exclusively gland specific50,51,52,53,54. Our RNA sequencing studies also suggested broader patterns of spidroin transcript expression across silk glands (Supplementary Fig. 9 and Supplementary Note). To better understand the regulation of spidroin genes and the mechanisms of silk gland specialization, we directly interrogated the degree of spidroin expression bias by using qPCR to measure the RNA transcript levels of the 28 N. clavipes spidroins in morphologically classified silk glands and control tissues collected from three adult females (Supplementary Fig. 1b and Supplementary Note). We prepared three cleanly separated isolates of all morphologically distinct gland types except for the aciniform and piriform glands, which because of their proximal anatomical locations could not be cleanly separated and were therefore treated as a combined sample (“other silk glands”; Supplementary Note). In every silk gland assayed in our experiments, spidroin transcripts belonging to more than one silk class were detected (Supplementary Figs. 10 and 11). As expected from the bulk of previous studies, we found examples of spidroin genes from each class that were highly expressed in their corresponding morphologically distinct silk gland (for example, MiSp-c in minor ampullate gland; Fig. 5a). Our data also identified several cases in which spidroin transcripts were expressed abundantly in glands that did not correspond with the gene name (for example, MaSp-h in tubuliform gland; Fig. 5a). Some spidroins appeared to be expressed in all of the silk glands assayed (for example, AgSp-d; Fig. 5a). However, it is important to note that spidroin gene names have historically been conferred on the basis of the gland from which the gene was first cloned and do not assume exclusivity in expression. While we detected transcripts for the tubuliform spidroin TuSp in several silk glands, the expression of this gene was not highest in the tubuliform gland (Supplementary Figs. 10 and 11), possibly because the adult females from which the tissue samples were collected were not in the process of preparing to spin egg casings or might have recently done so32,48. Several genes showed strong expression in a particular gland type, providing clues regarding their function (for example, Sp-5803 in flagelliform gland; Fig. 5a). When viewed as a profile across silk glands, we found that many of our unknown spidroins showed patterns of expression similar to those of members of known silk classes, providing hypotheses for their functionality (for example, the profile of Sp-8175 expression strongly correlated with the expression profile of AgSp spidroins, and the profile for Sp-74867 correlated with that of MaSp spidroins; Fig. 5b).
(a) Box-and-whiskers plots showing the relative expression of four N. clavipes spidroin loci in individual tissue dissections (n = 3 biological replicates per tissue) assayed by qPCR. Tissues including legs, venom glands, five anatomically distinct silk glands, and 'other' silk glands (aciniform and piriform glands attached to spinneret) are shown on the x axis, and expression (2−ΔΔCT method) is depicted on the y axis (log10 scale). Box-and-whiskers plots show the range of expression values of the given spidroin gene (left y axis) relative to RPL13a (housekeeping gene) expression and normalized to leg tissue. Thick black center lines represent median values. Upper whiskers represent largest observation ≤ upper quartile (Q3) + 1.5 interquartile range (IQR), and lower whiskers represent smallest observation ≥ lower quartile (Q1) −1.5(IQR). Red asterisks mark silk glands with significantly greater expression of a given gene over leg tissue, whereas black asterisks indicate a single silk gland type exhibiting significantly greater expression values of a given gene than all other silk gland types together (one-tailed Wilcoxon rank-sum tests): *P < 0.05, **P < 0.01. (b) Heat map showing patterns of co-expression among N. clavipes spidroins as assayed by qPCR across venom and silk glands. Co-expression scores were calculated using Pearson correlation of relative expression values (2−ΔΔCT) for each pair of genes and plotted using single-linkage hierarchical clustering. Owing to sequence similarity between MaSp-b and MaSp-c the data for these two transcripts are presented together as “MaSp-b,c”. (c) Box-and-whiskers plot showing the expression of three genes (FLAG-a, FLAG-b (VeSp), and PR-1), assayed using qPCR in flagelliform silk glands (left three boxes) and venom glands (right three boxes) collected from three mature N. clavipes females (n = 3 tissue samples for each type). Significantly greater expression of FLAG-a was detected in flagelliform silk glands, whereas significantly greater expression of PR-1 (a venom-specific toxin gene used as a control) and FLAG-b was detected in the venom glands. Box and whiskers show the range of relative expression values (calculated using the 2−ΔΔCT method) of each of the three genes for both tissue types relative to RPL13a (a housekeeping gene) expression in each tissue and normalized to leg tissue. Thick black center lines represent median expression values. Upper whiskers represent largest observation ≤ upper quartile (Q3) +1.5 interquartile range (IQR), and lower whiskers represent smallest observation ≥ lower quartile (Q1) − 1.5(IQR). Black asterisks indicate that a given gene exhibits significantly greater (FLAG-a) or lower (FLAG-b, PR-1) expression in flagelliform silk glands versus venom glands (n = 3 samples per gland, one-tailed Wilcoxon rank-sum test): *P = 0.05. (d) Evidence of alternative spliceoforms of MaSp-f. Junction reads mapping to two distinct isoforms were detected. Junction reads mapping to MaSp-f isoform 2 were observed in silk 1 and silk 2 gland isolates, as indicated by the heat map beneath the isoform cartoon.
It is generally assumed that spidroin expression is confined to silk glands, but this was not the case for novel FLAG-b (Fig. 5c). In fact, the highest abundance of FLAG-b transcripts was detected in venom glands, a finding consistent with results from our RNA sequencing studies (Supplementary Fig. 9). PR-1 (a known venom toxin55) and FLAG-a (the established flagelliform spidroin) transcripts were enriched in the expected tissues and served as controls (Fig. 5c). Normalized FLAG-b transcript levels were ∼1,000- to 5,000-fold higher in venom gland than they were in silk glands (Wilcoxon rank-sum test, P = 0.00075).
Extreme spidroin diversity and evolutionary origins
The multiexon structure of some spidroins has led to conjecture that alternative splicing may increase transcript diversity9,49, although evidence for spidroin spliceoforms has not previously been confirmed experimentally30,36. We detected split reads that are evidence of alternative splicing of MaSp-f transcripts into two spliceoforms: the major full-length isoform and a minor isoform lacking most of the second exon (Fig. 5d). Given that the second exon of the full-length isoform encodes the putative C-terminal domain, which has been shown in other MaSp spidroins to act as a switch between storage and assembly forms of silk proteins and as a facilitator of protein organization in silk formation56, this raises the possibility that the second, truncated MaSp-f isoform transitions or organizes in a different manner than the full-length isoform.
To identify non-spidroin candidate genes potentially involved in silk production, we cataloged transcripts that were (i) highly expressed and/or (ii) uniquely expressed in N. clavipes silk glands, resulting in a list of 649 candidates from our RNA sequencing data (Supplementary Fig. 12 and Supplementary Table 15); 183 of these genes exhibited homology to documented silk gland–specific transcripts50,51,52,53,54. The candidates included catalytic enzymes such as kinases, proteases, dehydrogenases, acetyltransferases, and synthases, many of which are active in eukaryotic secretory systems57. We expect this catalog to include genes encoding proteins involved in the conversion of liquid silk dope to solid silk thread52,53, such as enzymes that maintain the pH gradient along the gland body to spigots on the spinnerets50,54. Candidates that might generate ions for the pH gradient include three carbonic anhydrase orthologs (Ca10, Ca13, and Ca14), four thyroid peroxidase (Tpo) paralogs, and five chorion peroxidase (Pxt) paralogs (Supplementary Table 15).
