Abstract
Long noncoding RNAs (lncRNAs) are prevalent genes with frequently precise regulation but mostly unknown functions. Here we demonstrate that lncRNAs guide the organismal DNA damage response. DNA damage activated transcription of the DINO (Damage Induced Noncoding) lncRNA via p53. DINO was required for p53-dependent gene expression, cell cycle arrest and apoptosis in response to DNA damage, and DINO expression was sufficient to activate damage signaling and cell cycle arrest in the absence of DNA damage. DINO bound to p53 protein and promoted its stabilization, mediating a p53 auto-amplification loop. Dino knockout or promoter inactivation in mice dampened p53 signaling and ameliorated acute radiation syndrome in vivo. Thus, inducible lncRNA can create a feedback loop with its cognate transcription factor to amplify cellular signaling networks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Komarova, E.A. et al. Dual effect of p53 on radiation sensitivity in vivo: p53 promotes hematopoietic injury, but protects from gastro-intestinal syndrome in mice. Oncogene 23, 3265–3271 (2004).
Kirsch, D.G. et al. p53 controls radiation-induced gastrointestinal syndrome in mice independent of apoptosis. Science 327, 593–596 (2010).
Lee, C.L. et al. p53 functions in endothelial cells to prevent radiation-induced myocardial injury in mice. Sci. Signal. 5, ra52 (2012).
Johnson, S.M. et al. Mitigation of hematologic radiation toxicity in mice through pharmacological quiescence induced by CDK4/6 inhibition. J. Clin. Invest. 120, 2528–2536 (2010).
Westphal, C.H. et al. Loss of atm radiosensitizes multiple p53 null tissues. Cancer Res. 58, 5637–5639 (1998).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Forrest, A.R. et al. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Sánchez, Y. et al. Genome-wide analysis of the human p53 transcriptional network unveils a lncRNA tumour suppressor signature. Nat. Commun. 5, 5812 (2014).
Léveillé, N. et al. Genome-wide profiling of p53-regulated enhancer RNAs uncovers a subset of enhancers controlled by a lncRNA. Nat. Commun. 6, 6520 (2015).
Hung, T. et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 43, 621–629 (2011).
Dimitrova, N. et al. LincRNA-p21 activates p21 in cis to promote Polycomb target gene expression and to enforce the G1/S checkpoint. Mol. Cell 54, 777–790 (2014).
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Li, M. et al. An Apela RNA-containing negative feedback loop regulates p53-mediated apoptosis in embryonic stem cells. Cell Stem Cell 16, 669–683 (2015).
Riley, K.J. & Maher, L.J. III. p53 RNA interactions: new clues in an old mystery. RNA 13, 1825–1833 (2007).
Lakin, N.D. & Jackson, S.P. Regulation of p53 in response to DNA damage. Oncogene 18, 7644–7655 (1999).
Loewer, A., Batchelor, E., Gaglia, G. & Lahav, G. Basal dynamics of p53 reveal transcriptionally attenuated pulses in cycling cells. Cell 142, 89–100 (2010).
Ulitsky, I., Shkumatava, A., Jan, C.H., Sive, H. & Bartel, D.P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147, 1537–1550 (2011).
Quinn, J.J. et al. Rapid evolutionary turnover underlies conserved lncRNA-genome interactions. Genes Dev. 30, 191–207 (2016).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Tsai, M.C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
Chu, C., Qu, K., Zhong, F.L., Artandi, S.E. & Chang, H.Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol. Cell 44, 667–678 (2011).
Giono, L.E. & Manfredi, J.J. The p53 tumor suppressor participates in multiple cell cycle checkpoints. J. Cell. Physiol. 209, 13–20 (2006).
Parrinello, S. et al. Oxygen sensitivity severely limits the replicative lifespan of murine fibroblasts. Nat. Cell Biol. 5, 741–747 (2003).
Lowe, S.W., Schmitt, E.M., Smith, S.W., Osborne, B.A. & Jacks, T. p53 is required for radiation-induced apoptosis in mouse thymocytes. Nature 362, 847–849 (1993).
Ihrie, R.A. et al. Perp is a mediator of p53-dependent apoptosis in diverse cell types. Curr. Biol. 13, 1985–1990 (2003).
Jeffers, J.R. et al. Puma is an essential mediator of p53-dependent and -independent apoptotic pathways. Cancer Cell 4, 321–328 (2003).
Qiu, W. et al. PUMA regulates intestinal progenitor cell radiosensitivity and gastrointestinal syndrome. Cell Stem Cell 2, 576–583 (2008).
