Abstract
The opposition between Polycomb repressive complexes (PRCs) and BAF (mSWI/SNF) complexes has a critical role in both development and disease. Mutations in the genes encoding BAF subunits contribute to more than 20% of human malignancies, yet the underlying mechanisms remain unclear, owing largely to a lack of assays to assess BAF function in living cells. To address this, we have developed a widely applicable recruitment assay system through which we find that BAF opposes PRC by rapid, ATP-dependent eviction, leading to the formation of accessible chromatin. The reversal of this process results in reassembly of facultative heterochromatin. Surprisingly, BAF-mediated PRC eviction occurs in the absence of RNA polymerase II (Pol II) occupancy, transcription, and replication. Further, we find that tumor-suppressor and oncogenic mutant BAF complexes have different effects on PRC eviction. The results of these studies define a mechanistic sequence underlying the resolution and formation of facultative heterochromatin, and they demonstrate that BAF opposes PRC on a minute-by-minute basis to provide epigenetic plasticity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Kennison, J.A. & Tamkun, J.W. Trans. -regulation of homeotic genes in Drosophila. New Biol. 4, 91–96 (1992).
Simon, J.A. & Kingston, R.E. Occupying chromatin: Polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol. Cell 49, 808–824 (2013).
Morgan, M.A. & Shilatifard, A. Chromatin signatures of cancer. Genes Dev. 29, 238–249 (2015).
Piunti, A. & Shilatifard, A. Epigenetic balance of gene expression by Polycomb and COMPASS families. Science 352, aad9780 (2016).
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011).
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 45, 592–601 (2013).
Shain, A.H. & Pollack, J.R. The spectrum of SWI/SNF mutations, ubiquitous in human cancers. PLoS One 8, e55119 (2013).
Kosho, T. & Okamoto, N. Genotype–phenotype correlation of Coffin–Siris syndrome caused by mutations in SMARCB1, SMARCA4, SMARCE1, and ARID1A. Am. J. Med. Genet. C. Semin. Med. Genet. 166C, 262–275 (2014).
Santen, G.W. et al. Mutations in SWI/SNF chromatin remodeling complex gene ARID1B cause Coffin–Siris syndrome. Nat. Genet. 44, 379–380 (2012).
Tsurusaki, Y. et al. Mutations affecting components of the SWI/SNF complex cause Coffin–Siris syndrome. Nat. Genet. 44, 376–378 (2012).
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2014).
Ho, L. et al. esBAF facilitates pluripotency by conditioning the genome for LIF/STAT3 signalling and by regulating Polycomb function. Nat. Cell Biol. 13, 903–913 (2011).
Tamkun, J.W. et al. Brahma: a regulator of Drosophila homeotic genes structurally related to the yeast transcriptional activator SNF2/SWI2. Cell 68, 561–572 (1992).
Tkachuk, D.C., Kohler, S. & Cleary, M.L. Involvement of a homolog of Drosophila Trithorax by 11q23 chromosomal translocations in acute leukemias. Cell 71, 691–700 (1992).
Kim, K.H. & Roberts, C.W. Targeting EZH2 in cancer. Nat. Med. 22, 128–134 (2016).
Varambally, S. et al. The Polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002).
Lee, W. et al. PRC2 is recurrently inactivated through EED or SUZ12 loss in malignant peripheral nerve sheath tumors. Nat. Genet. 46, 1227–1232 (2014).
Wu, J.I., Lessard, J. & Crabtree, G.R. Understanding the words of chromatin regulation. Cell 136, 200–206 (2009).
Wu, J.I. et al. Regulation of dendritic development by neuron-specific chromatin remodeling complexes. Neuron 56, 94–108 (2007).
Versteege, I. et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 394, 203–206 (1998).
Wilson, B.G. et al. Epigenetic antagonism between Polycomb and SWI/SNF complexes during oncogenic transformation. Cancer Cell 18, 316–328 (2010).
Kia, S.K., Gorski, M.M., Giannakopoulos, S. & Verrijzer, C.P. SWI/SNF mediates Polycomb eviction and epigenetic reprogramming of the INK4b-ARF-INK4a locus. Mol. Cell. Biol. 28, 3457–3464 (2008).
Kadoch, C. & Crabtree, G.R. Reversible disruption of mSWI/SNF (BAF) complexes by the SS18-SSX oncogenic fusion in synovial sarcoma. Cell 153, 71–85 (2013).
