Abstract
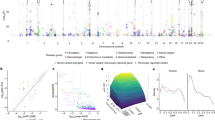
Clinical exome sequencing routinely identifies missense variants in disease-related genes, but functional characterization is rarely undertaken, leading to diagnostic uncertainty1,2. For example, mutations in PPARG cause Mendelian lipodystrophy3,4 and increase risk of type 2 diabetes (T2D)5. Although approximately 1 in 500 people harbor missense variants in PPARG, most are of unknown consequence. To prospectively characterize PPARγ variants, we used highly parallel oligonucleotide synthesis to construct a library encoding all 9,595 possible single–amino acid substitutions. We developed a pooled functional assay in human macrophages, experimentally evaluated all protein variants, and used the experimental data to train a variant classifier by supervised machine learning. When applied to 55 new missense variants identified in population-based and clinical sequencing, the classifier annotated 6 variants as pathogenic; these were subsequently validated by single-variant assays. Saturation mutagenesis and prospective experimental characterization can support immediate diagnostic interpretation of newly discovered missense variants in disease-related genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Majewski, J., Schwartzentruber, J., Lalonde, E., Montpetit, A. & Jabado, N. What can exome sequencing do for you? J. Med. Genet. 48, 580–589 (2011).
Gahl, W.A. et al. The National Institutes of Health Undiagnosed Diseases Program: insights into rare diseases. Genet. Med. 14, 51–59 (2012).
Barroso, I. et al. Dominant negative mutations in human PPARγ associated with severe insulin resistance, diabetes mellitus and hypertension. Nature 402, 880–883 (1999).
Jeninga, E.H., Gurnell, M. & Kalkhoven, E. Functional implications of genetic variation in human PPARγ. Trends Endocrinol. Metab. 20, 380–387 (2009).
Majithia, A.R. et al. Rare variants in PPARG with decreased activity in adipocyte differentiation are associated with increased risk of type 2 diabetes. Proc. Natl. Acad. Sci. USA 111, 13127–13132 (2014).
Flannick, J. et al. Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes. Nat. Genet. 45, 1380–1385 (2013).
Tennessen, J.A. et al. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 337, 64–69 (2012).
McLaughlin, H.M. et al. A systematic approach to the reporting of medically relevant findings from whole genome sequencing. BMC Med. Genet. 15, 134 (2014).
Manrai, A.K. et al. Genetic misdiagnoses and the potential for health disparities. N. Engl. J. Med. 375, 655–665 (2016).
Altshuler, D. et al. The common PPARγ Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat. Genet. 26, 76–80 (2000).
Claussnitzer, M. et al. Leveraging cross-species transcription factor binding site patterns: from diabetes risk loci to disease mechanisms. Cell 156, 343–358 (2014).
Tontonoz, P. & Spiegelman, B.M. Fat and beyond: the diverse biology of PPARγ. Annu. Rev. Biochem. 77, 289–312 (2008).
Agostini, M. et al. Non–DNA binding, dominant-negative, human PPARγ mutations cause lipodystrophic insulin resistance. Cell Metab. 4, 303–311 (2006).
Yu, S. et al. Adipocyte-specific gene expression and adipogenic steatosis in the mouse liver due to peroxisome proliferator–activated receptor γ1 (PPARγ1) overexpression. J. Biol. Chem. 278, 498–505 (2003).
Tontonoz, P., Nagy, L., Alvarez, J.G., Thomazy, V.A. & Evans, R.M. PPARγ promotes monocyte/macrophage differentiation and uptake of oxidized LDL. Cell 93, 241–252 (1998).
Barnes, M.R. & Gray, I.C. Bioinformatics for Geneticists (Wiley, 2003).
Chandra, V. et al. Structure of the intact PPAR-γ–RXR-α nuclear receptor complex on DNA. Nature 456, 350–356 (2008).
Garg, A. Acquired and inherited lipodystrophies. N. Engl. J. Med. 350, 1220–1234 (2004).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Myocardial Infarction Genetics Consortium Investigators. et al. Inactivating mutations in NPC1L1 and protection from coronary heart disease. N. Engl. J. Med. 371, 2072–2082 (2014).
