Abstract
We identify SMARCD2 (SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily D, member 2), also known as BAF60b (BRG1/Brahma-associated factor 60b), as a critical regulator of myeloid differentiation in humans, mice, and zebrafish. Studying patients from three unrelated pedigrees characterized by neutropenia, specific granule deficiency, myelodysplasia with excess of blast cells, and various developmental aberrations, we identified three homozygous loss-of-function mutations in SMARCD2. Using mice and zebrafish as model systems, we showed that SMARCD2 controls early steps in the differentiation of myeloid–erythroid progenitor cells. In vitro, SMARCD2 interacts with the transcription factor CEBPɛ and controls expression of neutrophil proteins stored in specific granules. Defective expression of SMARCD2 leads to transcriptional and chromatin changes in acute myeloid leukemia (AML) human promyelocytic cells. In summary, SMARCD2 is a key factor controlling myelopoiesis and is a potential tumor suppressor in leukemia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Krumsiek, J., Marr, C., Schroeder, T. & Theis, F.J. Hierarchical differentiation of myeloid progenitors is encoded in the transcription factor network. PLoS One 6, e22649 (2011).
Orkin, S.H. & Zon, L.I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008).
Krosl, J. et al. A mutant allele of the Swi/Snf member BAF250a determines the pool size of fetal liver hemopoietic stem cell populations. Blood 116, 1678–1684 (2010).
Friedman, A.D. Transcriptional control of granulocyte and monocyte development. Oncogene 26, 6816–6828 (2007).
Griffin, C.T., Brennan, J. & Magnuson, T. The chromatin-remodeling enzyme BRG1 plays an essential role in primitive erythropoiesis and vascular development. Development 135, 493–500 (2008).
Huang, H.T. et al. A network of epigenetic regulators guides developmental haematopoiesis in vivo. Nat. Cell Biol. 15, 1516–1525 (2013).
Álvarez-Errico, D., Vento-Tormo, R., Sieweke, M. & Ballestar, E. Epigenetic control of myeloid cell differentiation, identity and function. Nat. Rev. Immunol. 15, 7–17 (2015).
Cedar, H. & Bergman, Y. Epigenetics of haematopoietic cell development. Nat. Rev. Immunol. 11, 478–488 (2011).
Cairns, B.R. The logic of chromatin architecture and remodelling at promoters. Nature 461, 193–198 (2009).
de la Serna, I.L., Ohkawa, Y. & Imbalzano, A.N. Chromatin remodelling in mammalian differentiation: lessons from ATP-dependent remodellers. Nat. Rev. Genet. 7, 461–473 (2006).
Tolstorukov, M.Y. et al. Swi/Snf chromatin remodeling/tumor suppressor complex establishes nucleosome occupancy at target promoters. Proc. Natl. Acad. Sci. USA 110, 10165–10170 (2013).
Wilson, B.G. & Roberts, C.W. SWI/SNF nucleosome remodellers and cancer. Nat. Rev. Cancer 11, 481–492 (2011).
Buscarlet, M. et al. Essential role of BRG, the ATPase subunit of BAF chromatin remodeling complexes, in leukemia maintenance. Blood 123, 1720–1728 (2014).
Jojic, V. et al. Identification of transcriptional regulators in the mouse immune system. Nat. Immunol. 14, 633–643 (2013).
Alajem, A. et al. Differential association of chromatin proteins identifies BAF60a/SMARCD1 as a regulator of embryonic stem cell differentiation. Cell Rep. 10, 2019–2031 (2015).
Lickert, H. et al. Baf60c is essential for function of BAF chromatin remodelling complexes in heart development. Nature 432, 107–112 (2004).
Hall, C., Flores, M.V., Storm, T., Crosier, K. & Crosier, P. The zebrafish lysozyme C promoter drives myeloid-specific expression in transgenic fish. BMC Dev. Biol. 7, 42 (2007).
Liongue, C., Hall, C.J., O'Connell, B.A., Crosier, P. & Ward, A.C. Zebrafish granulocyte colony-stimulating factor receptor signaling promotes myelopoiesis and myeloid cell migration. Blood 113, 2535–2546 (2009).
Renshaw, S.A. et al. A transgenic zebrafish model of neutrophilic inflammation. Blood 108, 3976–3978 (2006).
Koulnis, M. et al. Identification and analysis of mouse erythroid progenitors using the CD71/TER119 flow-cytometric assay. J. Vis. Exp. 5, 2809 (2011).
Ramirez-Carrozzi, V.R. et al. A unifying model for the selective regulation of inducible transcription by CpG islands and nucleosome remodeling. Cell 138, 114–128 (2009).
Iriyama, N. et al. Enhancement of differentiation induction and upregulation of CCAAT/enhancer-binding proteins and PU.1 in NB4 cells treated with combination of ATRA and valproic acid. Int. J. Oncol. 44, 865–873 (2014).
Tanaka, M., Gombart, A.F., Koeffler, H.P. & Shiohara, M. Expression of bactericidal/permeability-increasing protein requires C/EBPɛ. Int. J. Hematol. 85, 304–311 (2007).