We found evidence for at least two different evolutionary mechanisms that might contribute to the diversification of spidroin loci. First, we saw evidence of new spidroin genes originating from tandem-duplication events, as hypothesized in earlier studies17,26,29,58,59,60. We identified pairs of tandem spidroins on two genomic scaffolds (scaffold_16392, MaSp-d and MaSp-e; scaffold_14910, Sp-14910-A and Sp-14910-B). We note here that the profiles of expression across silk glands between the pair-mates of the two pairs of tandem spidroins were strongly correlated (Fig. 5b). Second, we noted a ∼2-fold higher level of polymorphism in sequences encoding the N- and C-terminal domains of spidroins (mean θW = 0.002) in comparison to levels observed across the coding genome (mean θW = 0.0009; Wilcoxon rank-sum test, P = 5.4 × 10−6) (Supplementary Fig. 13a–c). This is a signature consistent with long-term balancing selection occurring at spidroin loci.
Discussion
To translate the remarkable properties of spider silks into innovative medical and industrial applications, it is necessary to expand knowledge of the diversity of the underlying gene structures, the relationships between structure and function, and final product synthesis. By generating the first annotated genome of an orb-weaving spider, we have taken an important step toward these ends. Our efforts—made possible by previous studies that documented spidroin sequences from several spider species18,27,43—have enabled us to catalog an extensive collection of 28 spidroins representing the full spectrum of araneoid silk classes, identifying 8 previously unreported spidroins and providing a wealth of new repetitive elements. Complemented by silk gland–specific expression profiling, this collection of data greatly expands understanding of spidroin gene diversity, structure, and expression across morphologically distinct silk glands.
Every silk gland that we profiled was found to express broad combinations of spidroin transcripts representing multiple classes. Together with analogous non-gland-specific expression patterns seen for individual spidroins in other spider species27,53, these data argue for complex, gland-specific models of spidroin expression and silk production. Future studies that measure the levels of spidroin proteins in silk glands would be useful to validate the correspondence of transcript abundance to translational output and will provide an additional dimension to understanding of silk production.
The generation of full-length sequences and assemblies has shed light on the evolutionary mechanisms acting on the N. clavipes spidroin gene family. Evidence of tandem gene duplication and high levels of polymorphism suggests that spidroins are naturally selected to maintain diversity. The presence of long arrays of tandem repeats in spidroins, reminiscent of those that facilitate rapid variation on microbial cell surfaces61 and morphological plasticity in mammals62, also suggests a gene family undergoing continuous evolution. In particular, N. clavipes spidroin gene phylogenies (Fig. 1 and Supplementary Figs. 2 and 3) provide evidence for the shared origin and nearly simultaneous expansion of the entire flagelliform and aggregate spidroin classes, supporting expectations that they are tightly linked to one another functionally and to the origin of the viscid orb web32.
Highlighting the diversity that has evolved within the spidroin family, perhaps the most remarkable example reported here is the novel flagelliform gene FLAG-b. This gene's sequence similarity and phylogenetic affinity to canonical flagelliform FLAG-a18 (Supplementary Figs. 2 and 3) suggest that it arose as a second genomic copy from a duplication event, yet FLAG-b transcripts are highly abundant in venom glands. This discovery is reminiscent of a puzzling detection of peptides from two spidroin-like proteins (S.m. Sp1 and S.m. Sp2b) in the venom gland of the velvet spider27. Our expression data add clarity to this issue, suggesting that FLAG-b has evolved functions beyond silk-related applications in N. clavipes, and refute the idea that spidroin expression is restricted to silk glands. As such, this flagelliform may represent a new kind of venom gland–expressed spidroin (VeSp). This observation suggests promising avenues for future research on links between spider silk and venom, both composed of complicated proteins whose production is a functional synapomorphy for the order Araneae. In this context, the spitting spider (Scytodes sp.) is particularly interesting, as it is the only spider to exude from its chelicerae fibrous 'venom' that is used to immobilize prey by gluing it to a substrate63,64,65. However, a recent transcriptomic and proteomic investigation did not find evidence of spidroins in the venom of Scytodes thoracica66, suggesting a different evolutionary route for this functional convergence. Proteomic studies could demonstrate whether FLAG-b is indeed translated into a protein in N. clavipes venom glands. If so, this raises the intriguing possibility that FLAG-b functions as a buffer, chaperone, adhesive, or preservative for the smaller bioactive compounds found in N. clavipes venom and thus may provide a novel use of spidroins in human medical applications.
Systematic characterization of the diversity, number, and structure of the repetitive elements found in N. clavipes spidroins yields the key observations that 66% of repeated motif variants are shared both within and across spidroin classes, whereas 95% of cassette variants are private to individual spidroins. Together, these observations suggest that the assembly of shared motifs into distinct cassettes, often organized into larger repeat ensembles29, may underlie the range of unique biophysical characteristics observed for the various spider silk classes—an idea also supported by recent results from transgenic spidroin expression in silkworm33,67. Given the relationships already observed between spidroin motif sequences and structural properties of silk proteins34, the extensive catalog of novel motifs and combinations reported here provides many candidates for transgenic studies. Our catalog can provide deeper understanding of the interplay between silk genes, silk protein structure, and the biomechanical properties of silks and will underlie future efforts to capture the extraordinary properties of spider silks in manmade materials.
Methods
For expanded methodological details, please refer to the separate Supplementary Note, Supplementary Figures 1–13, and Supplementary Tables 1–15. qPCR data and statistical test results are provided in the Supplementary Data.
Spider specimens.
Genomic DNA was extracted from three wild-caught N. clavipes adult females collected from Charleston County, South Carolina, USA. RNA for RNA sequencing experiments was obtained from four wild-caught N. clavipes adult females collected from Charleston County, South Carolina, USA. RNA for qPCR validation experiments was extracted from three wild-caught N. clavipes adult females from Citrus County, Florida, USA (Supplementary Table 1).
See the Supplementary Note for additional details.
DNA extraction and sequencing.
DNA was extracted using phenol:chloroform and column-based methods. Short-fragment (180-bp) paired-end sequencing libraries were constructed using TruSeq LT kits (Illumina). Paired-end long-insert jumping libraries were built using two protocols: the Illumina Mate Pair v2 (MPv2) and Nextera Mate Pair kits. MPv2 libraries featured inserts with mean sizes of of 3 kb, 5 kb, 7 kb, 9 kb, and 11 kb, whereas Nextera Mate Pair libraries featured inserts with mean sizes of 2 kb, 4 kb, 5 kb, 6 kb, 7 kb, 9 kb, 11 kb, 13 kb, and 17 kb (Supplementary Table 2).
Whole-genome shotgun sequencing was performed on the Illumina MiSeq (150 × 150 paired-end read lengths), Illumina HiSeq 2000 (100 × 100), and Illumina HiSeq 2500 (100 × 100) platforms using TruSeq v3 cluster kits and TruSeq sequencing-by-synthesis (SBS) chemistry.
See the Supplementary Note for additional details.
RNA extraction and sequencing.
For two individuals, RNA was extracted from the entirety of each specimen for two 'whole-body' RNA sequencing libraries. For the other two individuals, select tissues were microdissected (silk glands, venom glands, and brain tissue) and used for 14 tissue-specific RNA sequencing libraries (Supplementary Tables 1 and 2). In all cases, RNA was extracted using a combined TRIzol (Ambion, Life Technologies) and column-based protocol. Each of the 16 individual RNA samples was treated with TURBO-free DNase (Life Technologies), and rRNA content was depleted with the Ribo-Zero Gold kit (Epicentre, Human/Mouse/Rat). Strand-specific RNA sequencing libraries were constructed using the NEBnext Ultra-Directional RNA Library Prep kit (NEB, protocol B) and barcoded using TruSeq RNA adaptors (Illumina). All of the N. clavipes RNA sequencing libraries are listed in Supplementary Table 2.
High-throughput RNA sequencing was performed on the Illumina HiSeq 2000 (100 × 100) platform using TruSeq v3 cluster kits and TruSeq SBS chemistry (Illumina).
See the Supplementary Note for additional details.
De novo genome assembly.
Raw FASTQ read files were evaluated using FastQC (v0.11.2) and then trimmed using Trimmomatic (v0.32)68 to remove adaptor read-through, low-quality bases, and ambiguous base calls. All jumping mate-pair DNA libraries were processed using the program FastUniq (v1.1)69 to remove duplicate read pairs.
