Abstract
During the progression of pancreatic ductal adenocarcinoma (PDAC), heterogeneous subclonal populations emerge that drive primary tumor growth, regional spread, distant metastasis, and patient death. However, the genetics of metastases largely reflects that of the primary tumor in untreated patients, and PDAC driver mutations are shared by all subclones. This raises the possibility that an epigenetic process might operate during metastasis. Here we report large-scale reprogramming of chromatin modifications during the natural evolution of distant metastasis. Changes were targeted to thousands of large chromatin domains across the genome that collectively specified malignant traits, including euchromatin and large organized chromatin histone H3 lysine 9 (H3K9)-modified (LOCK) heterochromatin. Remarkably, distant metastases co-evolved a dependence on the oxidative branch of the pentose phosphate pathway (oxPPP), and oxPPP inhibition selectively reversed reprogrammed chromatin, malignant gene expression programs, and tumorigenesis. These findings suggest a model whereby linked metabolic–epigenetic programs are selected for enhanced tumorigenic fitness during the evolution of distant metastasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Embuscado, E.E. et al. Immortalizing the complexity of cancer metastasis: genetic features of lethal metastatic pancreatic cancer obtained from rapid autopsy. Cancer Biol. Ther. 4, 548–554 (2005).
Iacobuzio-Donahue, C.A. et al. DPC4 gene status of the primary carcinoma correlates with patterns of failure in patients with pancreatic cancer. J. Clin. Oncol. 27, 1806–1813 (2009).
Rahib, L. et al. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 74, 2913–2921 (2014).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R.A. & Feinberg, A.P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat. Genet. 41, 246–250 (2009).
Hansen, K.D. et al. Increased methylation variation in epigenetic domains across cancer types. Nat. Genet. 43, 768–775 (2011).
Timp, W. et al. Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors. Genome Med. 6, 61 (2014).
Feinberg, A.P., Koldobskiy, M.A. & Göndör, A. Epigenetic modulators, modifiers and mediators in cancer aetiology and progression. Nat. Rev. Genet. 17, 284–299 (2016).
Jones, S. et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008).
Campbell, P.J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010).
Makohon-Moore, A.P. et al. Limited heterogeneity of known driver gene mutations among the metastases of individual pancreatic cancer patients. Nat. Genet. http://dx.doi.org/10.1038/ng.3764 (2017).
McDonald, O.G., Wu, H., Timp, W., Doi, A. & Feinberg, A.P. Genome-scale epigenetic reprogramming during epithelial-to-mesenchymal transition. Nat. Struct. Mol. Biol. 18, 867–874 (2011).
Chen, W.H. et al. Human pancreatic adenocarcinoma: in vitro and in vivo morphology of a new tumor line established from ascites. In Vitro 18, 24–34 (1982).
Metzgar, R.S. et al. Antigens of human pancreatic adenocarcinoma cells defined by murine monoclonal antibodies. Cancer Res. 42, 601–608 (1982).
Kyriazis, A.A., Kyriazis, A.P., Sternberg, C.N., Sloane, N.H. & Loveless, J.D. Morphological, biological, biochemical, and karyotypic characteristics of human pancreatic ductal adenocarcinoma Capan-2 in tissue culture and the nude mouse. Cancer Res. 46, 5810–5815 (1986).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Rice, J.C. et al. Histone methyltransferases direct different degrees of methylation to define distinct chromatin domains. Mol. Cell 12, 1591–1598 (2003).
Hall, I.M. et al. Establishment and maintenance of a heterochromatin domain. Science 297, 2232–2237 (2002).
Duesberg, P. et al. Origin of metastases: subspecies of cancers generated by intrinsic karyotypic variations. Cell Cycle 11, 1151–1166 (2012).
Landt, S.G. et al. ChIP–seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Wen, B. et al. Euchromatin islands in large heterochromatin domains are enriched for CTCF binding and differentially DNA-methylated regions. BMC Genomics 13, 566 (2012).