Lekstrom-Himes, J. & Xanthopoulos, K.G. CCAAT/enhancer binding protein ɛ is critical for effective neutrophil-mediated response to inflammatory challenge. Blood 93, 3096–3105 (1999).
Lekstrom-Himes, J.A., Dorman, S.E., Kopar, P., Holland, S.M. & Gallin, J.I. Neutrophil-specific granule deficiency results from a novel mutation with loss of function of the transcription factor CCAAT/enhancer binding protein ɛ. J. Exp. Med. 189, 1847–1852 (1999).
Hu, H. et al. Maturity-dependent fractionation of neutrophil progenitors: a new method to examine in vivo expression profiles of differentiation-regulating genes. Exp. Hematol. 40, 675–681 (2012).
Gkikopoulos, T. et al. A role for Snf2-related nucleosome-spacing enzymes in genome-wide nucleosome organization. Science 333, 1758–1760 (2011).
Wada, T. et al. A novel in-frame deletion in the leucine zipper domain of C/EBPɛ leads to neutrophil-specific granule deficiency. J. Immunol. 195, 80–86 (2015).
Gallin, J.I. et al. Human neutrophil-specific granule deficiency: a model to assess the role of neutrophil-specific granules in the evolution of the inflammatory response. Blood 59, 1317–1329 (1982).
Breton-Gorius, J., Mason, D.Y., Buriot, D., Vilde, J.L. & Griscelli, C. Lactoferrin deficiency as a consequence of a lack of specific granules in neutrophils from a patient with recurrent infections. Detection by immunoperoxidase staining for lactoferrin and cytochemical electron microscopy. Am. J. Pathol. 99, 413–428 (1980).
Komiyama, A., Morosawa, H., Nakahata, T., Miyagawa, Y. & Akabane, T. Abnormal neutrophil maturation in a neutrophil defect with morphologic abnormality and impaired function. J. Pediatr. 94, 19–25 (1979).
Khanna-Gupta, A. et al. Growth factor independence-1 (Gfi-1) plays a role in mediating specific granule deficiency (SGD) in a patient lacking a gene-inactivating mutation in the C/EBPɛ gene. Blood 109, 4181–4190 (2007).
Koike, M. et al. C/EBP-ɛ: chromosomal mapping and mutational analysis of the gene in leukemia and preleukemia. Leuk. Res. 21, 833–839 (1997).
Verbeek, W., Wachter, M., Lekstrom-Himes, J. & Koeffler, H.P. C/EBPɛ−/− mice: increased rate of myeloid proliferation and apoptosis. Leukemia 15, 103–111 (2001).
Yamanaka, R. et al. Impaired granulopoiesis, myelodysplasia, and early lethality in CCAAT/enhancer binding protein ɛ–deficient mice. Proc. Natl. Acad. Sci. USA 94, 13187–13192 (1997).
Shi, J. et al. Role of SWI/SNF in acute leukemia maintenance and enhancer-mediated Myc regulation. Genes Dev. 27, 2648–2662 (2013).
Cruickshank, V.A. et al. SWI/SNF subunits SMARCA4, SMARCD2 and DPF2 collaborate in MLL-rearranged leukaemia maintenance. PLoS One 10, e0142806 (2015).
Madan, V. et al. Comprehensive mutational analysis of primary and relapse acute promyelocytic leukemia. Leukemia 30, 1672–1681 (2016).
Priam, P. et al. SMARCD2 subunit of the SWI/SNF chromatin-remodeling complex mediates granulopoiesis through a CEBPɛ- dependent mechanism. Nat. Genet. http://dx.doi.org/10.1038/ng.3812 (2017).
Wynn, R.F. et al. Intractable diarrhoea of infancy caused by neutrophil specific granule deficiency and cured by stem cell transplantation. Gut 55, 292–293 (2006).
Kotlarz, D. et al. Loss-of-function mutations in the IL-21 receptor gene cause a primary immunodeficiency syndrome. J. Exp. Med. 210, 433–443 (2013).
Glocker, E.O. et al. Inflammatory bowel disease and mutations affecting the interleukin-10 receptor. N. Engl. J. Med. 361, 2033–2045 (2009).
Bohn, G. et al. A novel human primary immunodeficiency syndrome caused by deficiency of the endosomal adaptor protein p14. Nat. Med. 13, 38–45 (2007).
Hamada, T. et al. Lipoid proteinosis maps to 1q21 and is caused by mutations in the extracellular matrix protein 1 gene (ECM1). Hum. Mol. Genet. 11, 833–840 (2002).
Krawitz, P. et al. Microindel detection in short-read sequence data. Bioinformatics 26, 722–729 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Ellett, F., Pase, L., Hayman, J.W., Andrianopoulos, A. & Lieschke, G.J. mpeg1 promoter transgenes direct macrophage-lineage expression in zebrafish. Blood 117, e49–e56 (2011).
Lin, H.F. et al. Analysis of thrombocyte development in CD41–GFP transgenic zebrafish. Blood 106, 3803–3810 (2005).
Gagnon, J.A. et al. Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS One 9, e98186 (2014).
Meeker, N.D., Hutchinson, S.A., Ho, L. & Trede, N.S. Method for isolation of PCR-ready genomic DNA from zebrafish tissues. Biotechniques 43, 610, 612, 614 (2007).