The N. clavipes genome was assembled de novo using a meta-assembly approach. Two draft assemblies were constructed in parallel using AllPaths-LG vR49967 (ref. 70) and SOAPdenovo2 (v2.04)71 and were then merged using Metassembler (v1.1)72. Genomic quality metrics were calculated for all N. clavipes assemblies using scripts from the Assemblathon 2 competition73, available at https://github.com/ucdavis-bioinformatics/assemblathon2-analysis/. To assess the genome's functional 'completeness', the Benchmarking Universal Single-Copy Orthologs (BUSCO) gene mapping method74 was also applied to all N. clavipes assemblies to identify conserved protein-coding genetic loci. All single-copy gene sequences from Ixodes scapularis (deer tick) were extracted from the BUSCO Arthropod gene set, a 95% refinement cutoff was applied, and 2,058 I. scapularis loci were used to query the completeness of all intermediate and final N. clavipes assemblies (Supplementary Table 5), as well as the all-isolate transcriptome assembly described below (Supplementary Table 7).
See the Supplementary Note for additional details.
De novo transcriptome assembly.
After quality control and filtering of reads, all RNA libraries were de novo assembled together as a primary all-isolate transcriptome using Trinity (rel_2.25.13)75,76 (Supplementary Table 5). Meanwhile, 16 tissue-specific transcriptomes were individually de novo assembled using Trinity (Supplementary Tables 1, 5, and 6). All transcripts were aligned back to the genome using the splice-aware mRNA/EST aligner GMAP (rel_10.22.14)77, and reads from each RNA library were aligned to the genome using RNA-STAR (2.4.2a)78 (Supplementary Table 6).
See the Supplementary Note for additional details.
Genome annotation.
Genomic features were defined on the final N. clavipes meta-assembly using four successive rounds of the annotation pipeline Maker2 (ref. 38). Repetitive regions were identified using RepeatRunner (supplied with Maker2), RepeatMasker v4.0.5 with RMblast, and RepBase repeat libraries79 and subsequently masked for downstream gene modeling. Tandem repeats were identified using Tandem Repeats Finder (v4.07b)80.
Gene models were based on multiple types of evidence: alternate species protein sequence alignments, alternate species EST/mRNA/cDNA sequence alignments, de novo–assembled transcripts from N. clavipes RNA–seq experiments, and ab initio gene predictions. Protein and EST/mRNA sequences were collected from online databases (Supplementary Table 8). Exon boundaries were marked using Exonerate (v2.2.0)81, and tRNAs were identified by tRNAscan-SE82. Feature boundaries were further polished by Maker2, directing successive rounds of trained predictions from SNAP (rel_11.29.13)39 and Augustus (v3.0.2)40 (WebAugustus41). In total, >32 million genomic features and 403,888 putative genes were modeled on the final N. clavipes annotated meta-assembly (Supplementary Table 9). Putative gene model identities were established by the reciprocal alignment of model protein sequences to the UniProtKB/SwissProt83 protein database (v6.3.15) using BLASTP84 and Maker2 accessory scripts.
Five tiers/sets of gene models with increasing stringency were defined on the basis of agreement among coding feature annotations, conserved protein domains, eukaryotic gene structure, and similarities with curated gene databases (Supplementary Table 9). The 'standard' gene set (54,186 genes, 58,132 mRNAs) contained only gene models that possessed known protein domains from the InterPro Pfam database85 and was produced using BLASTP84, HMMer (http://www.hmmer.org/), and Maker Standard scripts (K. Childs (Michigan State University), personal communication). The conservative 'gold' gene set (14,025 genes, 17,989 mRNAs) contained only the subset of gene models that were built from biological evidence (RNA, protein alignment). This gold gene set was used for all downstream analyses.
See the Supplementary Note for additional details.
Phylogenetic analyses.
Interspecific spidroin phylogenetic reconstructions were performed by aligning the first ∼130 N-terminal-domain residues of N. clavipes spidroins with those of all available spidroin sequences (Supplementary Table 10) using Geneious, Clustal, and BLOSUM62 (ref. 86). Trees were built with PhyML and RAxML and rooted with a Bothriocyrtum californicum fibroin sequence31, and bootstrap values were based on 1,000 replicates (Supplementary Fig. 2). The intraspecific N. clavipes spidroin phylogeny was similarly constructed using Geneious, Clustal, and BLOSUM62 (ref. 86), and unrooted spidroin trees were built with PhyML and BLOSUM62 with bootstrap values based on 1,000 replicates (Supplementary Fig. 3).
See the Supplementary Note for additional details.
Spidroin identification.
N. clavipes spidroins were identified by multiple BLAST84 searches of the genome, transcriptomes, and gene models using the spidroin query sequences detailed in Supplementary Tables 8, 10, and 12. Five loci exhibited complete coding sequences, but the majority of putative N. clavipes spidroins had internal sequence gaps, were only repeats, or encoded incomplete N- or C-terminal sequences at the ends of scaffolds. To find missing pieces, additional rounds of searching were performed by adding N. clavipes spidroin hits from the previous round to each new list of queries, ultimately yielding 349 genome hits, 364 transcriptome hits, 292 gene model protein hits, and 292 gene model transcript hits. Putative spidroin fragments were organized into five categories for validation and completion experiments: complete, internal gap, 5′ end, 3′ end, and repetitive sequence (Supplementary Table 12).
See the Supplementary Note for additional details.
Spidroin sequence validation using long-range PCR.
Putative N. clavipes spidroins were isolated and filled using a combination of long-range PCR (LR-PCR) and single-molecule real-time (SMRT) sequencing of a single N. clavipes adult female (Nep-010; Supplementary Table 1) at very high coverage. Multiple pairs of LR-PCR primers (Supplementary Table 11) were designed for each scaffold (Primer3), so that putative spidroin loci could be completely isolated by LR-PCR amplicons, and alternate primer pairs could be recruited in cases of suboptimal amplification. Pair mates were proposed using sequence similarity, orthologous alignments, and transcript tissue specificity. To 'bridge' two separate scaffolds, multiple combinations of cross-pair LR-PCR experiments were performed to identify scaffold pairs that were more cryptically related. LR-PCR reactions employed high-efficiency PrimeStar GXL polymerase (Clontech/TaKaRa), and amplicons were visualized on low-voltage 0.5% Bio-Rad Certified Megabase agarose gels. Amplicons were purified and pooled at equimolar ratios, with slightly higher volumes for the longest fragments (>20 kb). Two unique pools of spidroin amplicons were processed for SMRT library construction, as outlined in ref. 87, and sequenced using the P6-C4 sequencing enzyme, chemistry, and 4-h movie collection parameters (Pacific Biosciences). Quality-filtered FASTQ files of long SMRT reads were directly aligned to scaffolds that exhibited complete spidroins (five) or spidroins with internal gaps on single scaffolds (ten) using PBJelly (PBSuite 15.2.20.p1)88 and BLASR (v1.3.1)89. For putatively linked scaffold pairs (13 pairs), manual alignments were performed to effectively bridge gaps, correct errors, and resolve repeats. In total, 28 N. clavipes spidroin sequences (20 complete) were validated (Fig. 2 and Supplementary Table 12).
See the Supplementary Note for additional details.
Spidroin gene repeat motif identification and analyses.