Li, G. et al. Jarid2 and PRC2, partners in regulating gene expression. Genes Dev. 24, 368–380 (2010).
van der Stoop, P. et al. Ubiquitin E3 ligase Ring1b/Rnf2 of Polycomb repressive complex 1 contributes to stable maintenance of mouse embryonic stem cells. PLoS One 3, e2235 (2008).
Feldman, N. et al. G9a-mediated irreversible epigenetic inactivation of Oct-3/4 during early embryogenesis. Nat. Cell Biol. 8, 188–194 (2006).
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is an essential component of the core pluripotency transcriptional network. Proc. Natl. Acad. Sci. USA 106, 5187–5191 (2009).
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is essential for embryonic stem cell self-renewal and pluripotency. Proc. Natl. Acad. Sci. USA 106, 5181–5186 (2009).
Young, R.A. Control of the embryonic stem cell state. Cell 144, 940–954 (2011).
Hathaway, N.A. et al. Dynamics and memory of heterochromatin in living cells. Cell 149, 1447–1460 (2012).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Crabtree, G.R. & Olson, E.N. NFAT signaling: choreographing the social lives of cells. Cell 109 (Suppl.), S67–S79 (2002).
Crabtree, G.R. & Schreiber, S.L. Three-part inventions: intracellular signaling and induced proximity. Trends Biochem. Sci. 21, 418–422 (1996).
Spencer, D.M., Wandless, T.J., Schreiber, S.L. & Crabtree, G.R. Controlling signal transduction with synthetic ligands. Science 262, 1019–1024 (1993).
de Bruijn, D.R. et al. Targeted disruption of the synovial sarcoma–associated SS18 gene causes early embryonic lethality and affects PPARBP expression. Hum. Mol. Genet. 15, 2936–2944 (2006).
Banaszynski, L.A., Liu, C.W. & Wandless, T.J. Characterization of the FKBP.rapamycin.FRB ternary complex. J. Am. Chem. Soc. 127, 4715–4721 (2005).
Schuettengruber, B., Chourrout, D., Vervoort, M., Leblanc, B. & Cavalli, G. Genome regulation by Polycomb and Trithorax proteins. Cell 128, 735–745 (2007).
Blackledge, N.P. et al. Variant PRC1 complex–dependent H2A ubiquitylation drives PRC2 recruitment and Polycomb domain formation. Cell 157, 1445–1459 (2014).
Kadoch, C. & Crabtree, G.R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Yen, K., Vinayachandran, V., Batta, K., Koerber, R.T. & Pugh, B.F. Genome-wide nucleosome specificity and directionality of chromatin remodelers. Cell 149, 1461–1473 (2012).
Lorch, Y., Zhang, M. & Kornberg, R.D. Histone octamer transfer by a chromatin-remodeling complex. Cell 96, 389–392 (1999).
Mendenhall, E.M. et al. GC-rich sequence elements recruit PRC2 in mammalian ES cells. PLoS Genet. 6, e1001244 (2010).
Stanton, B.Z. et al. Smarca4 ATPase mutations disrupt direct eviction of PRC1 from chromatin. Nat. Genet. http://dx.doi.org/10.1038/ng.3735 (2016).
Deal, R.B., Henikoff, J.G. & Henikoff, S. Genome-wide kinetics of nucleosome turnover determined by metabolic labeling of histones. Science 328, 1161–1164 (2010).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Jadhav, U. et al. Acquired tissue-specific promoter bivalency is a basis for PRC2 necessity in adult cells. Cell 165, 1389–1400 (2016).
Schnitzler, G., Sif, S. & Kingston, R.E. Human SWI/SNF interconverts a nucleosome between its base state and a stable remodeled state. Cell 94, 17–27 (1998).
Ronan, J.L., Wu, W. & Crabtree, G.R. From neural development to cognition: unexpected roles for chromatin. Nat. Rev. Genet. 14, 347–359 (2013).
Khavari, P.A., Peterson, C.L., Tamkun, J.W., Mendel, D.B. & Crabtree, G.R. BRG1 contains a conserved domain of the SWI2/SNF2 family necessary for normal mitotic growth and transcription. Nature 366, 170–174 (1993).
Narayanan, R. et al. Loss of BAF (mSWI/SNF) complexes causes global transcriptional and chromatin state changes in forebrain development. Cell Rep. 13, 1842–1854 (2015).
Yoo, A.S., Staahl, B.T., Chen, L. & Crabtree, G.R. MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 460, 642–646 (2009).