Zhu, L.J., Holmes, B.R., Aronin, N. & Brodsky, M.H. CRISPRseek: a Bioconductor package to identify target-specific guide RNAs for CRISPR–Cas9 genome-editing systems. PLoS One 9, e108424 (2014).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Liu, Y., Zhou, J. & White, K.P. RNA–seq differential expression studies: more sequence or more replication? Bioinformatics 30, 301–304 (2014).
Sedlazeck, F.J., Rescheneder, P. & von Haeseler, A. NextGenMap: fast and accurate read mapping in highly polymorphic genomes. Bioinformatics 29, 2790–2791 (2013).
Liao, Y., Smyth, G.K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Rau, A., Gallopin, M., Celeux, G. & Jaffrezic, F. Data-based filtering for replicated high-throughput transcriptome sequencing experiments. Bioinformatics 29, 2146–2152 (2013).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA–seq data with DESeq2. Genome Biol. 15, 550 (2014).
Adli, M. & Bernstein, B.E. Whole-genome chromatin profiling from limited numbers of cells using nano-ChIP–seq. Nat. Protoc. 6, 1656–1668 (2011).
Cernilogar, F.M. et al. Chromatin-associated RNA interference components contribute to transcriptional regulation in Drosophila. Nature 480, 391–395 (2011).
Dahl, J.A. & Collas, P. A rapid micro chromatin immunoprecipitation assay (microChIP). Nat. Protoc. 3, 1032–1045 (2008).
Rahl, P.B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010).
Buenrostro, J.D., Wu, B., Chang, H.Y. & Greenleaf, W.J. ATAC–seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Croft, D. et al. The Reactome pathway knowledgebase. Nucleic Acids Res. 42, D472–D477 (2014).
Milacic, M. et al. Annotating cancer variants and anti-cancer therapeutics in Reactome. Cancers (Basel) 4, 1180–1211 (2012).
Newman, M.E. Modularity and community structure in networks. Proc. Natl. Acad. Sci. USA 103, 8577–8582 (2006).
Acknowledgements
We thank all medical and laboratory staff members involved in taking care of patients and performing scientific experiments, in particular R. Conca (FACS sorting), J. Hinke (genomic facility), and P. Robinson and S. Mundlos for next-generation sequencing expertise. We thank S. Hollizeck, D. Kotlarz, M. Lyszkiewicz, and N. Zietara for critical scientific discussion. We thank B. Zeller, R. Abdennour, and H. Nordgarden for clinical care of patients and A. Tierens for initial FACS and histological workup. We thank J. Lessard (IRIC, Université de Montréal) for providing antibodies to SMARCD1, SMARCD2, and SMARCD3 and for critical discussion.
The study has been supported by the European Research Council (ERC Advanced Grant 'Explore'), the Else Kröner-Fresenius-Stiftung, the DZIF (German Center for Infection Research), the Deutsche Forschungsgemeinschaft (Gottfried Wilhelm Leibniz Program), the German PID-NET (BMBF), and the Care-for-Rare Foundation.
V.P. was supported by a Monash International Postgraduate Research Scholarship (MIPRS) and a Monash Graduate Scholarship (MGS). G.L. was supported by the NHMRC (1069284, 1044754). The Australian Regenerative Medicine Institute (ARMI) is supported by grants from the State Government of Victoria and the Australian Government. This research was supported in part by the Intramural Research Program of the US National Institutes of Health, NLM. W.E. and C.Z. were supported by the Deutsche Forschungsgemeinschaft (DFG) through LMUexcellent and SFB1243 (subproject A14). J.G. was supported by the Bundesministerium für Bildung und Forschung, Juniorverbund in der Sytemmedizin 'mitOmics' grant FKZ 01ZX1405A, and C.M. is supported by EU Horizon2020 Collaborative Research Project SOUND (633974).
Author information
Authors and Affiliations
Contributions
M.W. designed, performed, and interpreted experiments and wrote and edited the manuscript. D.P. performed ATAC–seq and RNA–seq, Y.F., E.B., T.R., and M.R. were involved in genomic and biochemical analyses, J.P. led the computational biology efforts, C.M. and J.G. analyzed ATAC–seq and RNA–seq data, and C.Z. and W.E. performed mouse RNA–seq and digital gene expression analysis. A.S.-P., P.D.A., and M.R.A. provided clinical care for patients, V.P. and G.J.L. generated and analyzed zebrafish models, and P.M.K. analyzed whole-exome sequencing in initial patients. M.D., M.R.S., and E.W. generated mice. H.-P.H. performed immunohistochemistry analysis of bone marrow biopsies, H.S. provided expert clinical genetic consulting, and A.A.S. guided bioinformatics studies and helped write and edit the manuscript. C.K. designed and guided the study, supervised M.W., provided laboratory resources, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Syndromic features of human SMARCD2 deficiency.