All N. clavipes spidroins were translated into amino acid residues and then subjected to repeat motif identification using MEME and motif painting with MAST (v4.10)46. Repetitive motifs were manually curated to remove low-quality hits and motifs occurring in N- and C-terminal domains (using a hard cutoff of 100 residues) and were cataloged as unique 'motif variants' ranging from 4 to 87 residues in length. Motif variants were then organized into 'motif groups' on the basis of residue content and sequence, following the rules listed in Supplementary Table 13. Motifs that could not be informatively grouped were designated as 'unassigned'. The full catalog of motif variants was input in secondary rounds of motif searching with the custom pipeline Spider_pipeline.py, available at https://github.com/danich1/Spider-Pipeline (Figs. 2a,b and 3a–d, and Supplementary Fig. 7). Next, the pipeline was used to search for higher-order repetitive structures denoted as 'cassette variants', defined as two to four adjacent motif occurrences that were enriched across spidroins. Cassette variants were organized into 'cassette groups' (Supplementary Table 14) and curated to remove cassette types that exhibited inter-motif gaps of >20 residues or that occurred in N- and C-terminal domains (Fig. 4a,b and Supplementary Table 8).
See the Supplementary Note for additional details.
Amino acid content analyses.
To provide a background level of residue content for the N. clavipes coding genome, the proportion of each of the 20 different amino acids was computed for each of the 17,989 translated mRNA sequences from the gold gene set. The same was done for the 28 N. clavipes spidroin sequences (Supplementary Fig. 6). To test for significant enrichment of amino acid types, the distribution of each residue's proportions of non-spidroins was compared to those of spidroins using two-tailed unequal-variance Wilcoxon rank-sum tests.
See the Supplementary Note for additional details.
Polymorphism levels across the N. clavipes genome.
To quantify polymorphism in the N. clavipes genome, all fragment sequencing reads from a single individual (Nep-004) were remapped to the genome using BWA-MEM90 and variant calling of SNPs and small indels was performed using SAMtools91,92. Variants were hard-filtered to include only SNPs meeting minimum quality (20) and depth (20) thresholds and then subdivided into 14 categories (genome, noncoding, genes (gold set), CDS, mRNAs, 3′ UTRs, 5′ UTRs, exons, introns, gold N termini, gold C termini, spidroin genes, spidroin N termini, spidroin C termini) using VCFtools93. SNPs were counted for each category, and polymorphic levels were assessed on the basis of heterozygosity, number of segregating sites, SNP rate, and Watterson's estimator of theta (θW)94. The distribution of θW values of non-spidroins was compared to that of spidroins using the Wilcoxon rank-sum test.
See the Supplementary Note for additional details.
Expression and alternative splicing analyses.
To compare the relative abundance of N. clavipes gene models across the 16 different tissue isolates, reads from each RNA library were aligned to the final N. clavipes annotated meta-assembly using STAR (v2.4.2a). Next, the PORT v0.7.3 expression pipeline (https://github.com/itmat/Normalization) was applied to normalize and quantify the RNA–seq data between the libraries.
To identify putative alternatively spliced transcripts that existed among the gold genes and spidroins, the PORT pipeline was run in exon/intron mode to quantify reads mapping to genomic features at the splice-junction level. Normalized counts of the split-RNA reads mapping across each junction were summarized at each locus of putative alternative splicing. Proportions of split reads mapping to different alternative junctions were then calculated for each tissue type (Fig. 5d).
See the Supplementary Note for additional details.
qPCR analysis.
To test the relative expression of spidroin loci in discrete anatomical subsections, including specific silk gland types, qPCR analysis was performed with RNA transcripts isolated from three additional mature female N. clavipes individuals from Citrus County, Florida, USA. From the abdomen, silk glands were identified by relative position and morphology and then individually collected by severing their ducts near the spinnerets. From the cephalothorax, legs were collected, venom glands were collected after separation of the chelicerae from the cephalothorax, and the remaining cephalothorax tissue was retained as the 'head' sample. In total, each specimen was microdissected into 9 tissue subsections—venom glands, head (with no venom glands), legs, major ampullate silk gland (MA), minor ampullate silk gland (MI), flagelliform silk gland (FL), aggregate silk gland (AG), tubuliform silk glands, and 'other silk glands' (OTHER: piriform and aciniform glands, attached to the spinneret), yielding 27 experimental samples in total (Supplementary Table 1). RNA was extracted using TRIzol (Ambion, Life Technologies) and RNeasy Mini kit spin columns (Qiagen), and additional cleanup was performed using the RNA Clean & Concentrator-5 kit with DNase I treatment (Zymo Research). Small aliquots (∼5 μl) were used for quality control and quantification. cDNA was produced from each RNA sample with a High-Capacity cDNA Reverse Transcription kit (Life Technologies) and run alongside multiple and 'no reverse transcriptase' (NRT) negative controls. Primers were designed to target 30 loci (all 28 spidroins, 1 venom locus (CRiSP/Allergen/PR-1)55, and 1 housekeeping gene (RPL13a)95), as well as 22 genomic scaffold controls for all single-exon spidroin genes (Supplementary Table 11). qPCR reactions were set up in triplicate using standard SYBR Green PCR Master Mix (Life Technologies) and run on a ViiA 7 Real-Time PCR machine. Relative transcript abundance of targets in silk and venom gland samples was normalized to that of leg tissue samples and calculated using the  method96 (Fig. 5a–c and Supplementary Figs. 10 and 11). Co-expression scores were calculated using Pearson correlation of relative expression values (
method96 (Fig. 5a–c and Supplementary Figs. 10 and 11). Co-expression scores were calculated using Pearson correlation of relative expression values ( ) for each pair of genes and plotted using single-linkage hierarchical clustering (Fig. 5b).
) for each pair of genes and plotted using single-linkage hierarchical clustering (Fig. 5b).
See the Supplementary Note for additional details and the Supplementary Data for the complete listing of relative expression values ( ) for all replicates.
) for all replicates.
Identification of non-spidroin silk gland–specific transcripts.
The normalized expression data set of 14,025 'gold' genes was filtered to identify putative SST loci that could be categorized as (i) 'HighInSilk', with >1,000 absolute normalized mapped RNA reads in ≥1 silk gland and <200 reads in non-silk tissues; (ii) 'ExclusiveToSilk', with >100 absolute normalized mapped RNA reads in ≥1 silk gland and zero reads in non-silk tissues; (iii) 'GlandEnriched', with >400 absolute normalized mapped reads in only a single silk gland and <350 in all non-silk tissues; and (iv) 'Literature', corresponding to the BLASTP84 homologs (e value: ≤ 1 × 10−6) of 282 unique SSTs from studies of spider silk gland transcriptomes and proteomes50,51,52,53,54 plus peroxidase or anhydrase gene family members hypothesized to be involved in silk production50,54 (Supplementary Fig. 12 and Supplementary Table 15).
See the Supplementary Note for additional details.
Statistical methods.
To test for significant enrichment of amino acid types, the distribution of each residue's proportions for 17,989 translated non-spidroin mRNA sequences was compared to those for 28 spidroins using two-tailed unequal-variance Wilcoxon rank-sum tests. Of the 20 residues, glycine (W = 469,088.5, P = 2.8 × 10−15), alanine (W = 404,937.5, P = 2.595 × 10−8), and serine (W = 343,584, P = 8.505 × 10−4) occurred in significantly higher proportions among the 28 spidroins in comparison to background (Supplementary Figs. 5 and 6).
To compare SNP polymorphism levels in non-spidroin and spidroin loci, the value for four haploid individuals was applied when calculating θW for non-spidroin loci and the value for six haploid individuals was used for calculating θW in spidroin loci. The distribution of θW values for non-spidroins was compared to that of spidroins using the two-tailed Wilcoxon rank-sum test (W = 99,362, P = 5.402 × 10−6; Supplementary Fig. 13a–c).
