Abstract
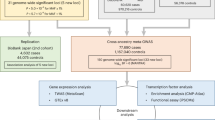
Atrial fibrillation affects more than 33 million people worldwide and increases the risk of stroke, heart failure, and death1,2. Fourteen genetic loci have been associated with atrial fibrillation in European and Asian ancestry groups3,4,5,6,7. To further define the genetic basis of atrial fibrillation, we performed large-scale, trans-ancestry meta-analyses of common and rare variant association studies. The genome-wide association studies (GWAS) included 17,931 individuals with atrial fibrillation and 115,142 referents; the exome-wide association studies (ExWAS) and rare variant association studies (RVAS) involved 22,346 cases and 132,086 referents. We identified 12 new genetic loci that exceeded genome-wide significance, implicating genes involved in cardiac electrical and structural remodeling. Our results nearly double the number of known genetic loci for atrial fibrillation, provide insights into the molecular basis of atrial fibrillation, and may facilitate the identification of new potential targets for drug discovery8.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
11 May 2017
In the version of this article initially published online, the authors were incorrectly defined as members of the AFGen consortium in the author list. The members of the consortium are listed in the Supplementary Note. The error has been corrected in the print, PDF and HTML versions of this article.
References
Chugh, S.S. et al. Worldwide epidemiology of atrial fibrillation: a Global Burden of Disease 2010 Study. Circulation 129, 837–847 (2014).
January, C.T. et al. 2014 AHA/ACC/HRS guideline for the management of patients with atrial fibrillation: executive summary: a report of the American College of Cardiology/American Heart Association Task Force on practice guidelines and the Heart Rhythm Society. J. Am. Coll. Cardiol. 65, 2246–2280 (2014).
Gudbjartsson, D.F. et al. Variants conferring risk of atrial fibrillation on chromosome 4q25. Nature 448, 353–357 (2007).
Benjamin, E.J. et al. Variants in ZFHX3 are associated with atrial fibrillation in individuals of European ancestry. Nat. Genet. 41, 879–881 (2009).
Ellinor, P.T. et al. Common variants in KCNN3 are associated with lone atrial fibrillation. Nat. Genet. 42, 240–244 (2010).
Ellinor, P.T. et al. Meta-analysis identifies six new susceptibility loci for atrial fibrillation. Nat. Genet. 44, 670–675 (2012).
Sinner, M.F. et al. Integrating genetic, transcriptional, and functional analyses to identify 5 novel genes for atrial fibrillation. Circulation 130, 1225–1235 (2014).
Nelson, M.R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Lloyd-Jones, D.M. et al. Lifetime risk for development of atrial fibrillation: the Framingham Heart Study. Circulation 110, 1042–1046 (2004).
Gudbjartsson, D.F. et al. Large-scale whole-genome sequencing of the Icelandic population. Nat. Genet. 47, 435–444 (2015).
Holm, H. et al. A rare variant in MYH6 is associated with high risk of sick sinus syndrome. Nat. Genet. 43, 316–320 (2011).
Weeke, P. et al. Exome sequencing implicates an increased burden of rare potassium channel variants in the risk of drug-induced long QT interval syndrome. J. Am. Coll. Cardiol. 63, 1430–1437 (2014).
Seals, D.F. et al. The adaptor protein Tks5/Fish is required for podosome formation and function, and for the protease-driven invasion of cancer cells. Cancer Cell 7, 155–165 (2005).
Laumet, G. et al. A study of the association between the ADAM12 and SH3PXD2A (SH3MD1) genes and Alzheimer's disease. Neurosci. Lett. 468, 1–2 (2010).
Cejudo-Martin, P. et al. Genetic disruption of the Sh3pxd2a gene reveals an essential role in mouse development and the existence of a novel isoform of Tks5. PLoS One 9, e107674 (2014).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Pe'er, I., Yelensky, R., Altshuler, D. & Daly, M.J. Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet. Epidemiol. 32, 381–385 (2008).
Hoggart, C.J., Clark, T.G., De Iorio, M., Whittaker, J.C. & Balding, D.J. Genome-wide significance for dense SNP and resequencing data. Genet. Epidemiol. 32, 179–185 (2008).
Dudbridge, F. & Gusnanto, A. Estimation of significance thresholds for genomewide association scans. Genet. Epidemiol. 32, 227–234 (2008).
Sham, P.C. & Purcell, S.M. Statistical power and significance testing in large-scale genetic studies. Nat. Rev. Genet. 15, 335–346 (2014).
Lumley, T., Brody, J., Dupuis, J. & Cupples, A. Meta-analysis of a rare-variant association test. (2012).
Lee, S., Wu, M.C. & Lin, X. Optimal tests for rare variant effects in sequencing association studies. Biostatistics 13, 762–775 (2012).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375, S1–S3 (2012).