Hon, G.C. et al. Global DNA hypomethylation coupled to repressive chromatin domain formation and gene silencing in breast cancer. Genome Res. 22, 246–258 (2012).
Klco, J.M. et al. Functional heterogeneity of genetically defined subclones in acute myeloid leukemia. Cancer Cell 25, 379–392 (2014).
Wellen, K.E. et al. ATP–citrate lyase links cellular metabolism to histone acetylation. Science 324, 1076–1080 (2009).
Lee, J.V. et al. Akt-dependent metabolic reprogramming regulates tumor cell histone acetylation. Cell Metab. 20, 306–319 (2014).
Mentch, S.J. et al. Histone methylation dynamics and gene regulation occur through the sensing of one-carbon metabolism. Cell Metab. 22, 861–873 (2015).
Cluntun, A.A. et al. The rate of glycolysis quantitatively mediates specific histone acetylation sites. Cancer Metab. 3, 10 (2015).
Kaelin, W.G. Jr. & McKnight, S.L. Influence of metabolism on epigenetics and disease. Cell 153, 56–69 (2013).
Carey, B.W., Finley, L.W., Cross, J.R., Allis, C.D. & Thompson, C.B. Intracellular α-ketoglutarate maintains the pluripotency of embryonic stem cells. Nature 518, 413–416 (2015).
Vander Heiden, M.G., Cantley, L.C. & Thompson, C.B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Liberti, M.V. & Locasale, J.W. The Warburg effect: how does it benefit cancer cells? Trends Biochem. Sci. 41, 211–218 (2016).
Shan, C. et al. Lysine acetylation activates 6-phosphogluconate dehydrogenase to promote tumor growth. Mol. Cell 55, 552–565 (2014).
Johnson, W.J. & McColl, J.D. 6-Aminonicotinamide—a potent nicotinamide antagonist. Science 122, 834 (1955).
Köhler, E., Barrach, H. & Neubert, D. Inhibition of NADP dependent oxidoreductases by the 6-aminonicotinamide analogue of NADP. FEBS Lett. 6, 225–228 (1970).
Cheung, W.K. et al. Control of alveolar differentiation by the lineage transcription factors GATA6 and HOPX inhibits lung adenocarcinoma metastasis. Cancer Cell 23, 725–738 (2013).
Li, C. et al. c-Met is a marker of pancreatic cancer stem cells and therapeutic target. Gastroenterology 141, 2218–2227.e5 (2011).
Ridky, T.W., Chow, J.M., Wong, D.J. & Khavari, P.A. Invasive three-dimensional organotypic neoplasia from multiple normal human epithelia. Nat. Med. 16, 1450–1455 (2010).
Andl, C.D. et al. Epidermal growth factor receptor mediates increased cell proliferation, migration, and aggregation in esophageal keratinocytes in vitro and in vivo. J. Biol. Chem. 278, 1824–1830 (2003).
Liu, X., Ser, Z. & Locasale, J.W. Development and quantitative evaluation of a high-resolution metabolomics technology. Anal. Chem. 86, 2175–2184 (2014).
Hansen, K.D., Langmead, B. & Irizarry, R.A. BSmooth: from whole genome bisulfite sequencing reads to differentially methylated regions. Genome Biol. 13, R83 (2012).
Hu, J., Ge, H., Newman, M. & Liu, K. OSA: a fast and accurate alignment tool for RNA–Seq. Bioinformatics 28, 1933–1934 (2012).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP–Seq (MACS). Genome Biol. 9, R137 (2008).
Song, Q. & Smith, A.D. Identifying dispersed epigenomic domains from ChIP–Seq data. Bioinformatics 27, 870–871 (2011).
Hawkins, R.D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6, 479–491 (2010).
Favorov, A. et al. Exploring massive, genome scale datasets with the GenometriCorr package. PLoS Comput. Biol. 8, e1002529 (2012).