Yoo, A.S. et al. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476, 228–231 (2011).
Victor, M.B. et al. Generation of human striatal neurons by microRNA-dependent direct conversion of fibroblasts. Neuron 84, 311–323 (2014).
Singhal, N. et al. Chromatin-remodeling components of the BAF complex facilitate reprogramming. Cell 141, 943–955 (2010).
Lickert, H. et al. Baf60c is essential for function of BAF chromatin remodelling complexes in heart development. Nature 432, 107–112 (2004).
Tea, J.S. & Luo, L. The chromatin remodeling factor Bap55 functions through the TIP60 complex to regulate olfactory projection neuron dendrite targeting. Neural Dev. 6, 5 (2011).
Weinberg, P., Flames, N., Sawa, H., Garriga, G. & Hobert, O. The SWI/SNF chromatin remodeling complex selectively affects multiple aspects of serotonergic neuron differentiation. Genetics 194, 189–198 (2013).
Dawson, M.A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Acknowledgements
We would like to dedicate this paper to Joe Calarco, who died tragically during the course of this work. His energy, enthusiasm, and insight will continue to guide our work and his warmth brightens our memories.
We are grateful to N. Hathaway, O. Bell, L. Chen, and J. Ronan, for insightful comments and technical assistance. We thank M. Alhamadsheh for synthesis of FK1012. We also thank J. Buenrostro and B. Wu for helpful advice and technical assistance with accessibility studies and V. Petkova. C.K. was supported by the NIH Director's New Innovator Award DP2 (1DP2CA195762-01), the American Cancer Society Scholar Award (RSG-14-051-01-DMC), the Pew Scholar Award, the A.P. Giannini Foundation, the Alex's Lemonade Stand Foundation (ALSF) Young Investigator Award, and the NIH SARC Sarcoma SPORE Career Development and SPORE Project (5U54 CA168512-04) awards. G.R.C. is supported by NIH grants (CA163915, NS046789), CIRM RB4-05886, SFARI and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.K. and G.R.C. conceived of the study and wrote the paper. C.K., R.T.W., J.P.C., C.M.W., E.L.M., S.M.G.B., and E.J.C. planned and performed the experiments. J.L.P. performed data analysis and statistics.
Corresponding authors
Ethics declarations
Competing interests
C.K. and G.R.C. are scientific co-founders, shareholders, and consultants of Foghorn Therapeutics, Inc.
Integrated supplementary information
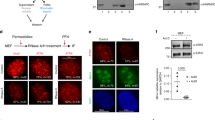
Supplementary Figure 1 Generation of a rapamycin-inducible recruitment system for mSWI/SNF (BAF) complexes to the Oct4 locus in MEFs.
(a) ChIP-seq tracks for Brg1 and Ezh2 over the Oct4 locus in ES cells and MEFs. (b) Lentiviral delivery vector design for Frb-V5-[BAF complex subunit, here shown, SS18] and direct fusion of FKBP to ZFHD1. (ZFHD1-FKBP) for tethering to binding array upstream of the modified Oct4 (Pou5f1) allele. (c) Enrichment of each mark over Oct4:WT and Oct4:CiA alleles in MEFs. The inserted allele and the wild-type allele show quantitatively similar chromatin landscapes. Error bars = Mean ± SD for n=3 experiments. (d) Baf47 and Baf57 Frb-V5 tagged complex subunits properly assemble into BAF complexes. (e) Density sedimentation analysis using 10-30% glycerol gradients indicate that introduced Frb-V5-Ss18 is stably incorporated as endogenous SS18 and is dedicated to the 2MDa BAF complexes. (f) Each Frb-V5 tagged BAF complex can be recruited by 20-40 fold upon 24 hours of rapamycin treatment. Error bars = Mean ± SD for n=3 experiments. (g) BAF complex recruitment (t=30’) to the Oct4:CiA locus in MEFs results in BAF complex occupancy levels similar to those in ES cells. Error bars = Mean ± SD for n=3 experiments. (h) BAF complex recruitment (fold enrichment by ChIP- qPCR) within the recruitment region at (left) +237bp, and (right)+377bp, from the ZHFD1 locus (-309bp and -169bp from the Oct4 TSS) using three different immunocapture antibodies.
Supplementary Figure 2 Chromatin marks, DNA accessibility, gene expression in BAF complex recruitment time course experiments.