Pers, T.H., Timshel, P. & Hirschhorn, J.N. SNPsnap: a Web-based tool for identification and annotation of matched SNPs. Bioinformatics 31, 418–420 (2015).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Boyle, A.P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Sherry, S.T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP–trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Arnold, M., Raffler, J., Pfeufer, A., Suhre, K. & Kastenmüller, G. SNiPA: an interactive, genetic variant–centered annotation browser. Bioinformatics 31, 1334–1336 (2015).
Deshmukh, A. et al. Left atrial transcriptional changes associated with atrial fibrillation susceptibility and persistence. Circ Arrhythm Electrophysiol 8, 32–41 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J.R. Stat. Soc. 57, 289–300 (1995).
Kuriyama, S. et al. The Tohoku Medical Megabank Project: design and mission. J. Epidemiol. 26, 493–511 (2016).
Pers, T.H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Neurology Working Group of the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium. Identification of additional risk loci for stroke and small vessel disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 15, 695–707 (2016).
Malik, R. et al. Low-frequency and common genetic variation in ischemic stroke: The METASTROKE collaboration. Neurology 86, 1217–1226 (2016).
Acknowledgements
A full list of acknowledgments appears in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
I.E.C., C.R., X.Y., T.T., K.L.L., E.J.B., S.A.L., M.R., B.G., and P.T.E. wrote and edited the manuscript. All authors contributed to and discussed the results and commented on the manuscript. GWAS and ExWAS analyses: A.V.S., N.A.B., M.M.-N., I.S., C.S., P.E.W., S.A., S. Thériault, J.A.B., J.C.B., H.L., J. Huffman, J.Y., X.G., F.R., M.N.N., D.E.A., G.P., S.-K.L., Y.K., M. Kähönen, A.C.P., A.R.H., J.S., L.-P.L., M.A., M.E.K., J.G.S., R.M., S.G., S. Trompet, M.D., S.W., J.A.W., D.I.C., M.V.P., Q.Y., T.B.H., M.F.S., J.S., D.R.v.W. Individual data set quality control and GWAS and ExWAS meta-analyses: I.E.C., K.L.L., C.R., X.Y., M.R., B.G., Y.P.H., N.V., J.E.S. Replication in METASTROKE and Neuro-CHARGE: Q.Y., J.C.H., S.D., G.C., B.B.W. Replication in UK Biobank: S.K., D.K., C.N.-C. Replication in BioBank Japan: S.-K.L., Y.K., M. Kubo, T.T. Replication in African-American population: R.D., D.J.R., S.H.S., A.S. CCAF eQTL analyses: J.B., M.K.C., D.v.W., J.D.S. Functional annotation: I.E.C., S.H.C., L.-C.W., M. Li, C.R., M.C., N.R.T., S.C. Pathway analyses: H.L.
Corresponding author
Ethics declarations
Competing interests
P.T.E. is the principal investigator on a grant from Bayer HealthCare to the Broad Institute focused on the genetics and therapeutics of atrial fibrillation. The remaining authors have no disclosures.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
Integrated supplementary information
Supplementary Figure 1 QQ plot for the combined-ancestry GWAS meta-analysis.
QQ plot displaying the observed versus the expected –log10 of the P value for each variant tested in the combined-ancestry GWAS meta-analysis. λ represents the genomic inflation factor.
Supplementary Figure 2 Regional plots from combined-ancestry ExWAS meta-analysis.
The most significant variant at each locus is plotted (purple, diamond-shaped) and identified by rsID. Each dot in the plots represents a single variant present in our results, and the color of the dot indicates the degree of linkage disequilibrium with the most significant variant, as shown in the top left color chart in each panel. The lower part of each panel shows the locations of genes in the respective loci. r2, degree of linkage disequilibrium; chr., chromosome. Regional plots were created using LocusZoom.
Supplementary Figure 3 QQ plot for the combined-ancestry ExWAS meta-analysis.
QQ plot displaying the observed versus the expected –log10 of the P value for each variant tested in the combined-ancestry ExWAS meta-analysis. λ represents the genomic inflation factor.
Supplementary Figure 4 Manhattan plot of results from African-American-ancestry GWAS meta-analysis.
Genetic loci that have also previously been associated with atrial fibrillation through GWAS are highlighted in blue. The dashed line represents the significance threshold (5 × 10–8). The gene names represent the gene in closest proximity to the most significant variant at each locus.
Supplementary Figure 5 Manhattan plot of results from European-ancestry GWAS meta-analysis.
New genetic loci for atrial fibrillation are highlighted in red, whereas loci that have also previously been associated with atrial fibrillation through GWAS are highlighted in blue. The dashed line represents the significance threshold (5 × 10–8). The gene names represent the gene in closest proximity to the most significant variant at each locus. There is a break in the y axis to increase the resolution of the genetic loci near the genome-wide significance threshold.
Supplementary Figure 6 Manhattan plot of results from Asian-ancestry GWAS meta-analysis.
New genetic loci for atrial fibrillation are highlighted in red, whereas loci that have also previously been associated with atrial fibrillation through GWAS are highlighted in blue. The dashed line represents the significance threshold (5 × 10–8). The gene names represent the gene in closest proximity to the most significant variant at each locus.