To assess the relative expression levels of spidroin loci in different tissues, we calculated  values from qPCR experiments as described by Livak and Schmittgen96. Each gene × tissue reaction was run in triplicate (three independent experiments) to control for technical variation. Cycling threshold (CT) values were averaged across technical replicates for each gene × tissue combination for each sample. The average CT values were then normalized to average RPL13a (housekeeping gene) CT values for the same tissue sample (ΔCT). ΔCT values for each gene × tissue combination were then normalized to the ΔCT values of the same gene for the leg tissue subsection of the same sample (ΔΔCT), which was then raised to the negative exponent of 2 (
values from qPCR experiments as described by Livak and Schmittgen96. Each gene × tissue reaction was run in triplicate (three independent experiments) to control for technical variation. Cycling threshold (CT) values were averaged across technical replicates for each gene × tissue combination for each sample. The average CT values were then normalized to average RPL13a (housekeeping gene) CT values for the same tissue sample (ΔCT). ΔCT values for each gene × tissue combination were then normalized to the ΔCT values of the same gene for the leg tissue subsection of the same sample (ΔΔCT), which was then raised to the negative exponent of 2 ( ). Biological replicates of each tissue (from three independent spiders) were kept separate for all calculations. The variances of relative expression values for each gene were compared across tissues using F tests, and their population means were tested using one-tailed unequal-variance Wilcoxon rank-sum tests (Fig. 5a–c and Supplementary Figs. 10 and 11). All F test and Wilcoxon rank-sum test input values and results are provided in the Supplementary Data. All statistical analyses were conducted with R v3.3.2 (R Foundation for Statistical Computing; https://www.r-project.org/foundation/). Circos plots were generated as described97.
). Biological replicates of each tissue (from three independent spiders) were kept separate for all calculations. The variances of relative expression values for each gene were compared across tissues using F tests, and their population means were tested using one-tailed unequal-variance Wilcoxon rank-sum tests (Fig. 5a–c and Supplementary Figs. 10 and 11). All F test and Wilcoxon rank-sum test input values and results are provided in the Supplementary Data. All statistical analyses were conducted with R v3.3.2 (R Foundation for Statistical Computing; https://www.r-project.org/foundation/). Circos plots were generated as described97.
Data availability.
All data are available as supplementary material or from the following databases as described. Data from this study are available through the central BioProject database at NCBI under project accession PRJNA356433 and are linked to BioSample accessions SAMN06132062 – SAMN06132080. The whole-genome sequence is available at the Whole-Genome Shotgun (WGS) database under accession MWRG00000000. All short-read sequencing data have been deposited in the NCBI Short Read Archive (study, SRP095945; experiments, SRX2458083–SRX2458130; runs, SRR5139318–SRR5139365), and transcriptome data are available at the Transcriptome Shotgun Assembly (TSA) under accession GFKT00000000.
URLs.
Repeat motif and cassette identification scripts, https://github.com/danich1/Spider-Pipeline; PORT v0.7.3 expression pipeline, https://github.com/itmat/Normalization; Assemblathon 2 assessment script, https://github.com/ucdavis-bioinformatics/assemblathon2-analysis/; Augustus, http://augustus.gobics.de/binaries/; BLASR, https://github.com/PacificBiosciences/blasr; BLAST, ftp://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/; BUSCO, http://busco.ezlab.org/; BWA-MEM, http://bio-bwa.sourceforge.net/; FastQC, http://www.bioinformatics.babraham.ac.uk/projects/fastqc/; FastUniq, https://sourceforge.net/projects/fastuniq/; Geneid, http://genome.crg.es/software/geneid/; Geneious Pro, http://www.geneious.com/; Hmmer, http://hmmer.org; Maker2, http://www.yandell-lab.org/software/maker.html; Metassembler, https://sourceforge.net/projects/metassembler/; PBSuite, https://sourceforge.net/projects/pb-jelly/; PicardTools, https://broadinstitute.github.io/picard/; R, https://www.r-project.org/foundation/; RepeatMasker, http://www.repeatmasker.org/; RNA-STAR, https://github.com/alexdobin/STAR/releases/; SAMtools, http://samtools.sourceforge.net/; SOAPdenovo2, https://github.com/aquaskyline/SOAPdenovo2/; Tandem Repeats Finder, https://tandem.bu.edu/trf/trf.html; Trimmomatic, http://www.usadellab.org/cms/?page=trimmomatic; Trinity, https://github.com/trinityrnaseq/trinityrnaseq/wiki; WebAugustus, http://bioinf.uni-greifswald.de/augustus/.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Accession codes
Primary accessions
BioProject
NCBI Reference Sequence
Sequence Read Archive
References
Natural History Museum Bern. The World Spider Catalog, version 18.0 http://wsc.nmbe.ch/ (accessed 9 November 2016).
Garrison, N.L. et al. Spider phylogenomics: untangling the Spider Tree of Life. PeerJ 4, e1719 (2016).
Blackledge, T.A. et al. Reconstructing web evolution and spider diversification in the molecular era. Proc. Natl. Acad. Sci. USA 106, 5229–5234 (2009).
Agnarsson, I., Kuntner, M. & Blackledge, T.A. Bioprospecting finds the toughest biological material: extraordinary silk from a giant riverine orb spider. PLoS One 5, e11234 (2010).
Swanson, B.O., Blackledge, T.A., Beltrán, J. & Hayashi, C.Y. Variation in the material properties of spider dragline silk across species. Appl. Phys., A Mater. Sci. Process. 82, 213–218 (2006).
Yang, Y. et al. Toughness of spider silk at high and low temperatures. Adv. Mater. 17, 84–88 (2005).
Steven, E. et al. Carbon nanotubes on a spider silk scaffold. Nat. Commun. 4, 2435 (2013).
Wright, S. & Goodacre, S.L. Evidence for antimicrobial activity associated with common house spider silk. BMC Res. Notes 5, 326 (2012).
Vollrath, F. & Knight, D.P. Liquid crystalline spinning of spider silk. Nature 410, 541–548 (2001).
Rising, A. & Johansson, J. Toward spinning artificial spider silk. Nat. Chem. Biol. 11, 309–315 (2015).
Gosline, J.M., DeMont, M.E. & Denny, M.W. The structure and properties of spider silk. Endeavour 10, 37–43 (1986).
Swanson, B.O., Blackledge, T.A., Summers, A.P. & Hayashi, C.Y. Spider dragline silk: correlated and mosaic evolution in high-performance biological materials. Evolution 60, 2539–2551 (2006).
Blasingame, E. et al. Pyriform spidroin 1, a novel member of the silk gene family that anchors dragline silk fibers in attachment discs of the black widow spider, Latrodectus hesperus. J. Biol. Chem. 284, 29097–29108 (2009).
Geurts, P. et al. Synthetic spider silk fibers spun from Pyriform Spidroin 2, a glue silk protein discovered in orb-weaving spider attachment discs. Biomacromolecules 11, 3495–3503 (2010).
Ayoub, N.A., Garb, J.E., Kuelbs, A. & Hayashi, C.Y. Ancient properties of spider silks revealed by the complete gene sequence of the prey-wrapping silk protein (AcSp1). Mol. Biol. Evol. 30, 589–601 (2013).
Hu, X. et al. Araneoid egg case silk: a fibroin with novel ensemble repeat units from the black widow spider, Latrodectus hesperus. Biochemistry 44, 10020–10027 (2005).
Garb, J.E. & Hayashi, C.Y. Modular evolution of egg case silk genes across orb-weaving spider superfamilies. Proc. Natl. Acad. Sci. USA 102, 11379–11384 (2005).
Hayashi, C.Y. & Lewis, R.V. Molecular architecture and evolution of a modular spider silk protein gene. Science 287, 1477–1479 (2000).
Adrianos, S.L. et al. Nephila clavipes Flagelliform silk-like GGX motifs contribute to extensibility and spacer motifs contribute to strength in synthetic spider silk fibers. Biomacromolecules 14, 1751–1760 (2013).
Higgins, L.E., Townley, M.A. & Tillinghast, E.K. Variation in the chemical composition of orb webs built by the spider Nephila clavipes (Araneae, Tetragnathidae). J. Arachnol. 29, 82–94 (2001).
Choresh, O., Bayarmagnai, B. & Lewis, R.V. Spider web glue: two proteins expressed from opposite strands of the same DNA sequence. Biomacromolecules 10, 2852–2856 (2009).