(a–d) Phenotype of patient AII.1. (a,b) Low-set, posteriorly rotated, simple ears, with prominent concha and broad incisura intertragica. (c) Hallux valgus (filled arrowhead) and plantar hyperkeratosis of left foot (empty arrowhead). (d) Sandal gap/increased interdigital space D1–D2 (asterisk) and dorsal feet hyperkeratosis D1 (filled arrowhead). (e–l) Phenotype of patient BII.1. (e,f) Thin hair, ear with dysplastic helix, neck with buffalo hump, and acanthosis nigricans. (g) Misaligned, dysplastic teeth, incomplete amelogenesis imperfecta (filled arrowhead), and accessory teeth in gingival plicae (empty arrowheads). (h,i) Features of metabolic syndrome and pubertas praecox vera, (j) Brachytelephalangy, longitudinal ridges on finger nails, and keratosis pilaris on forearm. (k) Bulging of right humerus diaphysis (empty arrowhead) and osteochondroma (filled arrowhead). (l) Increased interdigital space D1–D2 (asterisk), dorsal feet hyperkeratosis D1, D2 left (filled arrowhead), brittle (5th) nail (empty arrowhead), and pes planus. Images have been partially cropped.
Supplementary Figure 2 Development of MDS/AML in SMARCD2 deficiency.
(a–f) HE staining of bone marrow biopsies. (a,d) In 04/2008, HE-stained sections show altered bone marrow microarchitecture with reversion of the myeloid/erythroid ratio and dysplastic megakaryocytes. (b,e) In 10/2008, HE-stained sections show a dysplastic bone marrow with marked fibrosis (filled arrowheads). (c,f) In 04/2011, HE-stained sections show a hypercellular bone marrow with focal and peritrabecular infiltration by blast cells (asterisks). (g–i) Glycophorin C staining of bone marrow sections. (g) In 04/2008, irregular erythroid islands were observed. (h) In 10/2008, abnormal erythroid islands located along fibrotic strands were noted. (i) In 04/2011, erythroid islands are present in the immediate vicinity of blast cells (asterisk). (j–l) CD61 staining of bone marrow sections. (j) In 04/2008, small dysplastic megakaryocytes with hypolobated nuclei were observed. (k) In 10/2008, dysplastic micromegakaryocytes were noted. (l) In 04/2011, hypolobated dysplastic megakaryocytes in differing intensities, reminiscent of inadequate megakaryocyte maturation and megakaryocyte dysplasia, were detected. Images have been partially cropped. Scale bars, approximately 20 μm.
Supplementary Figure 3 Decreased expression of granulocyte-specific proteins in SMARCD2-deficient hematopoietic cells.
(a) Immunohistochemical detection of lactoferrin (LTF) in healthy donor (HD) bone marrow cells showing strong expression of LTF in myeloid cells. (b) Immunohistochemical detection of CD15 on healthy donor bone marrow, showing strong expression of CD15 by mature neutrophilic cells. (c) Immunohistochemical detection of myeloperoxidase showing normal amounts and distribution of myeloperoxidase (MPO)-expressing neutrophilic cells presenting at all stages of maturation. (d) Immunohistochemical detection of LTF in patient AII.1 showing lack of LTF-expressing cells. (e) Immunohistochemical detection of CD15 in patient AII.1 showing reduced intensity and decreased cell numbers of CD15-expressing cells. (f) Immunohistochemical detection of MPO in patient AII.1 bone marrow cells showing a hypercellular bone marrow with presence of neutrophilic cells and blast cells expressing MPO with considerably lower intensity as compared to HD. (g) Immunohistochemical detection of LTF in patient BII.2 shows false positive background staining in a postmortem sample (note that LTF deficiency in cells from BII.1 has previously been documented by ELISA). (h) Immunohistochemical detection of CD15 on patient BII.2 bone marrow cells shows a hypercellular bone marrow with almost complete absence of CD15-expressing cells. (i) Immunohistochemical detection of MPO in patient BII.2 bone marrow cells shows hypercellularity. MPO-positive neutrophilic cells are reduced in numbers and intensity in comparison to HD. (j) Immunohistochemical detection of LTF in patient CII.1 showing a hypercellular bone marrow and a lack of LTF-expressing cells. (k) Immunohistochemical detection of CD15 in patient CII.1 showing a hypercellular bone marrow with markedly reduced intensity and decreased cell numbers of CD15-expressing cells in comparison to HD. (l) Immunohistochemical detection of MPO in patient CII.1 bone marrow cells shows a hypercellular bone marrow with clusters of neutrophilic cells and blast cells. MPO expression is reduced. Images have been partially cropped. Scale bars, approximately 20 μm.
Supplementary Figure 4 Zebrafish models for Smarcd2 deficiency.
