Abstract
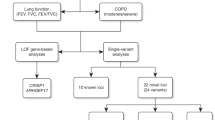
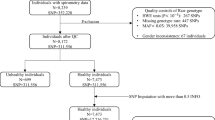
Chronic obstructive pulmonary disease (COPD) is characterized by reduced lung function and is the third leading cause of death globally. Through genome-wide association discovery in 48,943 individuals, selected from extremes of the lung function distribution in UK Biobank, and follow-up in 95,375 individuals, we increased the yield of independent signals for lung function from 54 to 97. A genetic risk score was associated with COPD susceptibility (odds ratio per 1 s.d. of the risk score (∼6 alleles) (95% confidence interval) = 1.24 (1.20–1.27), P = 5.05 × 10−49), and we observed a 3.7-fold difference in COPD risk between individuals in the highest and lowest genetic risk score deciles in UK Biobank. The 97 signals show enrichment in genes for development, elastic fibers and epigenetic regulation pathways. We highlight targets for drugs and compounds in development for COPD and asthma (genes in the inositol phosphate metabolism pathway and CHRM3) and describe targets for potential drug repositioning from other clinical indications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Fletcher, C. & Peto, R. The natural history of chronic airflow obstruction. BMJ 1, 1645–1648 (1977).
Lange, P., Celli, B. & Agustí, A. Lung-function trajectories and chronic obstructive pulmonary disease. N. Engl. J. Med. 373, 1575 (2015).
Lozano, R. et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380, 2095–2128 (2012).
Salvi, S.S. & Barnes, P.J. Chronic obstructive pulmonary disease in non-smokers. Lancet 374, 733–743 (2009).
Melén, E. Bridging genetics, epidemiology, and respiratory medicine. Am. J. Respir. Crit. Care Med. 190, 716–718 (2014).
Svanes, C. et al. Early life origins of chronic obstructive pulmonary disease. Thorax 65, 14–20 (2010).
Celli, B.R. et al. An official American Thoracic Society/European Respiratory Society statement: research questions in COPD. Eur. Respir. J. 45, 879–905 (2015).
Pillai, S.G. et al. A genome-wide association study in chronic obstructive pulmonary disease (COPD): identification of two major susceptibility loci. PLoS Genet. 5, e1000421 (2009).
Cho, M.H. et al. Variants in FAM13A are associated with chronic obstructive pulmonary disease. Nat. Genet. 42, 200–202 (2010).
Cho, M.H. et al. Risk loci for chronic obstructive pulmonary disease: a genome-wide association study and meta-analysis. Lancet Respir. Med. 2, 214–225 (2014).
Hancock, D.B. et al. Meta-analyses of genome-wide association studies identify multiple loci associated with pulmonary function. Nat. Genet. 42, 45–52 (2010).
Loth, D.W. et al. Genome-wide association analysis identifies six new loci associated with forced vital capacity. Nat. Genet. 46, 669–677 (2014).
Repapi, E. et al. Genome-wide association study identifies five loci associated with lung function. Nat. Genet. 42, 36–44 (2010).
Soler Artigas, M. et al. Genome-wide association and large-scale follow up identifies 16 new loci influencing lung function. Nat. Genet. 43, 1082–1090 (2011).
Soler Artigas, M. et al. Sixteen new lung function signals identified through 1000 Genomes Project reference panel imputation. Nat. Commun. 6, 8658 (2015).
Wain, L.V. et al. Novel insights into the genetics of smoking behaviour, lung function, and chronic obstructive pulmonary disease (UK BiLEVE): a genetic association study in UK Biobank. Lancet Respir. Med. 3, 769–781 (2015).
Nelson, M.R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Sanseau, P. et al. Use of genome-wide association studies for drug repositioning. Nat. Biotechnol. 30, 317–320 (2012).
Palmer, L.J. et al. Familial aggregation and heritability of adult lung function: results from the Busselton Health Study. Eur. Respir. J. 17, 696–702 (2001).
Wilk, J.B. et al. Evidence for major genes influencing pulmonary function in the NHLBI family heart study. Genet. Epidemiol. 19, 81–94 (2000).
Weiss, S.T. Lung function and airway diseases. Nat. Genet. 42, 14–16 (2010).
Gudbjartsson, D.F. et al. Large-scale whole-genome sequencing of the Icelandic population. Nat. Genet. 47, 435–444 (2015).
Hao, K. et al. Lung eQTLs to help reveal the molecular underpinnings of asthma. PLoS Genet. 8, e1003029 (2012).
Lamontagne, M. et al. Refining susceptibility loci of chronic obstructive pulmonary disease with lung eqtls. PLoS One 8, e70220 (2013).
Obeidat, M. et al. GSTCD and INTS12 regulation and expression in the human lung. PLoS One 8, e74630 (2013).