(a) (left) Occupancy of PRC2 complexes and H3K27me3 is reduced at 60 minutes post-rapamycin treatment. (right) Recruitment of Frb-V5-Stop does not recruit BAF complexes, nor alter the occupancy of PRC2 or the H3K27me3 mark at t=60 minutes of rapamycin treatment. Error bars = Mean ± SD for n=3 experiments. (b) Total H3, H3K9me3, and H2A.Z are unchanged upon 60- minute BAF complex recruitment. Error bars = Mean ± SD for n=3 experiments. (c) Recruitment of Frb-V5-SS18 (BAF) to the Ascl1 locus. CATCH-IT results indicate nucleosomal turnover at 60’ post treatment with rapamycin. ChIP using V5, H3, H2AK119Ub shows site-specific decreases in histone mark enrichment upon BAF complex recruitment. Error bars = Mean ± SD for n=2 experiments for ChIP and n=4 experiments for CATCH-IT. (d) Schematic for modified ATAC-seq accessibility assays using Tn5 transposase. (e) Tn5 accessibility landscape over the CiA Oct4 locus shows enhanced accessibility within 60 minutes of BAF complex recruitment. (f) Overlap of Brg, Ezh2, and Ring1b MACS-called peaks displayed as Venn diagrams (published datasets). (g) Rapamycin addition (BAF complex recruitment) at various time points indicated does not result in increased percentage of GFP-positive cells (left) nor Oct4 gene expression (right) in CiA Oct4 MEFs. Error bars = Mean ± SD for n=3 experiments.
(h) RNA Pol II ChIP-qPCR over the CiA Oct4 locus indicates lack of recruitment upon BAF complex recruitment over a time course. Error bars = Mean ± SD for n=2 experiments.
Supplementary Figure 3 Structures and downstream reformation of heterochromatin in FK1012 competition (washout) experiments.
(A) Structures of (left) Rapamycin and (right) FK1012 showing the regions that bind FKBP, but not FRB, making FK1012 an effective competitor for rapamycin-induced recruitment of BAF complexes. (B) FK1012 competition (washout) triggers downstream removal of recruited BAF complexes at the +237bp from ZFHD1 site (-309bp from Oct4 TSS). (C-D) Reformation of downstream (C) PRC2 and (D) PRC1 repression upon FK1012 washout at the +237bp (from ZNFHD1 array) site. (E) DNA accessibility is lost and heterochromatin reforms downstream upon FK1012 washout. All experiments are a representative n=1 except Ring1b where error bars = Mean ± SD for n=3 experiments.
Supplementary Figure 4 SS18-SSX- containing (SS-like) BAF complexes display enhanced occupancy and PcG displacement at the Oct4 locus in MEFs.
(A) Recruitment of BAF complexes is similar at the recruitment site (+0bp; -443bp of the Oct4 TSS). (B-D) Eviction of (B) Ezh2, (C) Ring1b, and (D) H3K27me3 at the recruitment site (+0) is similar between WT BAF complexes and SS-like BAF complexes. (E) SS-like BAF complexes show enhanced occupancy downstream (+1034bp) of recruitment site. (F-H) Eviction of (F) Ezh2, (G) Ring1b, and (H) H3K27me3 downstream (+1034bp) of recruitment site is enhanced by SS-like BAF complexes as compared to wild type. Error bars represent Mean ± SEM for n=3 experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1 and 2 (PDF 1231 kb)
Rights and permissions
About this article
Cite this article
Kadoch, C., Williams, R., Calarco, J. et al. Dynamics of BAF–Polycomb complex opposition on heterochromatin in normal and oncogenic states. Nat Genet 49, 213–222 (2017). https://doi.org/10.1038/ng.3734
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3734
This article is cited by
-
The golden key to open mystery boxes of SMARCA4-deficient undifferentiated thoracic tumor: focusing immunotherapy, tumor microenvironment and epigenetic regulation
Cancer Gene Therapy (2024)
-
The BAF chromatin remodeler synergizes with RNA polymerase II and transcription factors to evict nucleosomes
Nature Genetics (2024)
-
Energy-driven genome regulation by ATP-dependent chromatin remodellers
Nature Reviews Molecular Cell Biology (2024)
-
RNA polymerase II promoter-proximal pausing promotes BAF chromatin binding and remodeling
Nature Genetics (2024)
-
Synovial sarcoma X breakpoint 1 protein uses a cryptic groove to selectively recognize H2AK119Ub nucleosomes
Nature Structural & Molecular Biology (2024)