(a) Knockdown of smarcd2 by the splice-blocking morpholino oligonucleotides was examined by RT–PCR. Morpholino SB2 (MO SB2) failed to disrupt smarcd2 splicing. Top, amplification of smarcd2 exon boundaries; bottom, β-actin control; (+) and (–) indicate samples prepared with and without reverse transcriptase, i.e., monitoring for genomic DNA contamination. Image cropped from one gel. The full-length gel image is shown in Supplementary Data 3b. (b) Fluorescence image of Danio rerio strain Tg(mpx:EGFP)i114, control versus smarcd2-morphant embryo. Reduced numbers of GFP-expressing neutrophils are observed in morphant versus control fish embryos. Acquired images: CTRL (n = 15 images), ATG MO (n = 15 images), SB1 MO (n = 15 images) and SB2 MO (n = 15 images). (c) Comprehensive analysis of neutrophil numbers in Tg(mpx:EGFP)i114 at 72 h.p.f. after injection of morpholino oligonucleotides (nonspecific control versus translation-start-site blocker (ATG) and splice-site blocker (SB1 and SB2) MOs against smarcd2. Numbers represent fluorescence-labeled neutrophils per individual fish embryos. Data were pooled from two independent morpholino experiments: CTRL n = 15, ATG n = 15, SB1 n = 15, SB2 n = 15 fish. Center value, mean; error bars, s.d. P values were calculated by two-tailed unpaired t test. Replicates: 2. Please compare to Supplementary Tables 19 and 20. These results are an independent verification of the data in Figure 3b in another neutrophil reporter line. (d) Sanger chromatogram of the CRISPR/Cas9-edited frameshift allele smarcd21/1 (1/1) versus the smarcd2 wild-type allele (wt) indicating the induced mutation c.66dup (Mut).
Supplementary Figure 5 Smarcd2 knockdown in zebrafish is specific for granulopoiesis.
(a) Erythrocyte staining in zebrafish larvae using O-dianisidine staining. Zebrafish treated with smarcd2 SB morpholino (right) show staining equal to that of control-treated zebrafish (left). Gross comparison shows equal O-dianisidine staining in control and morphant fish. (b) Macrophage numbers in zebrafish treated with smarcd2 SB morpholino and control-treated zebrafish are visualized by mCherry-positive cells in the tail. The smarcd2 SB morpholino, inducing aberrant splicing and consequent smarcd2 deficiency, shows no effect on macrophage numbers. Center value, mean; error bars, s.e.m. P values were calculated by two-tailed unpaired t test. Acquired images: CTRL (n = 94 images) and SB1 MO (n = 48 images). (c) Thrombocyte numbers in zebrafish treated with smarcd2 SB morpholino and control-treated zebrafish are visualized by EGFP-positive cells in the tail. The smarcd2 SB morpholino, inducing aberrant splicing and consequent smarcd2 deficiency, shows no effect on thrombocyte numbers. Center value, mean; error bars, s.e.m. P values were calculated by two-tailed unpaired t test. Acquired images: CTRL (n = 48 images) and SB1 MO (n = 63 images). Data are pooled from two independent experiments in b and c. Replicates: 2, See Supplementary Table 20 for raw data.
Supplementary Figure 6 Morphology of smarcd2-deficient neutrophils in zebrafish.
(a) FACS sorting strategy of zebrafish neutrophils. (b) Developmental stages of zebrafish granulopoiesis (May–Grünwald Giemsa staining). (c) May–Grünwald Giemsa staining of sorted neutrophils in control versus morpholino knockdown zebrafish. (d) No gross histological differences were recognized nor was any overt difference revealed by a four-category differential count of the embryonic neutrophil population (n = 1 experiment, pooling of neutrophils from n = 150 embryos/group); see Supplementary Table 20 for raw data.
Supplementary Figure 7 Generation of Smarcd2-knockout mice.
(a) Mouse Smarcd2 (ENSMUST00000021052)-mutant embryonic stem cells (ES cells with heterozygous deletion of Smarcd2; clone AF4) were purchased from KOMP repository. Shown is Smarcd2tm1(KOMP)Vlcg (deletion allele). The positions of the primer pairs for the wild-type allele (empty arrowheads) and the knockout allele (filled arrowheads) are shown in relation to the Smarcd2 gene and deletion allele, respectively. (b) PCR-based genotyping. F1 intercrosses of heterozygous mice resulted in Smarcd2+/+, Smarcd2+/−, and Smarcd2−/− embryos. PCR was performed according to the manufacturer (KOMP repository): wild-type allele, 264 bp (empty arrowheads) and knockout allele, 569 bp (filled arrowheads). Embryos 13, 15, and 17 are of the knockout genotype (Smarcd2−/−), and embryos 12, 14, and 16 are of the heterozygous genotype (Smarcd2+/−). Images have been cropped; full-length images are shown in Supplementary Data 3d,e. (c–e) Mendelian inheritance pattern of mutant Smarcd2 alleles. (c) Knockout embryos are not viable after birth. The numbers of Smarcd2+/+ (n = 24), Smarcd2+/− (n = 52), and Smarcd2−/− (n = 1, perinatal death) offspring born are displayed. (d) The distributions of genotypes per litter (in percent) are displayed. In total, expected Mendelian ratios of the genotypes (percent of genotype per litter) are observed in utero. Data from 15 litters are shown; compare with Supplementary Table 20. Center value, mean; error bars, s.e.m. P values were calculated by two-tailed unpaired t test. (e) The number of embryos per genotype and gestational age are shown at 12.5, 13.5, 14.5, 15.5, and 16.5 d.p.c. In total, wild type n = 24, heterozygous n = 56, and knockout n = 27 embryos were observed. (f) Western blot analysis detecting members of the SWI/SNF complex in Smarcd2-knockout embryos. Protein expression of SMARCD1 (BAF60a), SMARCD2 (BAF60b), SMARCD2 (BAF60c) (kindly provided by J. Lessard, Montreal), and SWI/SNF members BRG1, BAF170, BAF155, and BAF47 are shown. Images of membranes have been cropped; full images are available in Supplementary Data 4. Protein expression was determined in crude embryonic tissue lysates by western blotting. Embryos from two litters were analyzed: wild type n = 3, heterozygous n = 3, knockout n = 3. Replicates: 1. (g) Enumeration of total white blood cells (CD45.2+) and the stem cell compartment (LSK cells and LSK CD150+CD48− cells) per embryo in wild-type, heterozygous, and knockout mouse fetal liver hematopoiesis. Smarcd2−/− embryos show reduced CD45.2+ blood cells but comparable amounts of stem cells (LSK, LSK CD150+CD48−). Data are from three litters: wild type n = 7; heterozygous n = 10; knockout n = 8. Center value, mean; error bars, s.e.m. P values were calculated by two-tailed unpaired t test. Replicates: 1.