Acknowledgements
We thank J. Zacharias for technical assistance with sample processing and members of the Johns Hopkins DNA sequencing core facility for ChIP–seq on SOLiD formats. This work was supported by NIH grant CA38548 (A.P.F.), NIH grants CA140599 and CA179991 (C.A.I.-D.), the AACR Pancreatic Cancer Action Network Pathway to Leadership grant (O.G.M.), the Vanderbilt GI SPORE (O.G.M.), the Vanderbilt–Ingram Cancer Center (O.G.M.), and NIH grant CA180682 (A.M.-M.).
Author information
Authors and Affiliations
Contributions
O.G.M., C.A.I.-D., and A.P.F. conceived the work and wrote the manuscript. A.P.F. oversaw epigenomic sequencing performed at Johns Hopkins University. C.A.I.-D. selected patient samples, performed pathological review, analyzed immunostaining data, and oversaw whole-genome sequencing studies performed at MSKCC. O.G.M. designed and performed or oversaw all experiments and data analysis conducted at Vanderbilt. X.L. and H.W. performed the bioinformatic and statistical analyses. O.G.M. and A.P.F. guided the bioinformatic analyses. T.S. performed immunohistochemical and immunofluorescence experiments. R.T. prepared sequencing libraries and performed sequencing runs, assisted by T.H.S. S.J.M., M.O.W., and J.W.L. performed the LC–HRMS measurements of metabolites. O.G.M. and A.E.W. analyzed and plotted LC–HRMS data. A.C. and K.E.W. performed and analyzed YSI glucose and lactate measurements. S.N. and K.M.S. maintained cell culture and performed NADPH assays and a subset of immunoblotting. Y.Z. performed animal injection experiments. A.M.-M. performed whole-genome sequencing. O.G.M. prepared the figures. All authors approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Global reprogramming of histone modifications during the evolution of distant metastasis.
(a) IHC staining performed on formalin-fixed, paraffin-embedded (FFPE) tissue sections revealed diffusely positive H3K9me2 and H3K9me3 staining across PDAC cells from the founder clone (top), the subclone that seeded metastasis (middle), and the peritoneal metastasis (bottom) in patient A141. Scale bars, 100 μm. (b) In contrast, heterogeneous staining for H3K9me2 and H3K9me3 was detected in FFPE tissue sections taken from the primary tumor subclone that seeded metastasis (middle) and in the matched liver metastasis itself (bottom) in patient A132. Scale bars, 100 μm. (c) FFPE tissue sections were available from primary tumor subclones that seeded both peritoneal and distant metastasis in patient A38. Similar to patients A124 (Fig. 1a) and A141, diffusely positive staining for heterochromatin modifications was observed in both the founder clone (first panel) and the subclone that seeded peritoneal metastasis (second panel). In contrast, similar to patients A125 (Fig. 1b) and A132, heterogeneous staining between PDAC cells emerged in the primary tumor subclone that seeded distant metastases (third panel). The matched lung metastasis itself was diffusely negative (fourth panel), similar to cell lines isolated from this site and a matched liver metastasis in this patient (Fig. 1c). Scale bars, 100 μm. (d) Immunoblots on cell lines showed minimal or inconsistent changes for the indicated histone modifications between regional PDAC samples, including A38Per (Per). (e) In contrast, immunoblots on cell lines isolated from a panel of distant metastatic subclones showed recurrent changes in specific modifications, as compared to A38Per. (f) Global reprogramming of H3K9me3 and H4K20me2 was also observed between cell lines isolated from primary tumor subclones (Pr1, Pr2) and a matched lung metastasis in the same patient (pA13).
Supplementary Figure 2 Specificity of reprogrammed histone modifications.