Vasanthavada, K. et al. Spider glue proteins have distinct architectures compared with traditional spidroin family members. J. Biol. Chem. 287, 35986–35999 (2012).
Townley, M.A. & Tillinghast, E.K. in Spider Ecophysiology (ed. Nentwif, W.) 283–302 (Springer 2013).
Townley, M.A., Pu, Q., Zercher, C.K., Neefus, C.D. & Tillinghast, E.K. Small organic solutes in sticky droplets from orb webs of the spider Zygiella atrica (Araneae; Araneidae): β-alaninamide is a novel and abundant component. Chem. Biodivers. 9, 2159–2174 (2012).
Blackledge, T.A. & Hayashi, C.Y. Unraveling the mechanical properties of composite silk threads spun by cribellate orb-weaving spiders. J. Exp. Biol. 209, 3131–3140 (2006).
Chaw, R.C. et al. Intragenic homogenization and multiple copies of prey-wrapping silk genes in Argiope garden spiders. BMC Evol. Biol. 14, 31 (2014).
Sanggaard, K.W. et al. Spider genomes provide insight into composition and evolution of venom and silk. Nat. Commun. 5, 3765 (2014).
Beckwitt, R. & Arcidiacono, S. Sequence conservation in the C-terminal region of spider silk proteins (Spidroin) from Nephila clavipes (Tetragnathidae) and Araneus bicentenarius (Araneidae). J. Biol. Chem. 269, 6661–6663 (1994).
Gatesy, J., Hayashi, C., Motriuk, D., Woods, J. & Lewis, R. Extreme diversity, conservation, and convergence of spider silk fibroin sequences. Science 291, 2603–2605 (2001).
Rising, A., Hjälm, G., Engström, W. & Johansson, J. N-terminal nonrepetitive domain common to dragline, flagelliform, and cylindriform spider silk proteins. Biomacromolecules 7, 3120–3124 (2006).
Garb, J.E., Ayoub, N.A. & Hayashi, C.Y. Untangling spider silk evolution with spidroin terminal domains. BMC Evol. Biol. 10, 243 (2010).
Blackledge, T.A., Kuntner, M. & Agnarsson, I. in Advances in Insect Physiology (ed. Casas, J.) Vol. 41, 175–262 (Burlington Academic Press, 2011).
Kuwana, Y., Sezutsu, H., Nakajima, K., Tamada, Y. & Kojima, K. High-toughness silk produced by a transgenic silkworm expressing spider (Araneus ventricosus) dragline silk protein. PLoS One 9, e105325 (2014).
Gosline, J.M., Guerette, P.A., Ortlepp, C.S. & Savage, K.N. The mechanical design of spider silks: from fibroin sequence to mechanical function. J. Exp. Biol. 202, 3295–3303 (1999).
Kuntner, M., Arnedo, M.A., Trontelj, P., Lokovšek, T. & Agnarsson, I. A molecular phylogeny of nephilid spiders: evolutionary history of a model lineage. Mol. Phylogenet. Evol. 69, 961–979 (2013).
Gaines, W.A.I. & IV & Marcotte, W.R.J. Jr. Identification and characterization of multiple Spidroin 1 genes encoding major ampullate silk proteins in Nephila clavipes. Insect Mol. Biol. 17, 465–474 (2008).
Bond, J.E. et al. Phylogenomics resolves a spider backbone phylogeny and rejects a prevailing paradigm for orb web evolution. Curr. Biol. 24, 1765–1771 (2014).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Hoff, K.J. & Stanke, M. WebAUGUSTUS—a web service for training AUGUSTUS and predicting genes in eukaryotes. Nucleic Acids Res. 41, W123–W128 (2013).
Colgin, M.A. & Lewis, R.V. Spider minor ampullate silk proteins contain new repetitive sequences and highly conserved non-silk-like “spacer regions”. Protein Sci. 7, 667–672 (1998).
Lewis, R.V. Spider silk: the unraveling of a mystery. Acc. Chem. Res. 25, 392–398 (1992).
Hayashi, C.Y. & Lewis, R.V. Evidence from flagelliform silk cDNA for the structural basis of elasticity and modular nature of spider silks. J. Mol. Biol. 275, 773–784 (1998).
Hayashi, C.Y., Shipley, N.H. & Lewis, R.V. Hypotheses that correlate the sequence, structure, and mechanical properties of spider silk proteins. Int. J. Biol. Macromol. 24, 271–275 (1999).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Vollrath, F. Spider webs and silks. Sci. Am. 266, 70–76 (1992).
Casem, M.L., Collin, M.A., Ayoub, N.A. & Hayashi, C.Y. Silk gene transcripts in the developing tubuliform glands of the Western black widow, Latrodectus hesperus. J. Arachnol. 38, 99–103 (2010).
Vollrath, F. Biology of spider silk. Int. J. Biol. Macromol. 24, 81–88 (1999).
Andersson, M. et al. Carbonic anhydrase generates CO2 and H+ that drive spider silk formation via opposite effects on the terminal domains. PLoS Biol. 12, e1001921 (2014).
Chaw, R.C., Correa-Garhwal, S.M., Clarke, T.H., Ayoub, N.A. & Hayashi, C.Y. Proteomic evidence for components of spider silk synthesis from black widow silk glands and fibers. J. Proteome Res. 14, 4223–4231 (2015).
Clarke, T.H. et al. Multi-tissue transcriptomics of the black widow spider reveals expansions, co-options, and functional processes of the silk gland gene toolkit. BMC Genomics 15, 365 (2014).
Lane, A.K., Hayashi, C.Y., Whitworth, G.B. & Ayoub, N.A. Complex gene expression in the dragline silk producing glands of the Western black widow (Latrodectus hesperus). BMC Genomics 14, 846 (2013).
Pouchkina, N.N., Stanchev, B.S. & McQueen-Mason, S.J. From EST sequence to spider silk spinning: identification and molecular characterisation of Nephila senegalensis major ampullate gland peroxidase NsPox. Insect Biochem. Mol. Biol. 33, 229–238 (2003).
Undheim, E.A.B. et al. A proteomics and transcriptomics investigation of the venom from the barychelid spider Trittame loki (brush-foot trapdoor). Toxins (Basel) 5, 2488–2503 (2013).
Hagn, F. et al. A conserved spider silk domain acts as a molecular switch that controls fibre assembly. Nature 465, 239–242 (2010).
Bard, F. et al. Functional genomics reveals genes involved in protein secretion and Golgi organization. Nature 439, 604–607 (2006).
Zhao, Y., Ayoub, N.A. & Hayashi, C.Y. Chromosome mapping of dragline silk genes in the genomes of widow spiders (Araneae, Theridiidae). PLoS One 5, e12804 (2010).
Hayashi, C.Y., Blackledge, T.A. & Lewis, R.V. Molecular and mechanical characterization of aciniform silk: uniformity of iterated sequence modules in a novel member of the spider silk fibroin gene family. Mol. Biol. Evol. 21, 1950–1959 (2004).
Starrett, J., Garb, J.E., Kuelbs, A., Azubuike, U.O. & Hayashi, C.Y. Early events in the evolution of spider silk genes. PLoS One 7, e38084 (2012).
Verstrepen, K.J., Jansen, A., Lewitter, F. & Fink, G.R. Intragenic tandem repeats generate functional variability. Nat. Genet. 37, 986–990 (2005).
Fondon, J.W. III & Garner, H.R. Molecular origins of rapid and continuous morphological evolution. Proc. Natl. Acad. Sci. USA 101, 18058–18063 (2004).
Suter, R.B. & Stratton, G.E. Scytodes vs. Schizocosa: predatory techniques and their morphological correlates. J. Arachnol. 33, 7–15 (2005).
Suter, R.B. & Stratton, G.E. Spitting performance parameters and their biomechanical implications in the spitting spider, Scytodes thoracica. J. Insect Sci. 9, 1–15 (2009).