Supplementary Figure 8 Colony count and flow cytometric analysis of GM colonies.
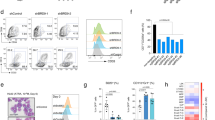
(a) Comprehensive analysis of myeloid cell colonies. Data are from five experiments with eight litters: wild type n = 10, heterozygous n = 13, knockout n = 12. Numbers of colonies have been normalized to the input of 500 LSK cells. Center value, mean; error bars, s.e.m. P values were calculated by two-tailed unpaired t test. The F test when performed to compare the variances of the wild-type, heterozygous, and knockout groups showed, that variances differed significantly in total colonies (wild type vs. heterozygous, ***P < 0.001; wild type vs. knockout, ***P < 0.001), GM colonies (wild type vs. heterozygous, ***P < 0.001; wild type vs. knockout, **P = 0.003), M colonies (wild type vs. knockout, *P = 0.011), and G colonies (wild type vs. heterozygous, *P = 0.036; wild type vs. knockout, ***P < 0.001). Unpaired t tests with Welch’s correction showed that differences in means are significant in total colonies (wild type vs. heterozygous, *P = 0.037; wild type vs. knockout, **P = 0.004), in GM colonies (wild type vs. heterozygous, *P = 0.019; wild type vs. knockout, *P = 0.013), in M colonies (wild type vs. knockout, *P = 0.022), and in G colonies (wild type vs. knockout, *P = 0.011). This is likely to be due to sampling of experiments, outliers, and low numbers of colonies in M/G knockout samples. (b) Representative FACS plots of myeloid GM colonies derived from a showing expression of myeloid surface markers. Numbers are representative of the percentage of CD11b+Gr1+ and CD11b+Ly6c+ cells. CD11b+Gr1+ and CD11b+Ly6c+ cells are absent in Smarcd2−/− colonies. Data shown are from one experiment (wild type n = 1, heterozygous n = 2, knockout n = 2), subsequently confirmed by a second experiment with wild type n = 5, heterozygous n = 4, and knockout n = 6, for which data are not shown. Replicates: 2.
Supplementary Figure 9 Differentially expressed genes in mouse Smarcd2-deficient early progenitors affect myeloid pathways.
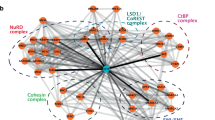
(a) Differentially expressed genes in Smarcd2+/+ versus Smarcd2−/− mouse LSK cells cluster in 11 groups by Reactome FI spectral clustering. Pathways cluster significantly in immune system and myeloid pathways. Smarcd2 is highlighted in red. Relevant pathways and interactions between pathways are indicated. The sources of pathway annotations are indicated in parentheses: C, CellMap; R, Reactome; K, KEGG; N, NCI PID; P, Panther; B, BioCarta. A complete list of pathways enriched per cluster is shown in Supplementary Table 6a. (b) Venn diagram of differentially expressed genes in Smarcd2+/+ versus Smarcd2−/− LSK cells and CEBPɛ target genes (Supplementary Table 6c). (c) Differentially expressed genes in Smarcd2+/+ versus Smarcd2−/− mouse LSK cells overlap with CEBPɛ targets (13 genes) and interact via MYC, FOS, EP300, and the ubiquitin system. Genes in green diamonds have been introduced as linker genes; differentially expressed genes are color-coded by continuous mapping of log(fold change) generated in LSK Smarcd2+/+ versus Smarcd2−/− RNA–seq experiments (blue, reduced expression in knockout; red, increased expression in knockout). Analysis and display of the network were carried out using Cytoscape 3.3.0 and the Reactome Functional Interaction (FI) plugin (solid line with arrow for activating/catalyzing FI, solid line with block for inhibitory FI, solid line (alone) for FIs extracted from complexes or inputs, and dotted lines for predicted FIs). The genes for all intersections are listed in Supplementary Table 6c.
Supplementary Figure 10 Absence of GMP progenitors in Smarcd2−/− fetal liver hematopoiesis.