(a) Ki67 staining showed similar cell cycle rates for peritoneal (A38Per) and matched lung (A38Lg) metastases from the same patient grown in medium with 10% FBS, and serum-free medium (SFM) greatly slowed growth in both subclones. Scale bars, 100 μm. (b) Serial cell counts for the indicated times confirmed similar growth rates and growth arrest in SFM. (c) GO analysis on RNA–seq data from cells cultured in serum versus SFM further confirmed growth arrest in SFM (lung data; the peritoneal cells gave identical results). (d) Immunoblots showed persistence of reprogrammed chromatin modifications in proliferative (FBS) and growth-arrested (SFM) cells. (e) Treatment of the peritoneal subclone with PDAC chemotherapies (Gem, gemcitabine; G+FU, gemcitabine + 5-fluorouracil) did not induce loss of methylation or gain of acetylation as seen between peritoneal and distant metastases (e.g., Fig. 1c,d), supporting the notion that those changes were independent of chemotherapy treatments. As expected, treatments resulted in increased levels of γH2AX, a signature of activated DNA repair pathways.
Supplementary Figure 3 Specificity of chromatin modifications within heterochromatin domains.
(a) Plots of ChIP–seq read densities normalized to input for histone modifications (y axes) showed that heterochromatin modifications (H3K9me2, H3K9me3, H3K27me3) were enriched in regions that were called heterochromatin (0% to 100%; bottom labels) for each subclone (indicated above). In contrast, euchromatin modifications (H3K36me3, H3K27ac) were depleted from the same regions. (b) Heterochromatin domains detected using alternative thresholds for domain calls produced similar results as shown in Figure 2a, demonstrating that the domain calls and reprogramming changes within them were robust across multiple sensitivity criteria (removal of short peaks less than 10 kb in length and merging of broad peaks separated by 15 kb are shown; other sensitivity tests using alternative thresholds produced nearly identical results that even more closely matched those in Fig. 2a).
Supplementary Figure 4 Reprogramming of H3K9me3 in LOCKs during PDAC subclonal evolution.
(a) ChIP–seq data from paired peritoneal (A38Per) and lung (A38Lg) metastatic subclones detected dramatic reduction of H3K9me3 in A38Lg that overlapped with H3K9me2 in LOCK domains. (b) Similar data were found for patient A13, which also showed loss of H3K9me3 from LOCK regions in the A13Pr2 and A13Lg subclones, as compared to the A13Pr1 primary tumor subclone.
Supplementary Figure 5 Localized reprogramming of DE gene loci within LOCK domains.
(a) Mapping DE genes from RNA–seq (distant metastases and precursors versus A38Per) to LOCKs revealed reciprocal changes in H3K27me3 and H3K27ac + H3K36me3 + DNA methylation around genes downregulated in LOCKs. (b) The opposite changes in H3K27me3 and H3K27ac + H3K36me3 were detected from genes upregulated in LOCKs. DNA methylation remained high in these regions.
Supplementary Figure 6 Specificity of chromatin modifications within ECDs.
(a) Plots of ChIP–seq read densities normalized to input for histone modifications (left labels) showed that euchromatin modifications (H3K36me3, H3K27ac; H3K4me3 is not shown) were enriched in regions that were called ECDs (0% to 100%; bottom labels) for each subclone (indicated above). In contrast, heterochromatin modifications (H3K9me2, H3K9me3, H3K27me3) were depleted from ECDs. (b) Euchromatin domains detected using alternative thresholds for domain calls (removal of short peaks less than 10 kb in length and merging of broad peaks separated by 15 kb) produced similar results as shown in Figure 2d, demonstrating that the domain calls were robust across multiple criteria. The y axes represent ChIP–seq read densities normalized to input for both panels.
Supplementary Figure 7 Reprogramming of large LOCK domains during PDAC evolution.