Clements, R. & Li, D.Q. Regulation and non-toxicity of the spit from the pale spitting spider Scytodes pallida (Araneae: Scytodidae). Ethology 111, 311–321 (2005).
Zobel-Thropp, P.A., Correa, S.M., Garb, J.E. & Binford, G.J. Spit and venom from Scytodes spiders: a diverse and distinct cocktail. J. Proteome Res. 13, 817–835 (2014).
Teulé, F. et al. Silkworms transformed with chimeric silkworm/spider silk genes spin composite silk fibers with improved mechanical properties. Proc. Natl. Acad. Sci. USA 109, 923–928 (2012).
Bolger, A.M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Xu, H. et al. FastUniq: a fast de novo duplicates removal tool for paired short reads. PLoS One 7, e52249 (2012).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl. Acad. Sci. USA 108, 1513–1518 (2011).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Wences, A.H. & Schatz, M.C. Metassembler: merging and optimizing de novo genome assemblies. Genome Biol. 16, 207 (2015).
Bradnam, K.R. et al. Assemblathon 2: evaluating de novo methods of genome assembly in three vertebrate species. Gigascience 2, 10 (2013).
Simão, F.A., Waterhouse, R.M., Ioannidis, P., Kriventseva, E.V. & Zdobnov, E.M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Grabherr, M.G. et al. Full-length transcriptome assembly from RNA–Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Haas, B.J. et al. De novo transcript sequence reconstruction from RNA–seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Wu, T.D. & Watanabe, C.K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Dobin, A. et al. STAR: ultrafast universal RNA–seq aligner. Bioinformatics 29, 15–21 (2013).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Slater, G.S.C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Lowe, T.M. & Eddy, S.R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Mitchell, A. et al. The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res. 43, D213–D221 (2015).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Rogers, M.B. et al. Intrahost dynamics of antiviral resistance in influenza A virus reflect complex patterns of segment linkage, reassortment, and natural selection. MBio 6, e02464–14 (2015).
English, A.C. et al. Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS One 7, e47768 (2012).
Chaisson, M.J. & Tesler, G. Mapping single molecule sequencing reads using basic local alignment with successive refinement (BLASR): application and theory. BMC Bioinformatics 13, 238 (2012).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv https://arxiv.org/abs/1303.3997 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Watterson, G.A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 7, 256–276 (1975).
Scharlaken, B. et al. Reference gene selection for insect expression studies using quantitative real-time PCR: the head of the honeybee, Apis mellifera, after a bacterial challenge. J. Insect Sci. 8, 1–10 (2008).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Acknowledgements
We thank our many colleagues who kindly shared their expertise, time, resources, and insights with this project: J. Coddington, V. Aggarwala, K. Siewert, K. Johnson, K. Lorenz, R. Aikens, O. Yörük, K. Gawronski, D. Cousminer, K. Redmond (for spider searching), R. Hansen, T. Abel, J. Geskes, S. Khetarpal, C. Brown, K. Hayer, A. Ahmad, G. Grant, J. Growe, L. Francey, Y. Lee, J. Schug, H. Zillges, J. Grubb, C. Theodorou, A. Srinivasan, C. Calafut, J. Szostek, R. Monyak, T. Jongens, L. Hennessy, S. Teegarden, G. FitzGerald, L. Hood, I. Silverman, B. Gregory, R. Sebra, K. Childs, C. Holt, A. English, F.J. Barton, M.L. Barton, J. Retief, and T. Orpin. We are also indebted to the three anonymous reviewers for helpful comments on the manuscript. B.F.V. is grateful for support of the work from the Alfred P. Sloan Foundation (BR2012-087). Genomic assembly was conducted on the PMACS HPC infrastructure at the University of Pennsylvania, funded in part by NIH Special Instrumentation Grant 1S10OD012312-NIH.
Author information
Authors and Affiliations
Contributions
B.F.V., L.H., and I.A. conceived of the project. P.L.B., B.F.V., N.F.L., and J.B.H. designed the experiments. B.F.V. and J.B.H. contributed reagents and materials. P.L.B., L.H., S.M.C.-G., and C.Y.H. provided samples. C.Y.H., S.M.C.-G., and L.H. performed specimen dissections. P.L.B. conducted all bench experiments. P.L.B. performed all assembly and annotation pipelines. P.L.B., B.F.V., and D.N.N. ran motif searches. P.L.B., N.F.L., and E.J.K. performed expression analyses. P.L.B., B.F.V., C.Y.H., J.B.H., M.K., L.H., and I.A. reviewed analyses. P.L.B. and B.F.V. prepared figures and tables. P.L.B. and B.F.V. wrote the first draft of the manuscript. All authors reviewed drafts of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 The golden orb-weaver spider’s morphology, reported silk gland anatomy, and web construction.
(a) Photographs of N. clavipes showing an adult female at the center hub of her orb web (left) and a view of the spinneret silk-extruding organs on the underside of the female abdomen (right). (b) Silk gland anatomy of N. clavipes, showing the seven different female araneoid gland morphologies found in the abdomen and the different classes of silk proteins produced. Each silk class has specific physical characteristics; for example, the minor and major ampullate spidroins produce silks with great tensile strength, flagelliform silk has great extensibility, aggregate silks are non-fibrous stick glue, etc. This illustration (inspired by ref. 52) exhibits one set of silk glands and spinnerets from a bilateral pair, and indicates that each gland type produces a specific type of silk. However our expression data (Fig. 5a) suggest that this is not the case, supporting previous findings48,50,53 that individual glands can express multiple classes of spidroins. Note: the gland type coloration scheme and corresponding silk use pictograms defined here are used in later figures. (c) Putative applications of spider silk types in web construction (web diagram adapted from ref. 54), as described in previous studies. (i) Web building and maintenance: major ampullate silk is used for bridgelines and web radii; minor ampullate silk is used for temporary spiral; piriform attaches fibers together and to substrates; flagelliform is used for the capture spiral; aggregate silks are sticky, aiding in adherence and prey capture. (ii) Prey wrapping: aciniform (top inset photo). (iii) Silk egg casings: tubuliform (bottom inset photo). References for silk classes and their purported uses are listed in the main text. (Photos provided by P.L.B.)
Supplementary Figure 2 Maximum-likelihood phylogenetic gene tree of 28 N. clavipes spidroins in the context of 55 spidroins from other spider taxa.
The spidroin gene tree is rooted with a Bothriocyrtum californicum fibroin sequence (B.c. fibroin1; accession HM752562) and is based on multiple-sequence alignment (MSA) using the first ~130 amino acid residues of the N-terminal domain encoded by each gene. MSA was performed with Geneious, Clustal, and BLOSUM62, and the consensus tree was built with PhyML (Supplementary Note). Bootstrap proportions >50 (based on 1,000 replicates) are shown to the left of their respective nodes. Colors follow the gland/spidroin class designations shown in Supplementary Figure 1. Codes and accession numbers for different spidroins and taxa are listed in Supplementary Table 10.
Supplementary Figure 3 Maximum-likelihood phylogenetic gene trees for the catalog of 28 spidroins identified in N. clavipes.
(a,b) Unrooted maximum-likelihood phylogenic trees for the catalog of 28 spidroins identified in N. clavipes, shown as both transformed (a) and non-transformed (b) layouts. Both trees are based on a multiple-sequence alignment (MSA) using the first ~130 amino acid residues of the N-terminal domain for each N. clavipes spidroin. MSA was performed with Geneious, Clustal, and BLOSUM62, and the consensus tree was built with PhyML (Supplementary Note). Bootstrap proportions >50 (based on 1,000 replicates) are shown to the left of their respective nodes. Colors follow the gland/spidroin class designations shown in Supplementary Figure 1. Codes for N. clavipes spidroins are listed in Supplementary Table 12.
Supplementary Figure 4 Agarose gel images of long-range PCR–amplified MiSp sequences used for validation of draft assembly, scaffold bridging, and gap closure.