(a–c) Representative example of wild-type (a) and knockout (b) fetal liver hematopoiesis (per embryo) and gating strategy (c). Cells were first discriminated by FSC/SSC (data not shown) and then by SSC-H/CD45. CD45+ cells were gated on Lin−c-Kit+ cells. CD45+Lin− Sca-1+c-Kit+ (LSK) cells were sorted, and CD45+Lin−CD117+Sca-1− (myeloid progenitor) cells were further separated into CD45+Lin−Sca-1−c-Kit+CD34−CD16/32lo MEPs, CD45+Lin−Sca-1−c-Kit+CD34+CD16/32int CMPs, and CD45+Lin−Sca-1−c-Kit+CD34+CD16/32hi GMPs. In comparison to a, GMP cells are almost absent in b. (c) Numbers in black boxes refer to the percentage of wild-type cells, and numbers in white boxes refer to the percentage of knockout cells.
Supplementary Figure 11 Promoter occupancy of target genes for the CEBPɛ and BRG1 proteins in SMARCD2-deficient NB4 cells.
(a–c) CEBPɛ and BRG1 bind to the promoter regions of MMP8 (a), CAMP (b), and SERPIN A1 (c). Although binding is not significantly reduced in cells transduced with shRNA1 (P = ns) or shRNA2 (P = ns) in comparison to cells transduced with a nonspecific control shRNA (CTRL), a trend can be observed. The experiment was performed two times. Data points from two experiments with a total of five biological replicates (five independent cultures and ChIP immunoprecipitations) are merged. Center value, mean; error bars, s.d. P values were calculated by two-tailed unpaired t test. Replicates: 3.
Supplementary Figure 12 Immunoprecipitation of endogenous CEBPɛ.
(a–d) Experiment 1. (a) Immunoblotting for SMARCD2 detects SMARCD2 in a CEBPɛ immunoprecipitate in short and long exposures; a distinct band is noted (red arrow) at the expected size of ~56 kDa, corresponding to input and coimmunoprecipitated SMARCD2 proteins. (b) Cropped lanes of a showing immunoprecipitation with isotype control and antibody to CEBPɛ (a2). (c) CEBPɛ is detected in input and immunoprecipitate; the image was cropped (c3) for summary. (d) Cropped images highlighting bands from experiment 1 shown with standard kDa marker (PageRuler Prestained Protein Ladder, Thermo Fisher). (e–g) Experiment 2. (e) Immunoblotting for SMARCD2 detects SMARCD2 in two independent CEBPɛ immunoprecipitates from the same lysate (technical replicates). Distinct bands are noted (red arrows) at the expected size of ~56 kDa, corresponding to input and coimmunoprecipitated SMARCD2 proteins (e1). Immunoblotting for GAPDH shows the presence of GAPDH in input but not in immunoprecipitates (e3). (f) CEBPɛ is detected in input and immunoprecipitates (red arrows) (f2). (g) Cropped images highlighting bands from experiment 2 shown with standard kDa marker (PageRuler Prestained Protein Ladder, Thermo Fisher). (h–j) Experiment 3. (h) Immunoblotting for SMARCD2 detects SMARCD2 in the CEBPɛ immunoprecipitate. A distinct band is noted (red arrow) at the expected size of ~56 kDa, corresponding to input and coimmunoprecipitated SMARCD2 (h1). Immunoblotting for GAPDH shows the presence of GAPDH in Input but not in immunoprecipitates (red arrow) (h3). (i) CEBPɛ is detected in input and immunoprecipitates (red arrows) (i2). (j) Cropped images highlighting bands from experiment 3 shown with standard kDa marker (Precision Plus Protein Kaleidoscope Prestained Protein Standards, Bio-Rad). The protein standard ladders differ between experiment 1 and 2 and experiment 3.
Supplementary Figure 13 SMARCD2 knockdown in human NB4 AML cells compromises myeloid pathways.
(a) Differentially expressed genes in undifferentiated AML-NB4 cells (shRNA-mediated SMARCD2 knockdown versus control) cluster in 11 groups by Reactome FI spectral partition and show significant enrichment for myeloid pathways. SMARCD2 is highlighted in red. Relevant pathways and interactions between pathways are indicated. The sources of pathway annotations are indicated in parentheses: C, CellMap; R, Reactome; K, KEGG; N, NCI PID; P, Panther; B, BioCarta. A complete list of the pathways enriched per cluster is given in Supplementary Table 6e. (b) Differentially expressed genes in ATRA-differentiated AML-NB4 cells (shRNA-mediated SMARCD2 knockdown versus control) cluster in 12 groups by Reactome FI spectral partition and show significant enrichment for myeloid pathways. SMARCD2 is highlighted in red. Relevant pathways and interactions between pathways are indicated. The sources of pathway annotations are indicated in parentheses: C, CellMap; R, Reactome; K, KEGG; N, NCI PID; P, Panther; B, BioCarta. A complete list of the pathways enriched per cluster is shown in Supplementary Table 6f. (c) Venn diagram of differentially expressed genes (undifferentiated and ATRA differentiated) and CEBPɛ target genes. Genes from all intersections are listed in Supplementary Table 6d. (d) Differentially expressed genes in ATRA-differentiated AML-NB4 cells overlap with CEBPɛ targets (21 genes) and interact via EP300, TP53, STAT1, and the ubiquitin system. Genes in green diamonds have been introduced as linker genes; differentially expressed genes are color-coded by continuous mapping of log(fold change) in SMARCD2-knockdown cells versus control cells (blue, reduced expression in SMARCD2-knockdown cells; red, increased expression in SMARCD2-knockdown cells). Analysis and display of the network were carried out with Cytoscape 3.3.0 and the Reactome Functional Interaction (FI) plugin: solid lines with arrows for activating/catalyzing FIs, solid lines with blocks for inhibitory FIs, solid lines (alone) for FIs extracted from complexes or inputs, and dotted lines for predicted FIs. This data set is derived from NB4 cells exposed to 1 μM ATRA for 2 d and, thus, cannot directly be compared to the qPCR data shown in Figure 5a.