González-Pérez, A. & López-Bigas, N. Improving the assessment of the outcome of nonsynonymous SNVs with a consensus deleteriousness score, Condel. Am. J. Hum. Genet. 88, 440–449 (2011).
Fowler, D.M. et al. High-resolution mapping of protein sequence–function relationships. Nat. Methods 7, 741–746 (2010).
Starita, L.M. et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 200, 413–422 (2015).
Demir, T. et al. Familial partial lipodystrophy linked to a novel peroxisome proliferator activator receptor-γ (PPARG) mutation, H449L: a comparison of people with this mutation and those with classic codon 482 Lamin A/C (LMNA) mutations. Diabet. Med. 33, 1445–1450 (2016).
Findlay, G.M., Boyle, E.A., Hause, R.J., Klein, J.C. & Shendure, J. Saturation editing of genomic regions by multiplex homology-directed repair. Nature 513, 120–123 (2014).
Fowler, D.M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Melnikov, A., Rogov, P., Wang, L., Gnirke, A. & Mikkelsen, T.S. Comprehensive mutational scanning of a kinase in vivo reveals substrate-dependent fitness landscapes. Nucleic Acids Res. 42, e112 (2014).
Brinkman, E.K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Forman, B.M. et al. 15-deoxy-δ12,14-prostaglandin J2 is a ligand for the adipocyte determination factor PPARγ. Cell 83, 803–812 (1995).
Ellard, S. et al. Improved genetic testing for monogenic diabetes using targeted next-generation sequencing. Diabetologia 56, 1958–1963 (2013).
Acknowledgements
This work was supported by grants from the National Institute of Diabetes and Digestive and Kidney Diseases (1K08DK102877-01, to A.R.M.; 1R01DK097768-01, to D.A.), NIH/Harvard Catalyst (1KL2TR001100-01, to A.R.M.), the Broad Institute (SPARC award, to A.R.M. and T.M.), and the Wellcome Trust (095564, to K.C.; 107064, to D.B.S.).
We thank J. Doench, C. Zhu, D. O'Connell, G. Cowley, M. Sullender, D. MacArthur, E. Minkel, B. Bulik-Sullivan, and J. Avruch for helpful discussions, laboratory assistance, and manuscript review.
This paper is dedicated to the memory of Promila Nandi (30 April 1933–27 December 2013).
Author information
Authors and Affiliations
Consortia
Contributions
A.R.M., T.M., and D.A. designed the study. A.R.M., B.T., M.A., and K.G. performed experiments with help from R.R., X.Z., M.F.B., and E.K. A.R.M. and N.P. analyzed the data with help from B.T., T.S., G.P., K.A.P., M.D., and T.M. I.B., S.E., S.K., S.O'R., K.C., and D.B.S. contributed clinical data and genotypes. A.R.M. and D.A. wrote the manuscript. D.B.S., S.O'R., K.C., E.D.R., and J.C.F. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of members and affiliations appears in the Supplementary Note.
A list of members and affiliations appears in the Supplementary Note.
A list of members and affiliations appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–4 and Supplementary Note. (PDF 2601 kb)
Source data
Rights and permissions
About this article
Cite this article
Majithia, A., Tsuda, B., Agostini, M. et al. Prospective functional classification of all possible missense variants in PPARG. Nat Genet 48, 1570–1575 (2016). https://doi.org/10.1038/ng.3700
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3700
This article is cited by
-
MmisAT and MmisP: an efficient and accurate suite of variant analysis toolkit for primary mitochondrial diseases
Human Genomics (2023)
-
satmut_utils: a simulation and variant calling package for multiplexed assays of variant effect
Genome Biology (2023)
-
PFAS Exposures and the Human Metabolome: A Systematic Review of Epidemiological Studies
Current Pollution Reports (2023)
-
Genotype-stratified treatment for monogenic insulin resistance: a systematic review
Communications Medicine (2023)
-
Saturation variant interpretation using CRISPR prime editing
Nature Biotechnology (2022)