(a–c) H3K9me3 was enriched and DNA was hypomethylated within large LOCK domains. (b,c) Striking reciprocal reprogramming of H3K9me2 and H3K9me3 and DNA methylation was detected in a subset of large LOCKs in patient A38. (d) H3K9me3 was enriched across large HPDE LOCKs. (e) Examples of reprogrammed domains between samples across a 25-Mb locus on chromosome 13.
Supplementary Figure 8 Malignant heterogeneity between A38 subclones.
(a) Oxidoreductase capacity was measured with MTT assays performed on equal numbers of growth-arrested cells in the absence of serum, and MTT signals were normalized to total cell number per well. Consistent with GO results, A38Lg possessed higher oxidoreductase activity. n = 4 technical replicates; error bars, s.d. P values were calculated by two-tailed t test. (b) NADPH/NADP levels were measured on equal numbers of growth-arrested cells. More NADPH per 1 million cells was detected in A38Lg. n = 2 biological replicates; error bars, s.d. P values were calculated by two-tailed t test. (c) A38Per and A38Lg maintained well- or poorly differentiated morphology in patient tissues and across three separate in vitro culture conditions as indicated. (d) IF performed on fixed tissues from the primary tumor showed loss of E-cadherin with gain of vimentin in the precursor subclone that seeded A38Lg, consistent with EMT. (e,f) RNAi knockdown of KRAS (e) blocked 3D tumor formation in suspension assays (f) more efficiently in A38Per than A38Lg. n = 4 technical replicates; error bars, s.d. P values were calculated by two-tailed t test. Scale bars, 400 μm.
Supplementary Figure 9 Rearrangements were targeted to large LOCKs and ECDs in patient A38.
(a) Total breakpoints in the A38 genome were not significantly enriched within large LOCKs or ECDs. P values represent the results of χ2 analyses of observed breakpoints in each domain versus those expected if rearrangements were uniformly distributed across the genome. (b) Unlike typical LOCKs, rearrangements (breakpoint fusions) located within large LOCKs and ECDs were significantly enriched for joining of the breakpoints within homologous domains (e.g., ECD to ECD). χ2-test P values were calculated as above. (c) Examples of large LOCK domains where instability hotspots clustered, including both intrachromosomal (amplifications) and interchromosomal (translocations) rearrangements.
Supplementary Figure 10 Enhanced glucose metabolism with depleted 6PG across distant metastatic and precursor subclones.
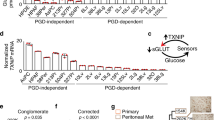
(a) Extracellular glucose consumption and lactate secretion were elevated in A38Lg, A13Pr1, and A13Lg relative to A38Per. (b) Schematic of glycolytic (outside) and PPP (boxed) metabolites, with intracellular metabolite levels detected by LC–HRMS plotted for each sample. Data represent LC–HRMS signals normalized to total protein for each sample, as indicated on the y axis.
Supplementary Figure 11 6AN targets glucose metabolism and the PGD step of the oxPPP.
(a) 6AN selectively slowed rates of extracellular glucose consumption and lactate secretion in precursor (A13Pr1) and metastatic (A13Lg, A38Lg) subclones, with no effect on glutamine or glutamate. (b) 6AN reduced incorporation of intracellular 13C-labeled glucose into metabolites downstream of the oxPPP. (c) 6AN greatly increased levels of the PGD substrate (6PG) and upstream metabolites with corresponding reductions in the levels of downstream products. (d) Scatterplots with correlation coefficients confirmed that 6AN gene expression changes were targeted to DE genes detected in A38Per (versus A38Lg) and that this was a partial, quantitative effect.
Supplementary Figure 12 6AN selectively modulated the reprogrammed chromatin state of distant metastatic subclones.
(a) 6AN treatments generally increased global H3K9me2 with corresponding decreased acetylation in distant metastatic subclones. (b) Normal cells and local regional PDACs did not show such changes. (c) 6AN changes persisted after 3 d of treatment (+) followed by removal of 6AN from the medium (+–) for an additional 3 d.