Supplementary Figure 7 Manhattan plot of results from incident atrial fibrillation GWAS meta-analysis in Europeans.
Genetic loci that have also previously been associated with atrial fibrillation through GWAS are highlighted in blue. The dashed line represents the significance threshold (5 × 10–8). The gene names represent the gene in closest proximity to the most significant variant at each locus. There is a break in the y axis to increase the resolution of the genetic loci near the genome-wide significance threshold.
Supplementary Figure 8 Manhattan plot of results from prevalent atrial fibrillation GWAS meta-analysis in Europeans.
New genetic loci for atrial fibrillation are highlighted in red, whereas loci that have also previously been associated with atrial fibrillation through GWAS are highlighted in blue. The dashed line represents the significance threshold (5 × 10–8). The gene names represent the gene in closest proximity to the most significant variant at each locus. There is a break in the y axis to increase the resolution of the genetic loci near the genome-wide significance threshold.
Supplementary Figure 9 Distinct loci on chromosomes 1 and 10, as demonstrated using approximate joint and conditional association analysis in European-ancestry studies with GCTA software.
All conditional analyses were performed using the European-ancestry results only, with a European-ancestry reference population from the Framingham Heart Study. Regional plots were created using Locus Zoom software1 with LD information from the European ancestry 1000 Genomes reference population. (a,b) Regional plots of the independent signals at chromosome 1q24; the new METTL11B locus (a) and the replicated PRRX1 locus (b). (c,d) Regional plots of the independent signals at chromosome 10q24; the new SH3PXD2A locus (c) and the replicated NEURL1 locus (d).
Supplementary Figure 10 Atrial fibrillation–associated loci display pleiotropy across clinical, electrocardiographic, and echocardiographic cardiac phenotypes.
Diagram showing overlap of genetic associations between cardiac phenotypes, identified through interrogation of the NHGRI-EBI GWAS catalog2. Gene names are in italics and represent genetic loci identified through GWAS (Supplementary Table 13). AF, atrial fibrillation; HR, heart rate; LVIDD, left-ventricle internal diastolic diameter on echocardiography; PR-I, PR interval; PR-S, PR segment; QRS, QRS interval; QT, QT interval.
Supplementary Figure 11 Atrial fibrillation–associated loci are enriched for functional elements.
(a–f) Median overlap of loci by phastCons 46-way primate conserved elements (a), phastCons 46-way mammalian conserved elements (b), Roadmap Epigenomics cardiac H3K27ac gapped peaks (R atrium, L ventricle, R ventricle, aorta) (c), any Roadmap Epigenomics H3K27ac gapped peak (98 cell types) (d), ENCODE DNaseHS cardiac sites (cardiac fibroblasts, atrial fibroblasts, cardiac myocytes) (e), and ENCODE DNaseHS master sites (125 cell types) (f). *P < 0.05, **P < 0.01, ***P < 0.001, one-tailed bootstrapping (n = 1,000). Whiskers, interquartile range; AF, atrial fibrillation–associated loci (n = 24); GWAS, NHGRI-EBI GWAS catalog associated loci (n = 3,381); 1000G, 1000 Genomes control loci based on SNPsnap matched variants to atrial fibrillation GWAS hits (n = 9,093).
Supplementary Figure 12 Atrial fibrillation–associated loci are enriched for eQTLs.
The figure shows the proportion of loci with at least one GTEx eQTL for all tissues available in the GTEx database. Only the three top tissues showed significant enrichment of eQTLs for atrial fibrillation loci (pancreas, left ventricle, and tibial artery tissue) AF, atrial fibrillation–associated loci (n = 24); GWAS, NHGRI-EBI GWAS catalog associated loci (n = 3,381); 1000G, 1000 Genomes control loci based on SNPsnap matched variants to atrial fibrillation GWAS hits (n = 9,093).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 3–9, 12–16 and 21–29, and Supplementary Note. (PDF 4541 kb)
Supplementary Tables
Supplementary Tables 1, 2, 10, 11 and 17–20. (XLSX 268 kb)
Rights and permissions
About this article
Cite this article
Christophersen, I., Rienstra, M., Roselli, C. et al. Large-scale analyses of common and rare variants identify 12 new loci associated with atrial fibrillation. Nat Genet 49, 946–952 (2017). https://doi.org/10.1038/ng.3843
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3843
This article is cited by
-
Sex-specific genetic architecture of blood pressure
Nature Medicine (2024)
-
Exploring the causal association of rheumatoid arthritis with atrial fibrillation: a Mendelian randomization study
Clinical Rheumatology (2024)
-
SNX17 Mediates Dendritic Spine Maturation via p140Cap
Molecular Neurobiology (2024)
-
Pathophysiology and clinical relevance of atrial myopathy
Basic Research in Cardiology (2024)
-
Genomic approaches to identify and investigate genes associated with atrial fibrillation and heart failure susceptibility
Human Genomics (2023)