The top panel highlights a single lane with an LR-PCR reaction (golden rectangle) for MiSp-c. The bottom panel highlights four lanes with LR-PCR reactions (golden rectangle) for MiSp-d. In both cases, multiple large bands are visible, indicating amplification of multiple targets that presumably represent genomic regions with high sequence similarity to the binding sites of the oligonucleotide primers used to isolate both MiSp types.
Supplementary Figure 5 Distribution of amino acid frequency for N. clavipes ‘gold’ gene models.
Amino acid frequency distributions were calculated for all 20 amino acids for all mRNA transcripts from the gold gene model set (n = 17,989 mRNA sequences). Several spidroins were found at the extreme ends of the individual amino acid distributions (Supplementary Fig. 5 and Supplementary Note). Box-and-whisker plots show the range of frequency values (y axis) for the given amino acid residues (x axis). Thick black center lines represent median values. Upper whiskers represent the largest observation ≤ the upper quartile (Q3) + 1.5 interquartile range (IQR), and lower whiskers represent the smallest observation ≥ the lower quartile (Q1) – 1.5 IQR.
Supplementary Figure 6 Distribution of amino acid frequency for 28 N. clavipes spidroin genes.
Amino acid frequency distributions were calculated for all 20 amino acids for all N. clavipes spidroin genes (n = 28 sequences). Box-and-whisker plots show the range of frequency values (y axis) for the given amino acid residues (x axis). Thick black center lines represent median values. Upper whiskers represent the largest observation ≤ the upper quartile (Q3) + 1.5 interquartile range (IQR), and the lower whiskers represent the smallest observation ≥ the lower quartile (Q1) – 1.5 IQR. Overall, spidroins exhibit enrichment of alanine, glycine, and serine residues, which have significantly different proportions when compared to 17,989 mRNA sequences from the gold gene set (Wilcoxon rank-sum test; Supplementary Fig. 5 and Supplementary Note). **P < 0.01.
Supplementary Figure 7 Shared and private motif occurrences in N. clavipes spidroins.
Bar graph comparing the number of shared (gold) versus private (dark gray) distinct repetitive motif occurrences observed in the different N. clavipes spidroins (n = 28 sequences).
Supplementary Figure 8 Shared and private cassette occurrences in N. clavipes spidroins.
Bar graph comparing the number of shared (gold) and private (dark gray) distinct repetitive cassette occurrences observed in the different N. clavipes spidroins (n = 28 sequences).
Supplementary Figure 9 RNA–seq expression patterns of spidroin genes in 13 N. clavipes tissue samples.
Heat map showing the absolute number of normalized RNA–seq reads that map to spidroin transcripts, assayed in ten individual silk glands, one venom gland isolate, and two brain isolates collected from two non-gravid females, Nep-008 and Nep-009. Owing to extensive sequence similarity between MaSp-b and MaSp-c, it was not possible to distinguish between reads that mapped to these two spidroins; thus, data for these two transcripts are presented together as “MaSp-b,c”. Reads mapping to MaSp-h and AgSp-c exceeded the heat map’s informative range; thus, we have included bar graph insets (right) confirming that reads mapping to MaSp-h (top inset) and AgSp-c (bottom inset) are substantially more abundant in silk glands than in venom gland or brain.
Supplementary Figure 10 Distributions of relative expression values for 29 N. clavipes genes in seven tissue types.
Box-and-whisker plots of the relative expression for all 28 N. clavipes spidroin genes and 1 venom gene (PR-1) in tissue dissections (n = 3 independent-specimen biological replicates per tissue) assayed by qPCR. Tissues included venom glands, five anatomically distinct silk glands, and ‘other’ silk glands (aciniform and piriform glands attached to the spinneret) and are shown left of the y axis, whereas relative expression (2−ΔΔCT method46) is depicted on the y axis (log10 scale) organized in rows by tissue type. Box-and-whisker plots show the range of expression values for the given genes (x axis) relative to RPL13a (housekeeping gene) expression and normalized to leg tissue. Thick black center lines represent median values. Upper whiskers represent the largest observation ≤ the upper quartile (Q3) + 1.5 interquartile range (IQR), and the lower whiskers represent the smallest observation ≥ the lower quartile (Q1) – 1.5 IQR. Asterisks indicate a single silk gland type exhibiting significantly greater expression values for a given gene versus all other silk gland types together (Wilcoxon rank-sum test). **P < 0.01.
Supplementary Figure 11 Mean relative expression values of 29 N. clavipes genes in seven tissue types.
Heat map showing the relative expression of N. clavipes spidroin loci in tissue dissections (n = 3 biological replicates per tissue) assayed by qPCR. Tissues included venom glands, five anatomically distinct silk glands, and ‘other’ silk glands (aciniform and piriform glands attached to the spinneret) and are shown on the x axis, with spidroins arranged on the y axis. The heat map panels depict relative mean fold change in gene expression (2−ΔΔCT method46) per tissue (distinct tissue dissections from n = 3 individuals) over RPL13a and normalized to leg tissue.
Supplementary Figure 12 RNA–seq expression patterns of SSTs in 13 N. clavipes tissue samples.
Heat map showing the absolute number of normalized reads that map to 649 non-spidroin silk gland–specific transcripts (SSTs), assayed in ten individual silk glands, one venom gland, and two brain isolates collected from two non-gravid females, Nep-008 and Nep-009. SSTs are vertically clustered based on the filtering method used for discovery (Supplementary Note), as noted by colored vertical bars at the right of the heat map. The categories defined on the left are described in Supplementary Table 15.
Supplementary Figure 13 Polymorphism levels of genes and genic features in the N. clavipes genome.
(a) Box-and-whisker plot comparing the distribution of θW values43 derived (from SNP counts) for 14,025 gold gene sequences in comparison to the distribution of θW values for 28 N. clavipes spidroins. Box-and-whisker plots show the range of θW values for each gene set. Thick black center lines represent median values. Upper whiskers represent the largest observation ≤ the upper quartile (Q3) + 1.5 interquartile range (IQR), and the lower whiskers represent the smallest observation ≥ the lower quartile (Q1) – 1.5 IQR. Asterisks indicate the 28 N. clavipes spidroin genes that exhibit significantly greater θW values than the collected gold gene set (Wilcoxon rank-sum test; Supplementary Note). **P < 0.01. (b) Vertical bar graph showing the mean θW values for 11 genomic feature categories, including many gold gene model subfeatures, in comparison to the mean θW values for N. clavipes spidroins, silk N termini, and silk C termini. (c) Bar graph depicting the θW values for individual N. clavipes spidroins.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 1–12 and Supplementary Note (PDF 4813 kb)
Supplementary Tables 13–15
Supplementary Tables 13–15 (XLSX 123 kb)
Supplementary Data
N. clavipes qPCR 2–ΔΔCT values for all loci and replicates. Includes all results of statistical tests. (XLSX 168 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) License. The images or other third party material in this article is included in the article’s Creative Commons license, unless otherwise indicated in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Babb, P., Lahens, N., Correa-Garhwal, S. et al. The Nephila clavipes genome highlights the diversity of spider silk genes and their complex expression. Nat Genet 49, 895–903 (2017). https://doi.org/10.1038/ng.3852
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3852
This article is cited by
-
Chromosome-level genome of spider Pardosa pseudoannulata and cuticle protein genes in environmental stresses
Scientific Data (2024)
-
Genomic and transcriptomic analyses support a silk gland origin of spider venom glands
BMC Biology (2023)
-
Lampshade web spider Ectatosticta davidi chromosome-level genome assembly provides evidence for its phylogenetic position
Communications Biology (2023)
-
Digestive enzymes and sphingomyelinase D in spiders without venom (Uloboridae)
Scientific Reports (2023)
-
Regionalization of cell types in silk glands of Larinioides sclopetarius suggest that spider silk fibers are complex layered structures
Scientific Reports (2023)