Supplementary Figure 13 6AN targeted reprogrammed LOCK regions.
(a) Mapping 6AN-repressed DE genes to A38Lg LOCKs revealed that these were located in reprogrammed LOCK-EI regions. (b) ChIP–seq on DMSO- versus 6AN-treated A38Lg detected a quantitative increase in LOCK-wide H3K9me2 levels from reprogrammed regions (as aligned to A38Per LOCKs). (c) ChIP–seq also detected 6AN-induced quantitative reductions in H3K27ac specifically from genes repressed from LOCKs. (d) Scatterplots with correlation coefficients showed that the quantitative reductions in H3K27ac from 6AN-downregulated LOCK genes also partially corresponded to genes that were co-(down)-regulated in A38Per (top), relative to all LOCK genes examined together (bottom). (e) Unlike H3K27ac, no significant differences in H3K27me3 levels were detected from genes repressed in LOCKs or other genes, similar to immunoblot findings.
Supplementary Figure 14 6AN selectively blocked 3D tumor formation in distant metastatic subclones.
(a,b) 6AN did not interfere with the ability of regional PDAC samples to form tumors in 3D Matrigel assays (a) or in 3D suspension tumorsphere assays (b). n = 2–4 technical replicates; error bars, s.d. (c) In contrast, 6AN strongly blocked the ability of distant metastatic subclones to form tumors in 3D suspension tumorsphere assays. n = 4 technical replicates; error bars, s.d. *P < 0.003, two-tailed t test. Scale bars, 400 μm.
Supplementary Figure 15 Reprogramming of the TOP2B locus in response to 6AN.
(a) 6AN did not alter mRNA expression (top) or levels of H3K9me2 and H3K27ac (bottom) across the TOP2B genomic locus in the peritoneal (A38Per) subclone (ChIP–qPCR primer positions are shown with respect to the TOP2B gene). (b) In contrast, 6AN selectively repressed mRNA expression (n = 4 technical replicates; *P < 0.002, two-tailed t test) and induced enrichment of H3K9me2 across the TOP2B locus in the lung metastatic subclone matched from the same patient (A38Lg). (c,d) Representative RT–qPCR (A38Lg) verified RNAi knockdown of CDH2 and TOP2B with minimal effect on VIM (vimentin) expression. n = 3 technical replicates, each normalized to ERK; error bars, s.d.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Tables 1, 5, 6 and 8–16, and Supplementary Note (PDF 4626 kb)
Supplementary Table 2
Data sets and correlation coefficients. (XLSX 30 kb)
Supplementary Table 3
Domain characteristics. (XLSX 14 kb)
Supplementary Table 4
Reprogramming within domains and sensitivity analyses. (XLSX 19 kb)
Supplementary Table 7
Gene expression changes. (XLSX 9329 kb)
Rights and permissions
About this article
Cite this article
McDonald, O., Li, X., Saunders, T. et al. Epigenomic reprogramming during pancreatic cancer progression links anabolic glucose metabolism to distant metastasis. Nat Genet 49, 367–376 (2017). https://doi.org/10.1038/ng.3753
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3753
This article is cited by
-
Pancreatic cancer acquires resistance to MAPK pathway inhibition by clonal expansion and adaptive DNA hypermethylation
Clinical Epigenetics (2024)
-
Oncogenic enhancers prime quiescent metastatic cells to escape NK immune surveillance by eliciting transcriptional memory
Nature Communications (2024)
-
Epigenetic reprogramming-induced guanidinoacetic acid synthesis promotes pancreatic cancer metastasis and transcription-activating histone modifications
Journal of Experimental & Clinical Cancer Research (2023)
-
Impaired histone inheritance promotes tumor progression
Nature Communications (2023)
-
Chromatin accessibility uncovers KRAS-driven FOSL2 promoting pancreatic ductal adenocarcinoma progression through up-regulation of CCL28
British Journal of Cancer (2023)