Regan, E.A. et al. Genetic epidemiology of COPD (COPDGene) study design. COPD 7, 32–43 (2010).
Vestbo, J. et al. Evaluation of COPD Longitudinally to Identify Predictive Surrogate End-points (ECLIPSE). Eur. Respir. J. 31, 869–873 (2008).
Fishman, A. et al. A randomized trial comparing lung-volume-reduction surgery with medical therapy for severe emphysema. N. Engl. J. Med. 348, 2059–2073 (2003).
Bell, B., Rose, C.L. & Damon, H. The Normative Aging Study: an interdisciplinary and longitudinal study of health and aging. Aging Hum. Dev. 3, 5–17 (1972).
Dewey, F.E. et al. Inactivating variants in ANGPTL4 and risk of coronary artery disease. N. Engl. J. Med. 374, 1123–1133 (2016).
Szklarczyk, D. et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Wilk, J.B. et al. Genome-wide association studies identify CHRNA5/3 and HTR4 in the development of airflow obstruction. Am. J. Respir. Crit. Care Med. 186, 622–632 (2012).
Jiang, Z. et al. A chronic obstructive pulmonary disease susceptibility gene, FAM13A, regulates protein stability of β-catenin. Am. J. Respir. Crit. Care Med. 194, 185–197 (2016).
Obeidat, M. et al. Molecular mechanisms underlying variations in lung function: a systems genetics analysis. Lancet Respir. Med. 3, 782–795 (2015).
Carter, R.I., Ungurs, M.J., Mumford, R.A. & Stockley, R.A. Aα-Val360: a marker of neutrophil elastase and COPD disease activity. Eur. Respir. J. 41, 31–38 (2013).
Sand, J.M. et al. Accelerated extracellular matrix turnover during exacerbations of COPD. Respir. Res. 16, 69 (2015).
Jhingran, A. et al. Compartment-specific and sequential role of MyD88 and CARD9 in chemokine induction and innate defense during respiratory fungal infection. PLoS Pathog. 11, e1004589 (2015).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Cogan, J.D. et al. Rare variants in RTEL1 are associated with familial interstitial pneumonia. Am. J. Respir. Crit. Care Med. 191, 646–655 (2015).
Todd, J.A., Bell, J.I. & McDevitt, H.O. HLA-DQβ gene contributes to susceptibility and resistance to insulin-dependent diabetes mellitus. Nature 329, 599–604 (1987).
Hobbs, B.D. et al. Genetic loci associated with chronic obstructive pulmonary disease overlap with loci for lung function and pulmonary fibrosis. Nat. Genet. http://dx.doi.org/10.0138/ng.3752 (2017).
Soler Artigas, M. et al. Effect of five genetic variants associated with lung function on the risk of chronic obstructive lung disease, and their joint effects on lung function. Am. J. Respir. Crit. Care Med. 184, 786–795 (2011).
Poole, C. A history of the population attributable fraction and related measures. Ann. Epidemiol. 25, 147–154 (2015).
Fadista, J., Manning, A.K., Florez, J.C. & Groop, L. The (in)famous GWAS P-value threshold revisited and updated for low-frequency variants. Eur. J. Hum. Genet. 24, 1202–1205 (2016).
Miller, M.R. et al. Standardisation of spirometry. Eur. Respir. J. 26, 319–338 (2005).
Abecasis, G.R. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Huang, J. et al. Improved imputation of low-frequency and rare variants using the UK10K haplotype reference panel. Nat. Commun. 6, 8111 (2015).
Walter, K. et al. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015).
Wilk, J.B. et al. A genome-wide association study of pulmonary function measures in the Framingham Heart Study. PLoS Genet. 5, e1000429 (2009).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Dastani, Z. et al. Novel loci for adiponectin levels and their influence on type 2 diabetes and metabolic traits: a multi-ethnic meta-analysis of 45,891 individuals. PLoS Genet. 8, e1002607 (2012).
Wakefield, J. A Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am. J. Hum. Genet. 81, 208–227 (2007).
van de Bunt, M., Cortes, A., Brown, M.A., Morris, A.P. & McCarthy, M.I. Evaluating the performance of fine-mapping strategies at common variant GWAS loci. PLoS Genet. 11, e1005535 (2015).
Jia, X. et al. Imputing amino acid polymorphisms in human leukocyte antigens. PLoS One 8, e64683 (2013).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP–trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Eicher, J.D. et al. GRASP v2.0: an update on the Genome-Wide Repository of Associations between SNPs and phenotypes. Nucleic Acids Res. 43, D799–D804 (2015).
Higgins, J.P., Thompson, S.G., Deeks, J.J. & Altman, D.G. Measuring inconsistency in meta-analyses. Br. Med. J. 327, 557–560 (2003).