Supplementary Figure 14 Regions of altered chromatin accessibility cluster around transcription start sites.
(a) Regions of open chromatin cluster around transcription start sites (TSSs) in DMSO/mock-treated NB4 cells. Peaks were binned into 500-bp windows with respect to their distance to the TSS. Each bar represents the fraction of peaks falling into one bin against all peaks of one group. The peaks are grouped as all peaks (black), upregulated peaks (red), and downregulated peaks (blue). The depicted peak distribution is based on undifferentiated (DMSO vehicle control) NB4 cells. (b) As in a but based on ATRA-differentiated NB4 cells.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14, Supplementary Note and Supplementary Tables 1, 2 and 18 (PDF 3733 kb)
Supplementary Data 1–5
Supplementary Data 1–5 (PDF 13450 kb)
Supplementary Table 3
Primers for human, mouse, and zebrafish studies; morpholinos and sgRNAs for zebrafish studies. (XLSX 16 kb)
Supplementary Table 4
Differentially expressed genes in Smarcd2+/+ vs. Smarcd2−/− LSK cells. (XLSX 517 kb)
Supplementary Table 5
Up- and downregulated clusters of genes in Smarcd2+/+ vs. Smarcd2−/− LSK cells. (XLSX 34 kb)
Supplementary Table 6
CEBPɛ target genes, Venn intersections, and pathways. (XLSX 53 kb)
Supplementary Table 7
Differentially expressed genes in Smarcd2+/+ vs. Smarcd2−/− LSK cells. (XLSX 15 kb)
Supplementary Table 8
Differentially expressed genes in Smarcd2+/+ vs. Smarcd2−/− CMP cells. (XLSX 159 kb)
Supplementary Table 9
Differentially expressed genes in Smarcd2+/+ vs. Smarcd2−/− GMP cells. (XLSX 77 kb)
Supplementary Table 10
Differentially expressed genes in Smarcd2+/+ vs. Smarcd2−/− MEP cells. (XLSX 38 kb)
Supplementary Table 11
Up- and downregulated GO terms in Smarcd2+/+ vs. Smarcd2−/− CMP cells. (XLSX 21 kb)
Supplementary Table 12
Up- and downregulated GO terms in Smarcd2+/+ vs. Smarcd2−/− GMP cells (XLSX 19 kb)
Supplementary Table 13
Up- and downregulated GO terms in Smarcd2+/+ vs. Smarcd2−/− LSK cells. (XLSX 10 kb)
Supplementary Table 14
Up- and downregulated GO terms in Smarcd2+/+ vs. Smarcd2−/− MEP cells. (XLSX 20 kb)
Supplementary Table 15
Differentially expressed genes in ATAC–seq and RNA–seq, CTRL vs. shRNA1-, shRNA2-treated NB4 cells, vehicle DMSO treated. (XLSX 7374 kb)
Supplementary Table 16
Differentially expressed genes in ATAC–seq and RNA–seq, CTRL vs. shRNA1-, shRNA2-treated NB4 cells, ATRA treated. (XLSX 2427 kb)
Supplementary Table 17
Available blood counts of patients A, B, and C, chronologically arranged. (XLSX 31 kb)
Supplementary Table 19
Definition and number of replicates performed per experiment. (XLSX 31 kb)
Supplementary Table 20
Numbers used for dot-bot diagrams and statistical analysis, statistical test results. (XLSX 464 kb)
Rights and permissions
About this article
Cite this article
Witzel, M., Petersheim, D., Fan, Y. et al. Chromatin-remodeling factor SMARCD2 regulates transcriptional networks controlling differentiation of neutrophil granulocytes. Nat Genet 49, 742–752 (2017). https://doi.org/10.1038/ng.3833
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3833
This article is cited by
-
SWI/SNF complexes in hematological malignancies: biological implications and therapeutic opportunities
Molecular Cancer (2023)
-
Zebrafish: a convenient tool for myelopoiesis research
Cell Regeneration (2023)
-
Multi-omics and machine learning reveal context-specific gene regulatory activities of PML::RARA in acute promyelocytic leukemia
Nature Communications (2023)
-
BRD9 determines the cell fate of hematopoietic stem cells by regulating chromatin state
Nature Communications (2023)
-
Novel Genetic Discoveries in Primary Immunodeficiency Disorders
Clinical Reviews in Allergy & Immunology (2022)