Westra, H.J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Kamburov, A., Stelzl, U., Lehrach, H. & Herwig, R. The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res. 41, D793–D800 (2013).
Segrè, A.V., Groop, L., Mootha, V.K., Daly, M.J. & Altshuler, D. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Trynka, G. et al. Disentangling the Effects of Colocalizing Genomic Annotations to Functionally Prioritize Non-coding Variants within Complex-Trait Loci. Am. J. Hum. Genet. 97, 139–152 (2015).
Acknowledgements
This work was funded by a Medical Research Council (MRC) strategic award to M.D.T., I.P.H., D.S. and L.V.W. (MC_PC_12010). This research has been conducted using the UK Biobank Resource under application 648. This article presents independent research funded partially by the National Institute for Health Research (NIHR). The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the UK Department of Health. This research used the ALICE and SPECTRE High-Performance Computing Facilities at the University of Leicester. Additional acknowledgments and funding details can be found in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
L.V.W., D.J.P., M.-R.J., A.L.J., N.J.W., J.F.W., B.S., H.S., N.M.P.-H., S.K., C.G., I.J.D., I. Rudan, S.M.K., O.P., M.K., C.H., T.L., O.T.R., A.J.H., C.E.P., P.D.S., A.G., P.S.B., J.D.C., T.H.B., N.N.H., R.A.M., I. Ruczinski, K.C.B., Y.B., P.J., P.D.P., D.D.S., K.H., E.P.B., R.J.F.L., R.G.W., Z.C., I.Y.M., L.L., E.Z., I. Sayers, D.P.S., I.P.H., U.G. and M.D.T. contributed to the conception and study design. L.V.W., N.S., M.S., A.M.E., B.N., L.B.-C., M.O., A.P.H., M.A.P., R.J.H., C.K.B., T.L.R., A.G.F., C.J., T.B., V.E.J., R.J.A., B.P.P., A.C., M.W., J.H., J.Z., P.K.J., B.S., R.R., M.I., N.M.P.-H., S.E.H., J.M., S.E., I. Surakka, V.V., C.H., T.L., D.M.E., C.A.W., E.S.W., R.B., B.D.H., A.A.L., D.W.S., M.v.d.B., C.-A.B., D.C.N., O.G., F.E.D., S.E.B., D.J.C., H.L.K., S.J., G. Thorleifsson, I.J., T.G., K.S., C.S., G.N., R.G.W., J.V., O.P.K., M.H.C., E.K.S., G. Trynka, J.H.Z. and D.P.S. contributed to data analysis. L.V.W., N.S., M.S., A.M.E., B.N., M.O., A.P.H., M.A.P., R.J.H., C.K.B., T.L.R., A.G.F., C.J., V.E.J., A.C., M.-R.J., B.S., R.R., H.S., M.I., N.M.P.-H., S.K., C.G., C.H., A.G., C.S., G.N., R.J.F.L., A.L.H., C.B., I. Sayers, A.P.M., D.P.S., I.P.H. and M.D.T. contributed to data interpretation.
Corresponding authors
Ethics declarations
Competing interests
F.E.D. and S.E.B. are employed by Regeneron Pharmaceuticals. D.C.N. is employed by Merck. In the past three years, E.K.S. received honoraria and consulting fees from Merck, grant support and consulting fees from GlaxoSmithKline, and honoraria and travel support from Novartis. S.J., G. Thorleifsson, I.J. and K.S. are employed by deCODE Genetics/Amgen. M.H.C. receives grant funding from GlaxoSmithKline.
Additional information
A list of members and affiliations appears in the Supplementary Note.
A list of members and affiliations appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1, 2, 5–9, 11 and 13–21, and Supplementary Note (PDF 8709 kb)
Supplementary Tables 3, 4, 10 and 12
Supplementary Tables 3, 4, 10 and 12 (XLSX 95 kb)
Rights and permissions
About this article
Cite this article
Wain, L., Shrine, N., Artigas, M. et al. Genome-wide association analyses for lung function and chronic obstructive pulmonary disease identify new loci and potential druggable targets. Nat Genet 49, 416–425 (2017). https://doi.org/10.1038/ng.3787
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3787
This article is cited by
-
Lipid metabolism-related genes are involved in the occurrence of asthma and regulate the immune microenvironment
BMC Genomics (2024)
-
Human genetic associations of the airway microbiome in chronic obstructive pulmonary disease
Respiratory Research (2024)
-
Genetics of chronic respiratory disease
Nature Reviews Genetics (2024)
-
Prediction and stratification of longitudinal risk for chronic obstructive pulmonary disease across smoking behaviors
Nature Communications (2023)
-
A genome-wide association study identifies distinct variants associated with pulmonary function among European and African ancestries from the UK Biobank
Communications Biology (2023)