Leibowitz, B.J. et al. Ionizing irradiation induces acute haematopoietic syndrome and gastrointestinal syndrome independently in mice. Nat. Commun. 5, 3494 (2014).
Hemann, M.T. et al. An epi-allelic series of p53 hypomorphs created by stable RNAi produces distinct tumor phenotypes in vivo. Nat. Genet. 33, 396–400 (2003).
Zhou, Y. et al. Activation of p53 by MEG3 non-coding RNA. J. Biol. Chem. 282, 24731–24742 (2007).
Wang, K.C. & Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011).
Kenzelmann Broz, D. et al. Global genomic profiling reveals an extensive p53-regulated autophagy program contributing to key p53 responses. Genes Dev. 27, 1016–1031 (2013).
Zhang, C. & Darnell, R.B. Mapping in vivo protein-RNA interactions at single-nucleotide resolution from HITS-CLIP data. Nat. Biotechnol. 29, 607–614 (2011).
Liu, T.X. et al. Knockdown of zebrafish Fancd2 causes developmental abnormalities via p53-dependent apoptosis. Dev. Cell 5, 903–914 (2003).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Rashid, N.U., Giresi, P.G., Ibrahim, J.G., Sun, W. & Lieb, J.D. ZINBA integrates local covariates with DNA-seq data to identify broad and narrow regions of enrichment, even within amplified genomic regions. Genome Biol. 12, R67 (2011).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Ideue, T., Hino, K., Kitao, S., Yokoi, T. & Hirose, T. Efficient oligonucleotide-mediated degradation of nuclear noncoding RNAs in mammalian cultured cells. RNA 15, 1578–1587 (2009).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nat. Protoc. 1, 1610–1616 (2006).
Das, R., Laederach, A., Pearlman, S.M., Herschlag, D. & Altman, R.B. SAFA: semi-automated footprinting analysis software for high-throughput quantification of nucleic acid footprinting experiments. RNA 11, 344–354 (2005).
Gherghe, C. et al. Definition of a high-affinity Gag recognition structure mediating packaging of a retroviral RNA genome. Proc. Natl. Acad. Sci. USA 107, 19248–19253 (2010).
Reuter, J.S. & Mathews, D.H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010).
Ouyang, Z., Snyder, M.P. & Chang, H.Y. SeqFold: genome-scale reconstruction of RNA secondary structure integrating high-throughput sequencing data. Genome Res. 23, 377–387 (2013).
Acknowledgements
We thank K. Nabel for able assistance. This study was supported by NIH R01-ES023168, R01-CA118750, R01-HG004361, P50-HG007735 to H.Y.C., NIH DP1-HD075622 to J.K.C., and ASCO Conquer Cancer Foundation/Susan G. Komen Young Investigator Award and Stanford Radiation Oncology Kaplan Funds to A.M.S. This research was funded in part through the NIH/NCI Cancer Center Support Grant P30 CA008748.
Author information
Authors and Affiliations
Contributions
H.Y.C. conceived and supervised the study. A.M.S., T.H., and H.Y.C. designed the experiments. A.M.S., J.T.G., T.H., R.A.F., A.Y.P., A.P.-d.-S., and R.B. performed experiments. A.M.S., J.T.G., T.H., R.A.F., Y.S., and K.Q. performed statistical analyses and analyzed the data. D.K.B., S.G., J.K.C., and L.D.A. contributed materials and advice. A.M.S., T.H., and H.Y.C. wrote the paper.
Corresponding author
Integrated supplementary information
Supplementary Figure 1 Identification of a DNA damage-inducible lncRNA.
(a) DINO tracks from RNA-seq of diploid human fibroblasts treated with doxorubicin and untreated controls (from ref. 16). DINO and CDKN1A exon 1 diagramed below. (b) DINO tracks from RNA-seq of TP53+/+ HCT116 cells treated with 5-fluorouracil and untreated control (from ref. 17). DINO and CDKN1A exon 1 diagramed below. (c) Northern blot of poly(A) selected RNA from human primary diploid fibroblasts treated with doxorubicin (16 h). Antisense RNA probe to DINO. (d) ChIP-qPCR results for DINO, CCNB1, and GAPDH loci in DNA damaged and untreated cells. Mean + standard deviation (s.d.) are shown for all bar graphs, n = 3. (e) Silver stained peptide gel of in vitro translation products from negative control (control), GFP, and DINO constructs. (f) Quantification of DINO copy number using an IVT DINO standard. (g) qRT-PCR of DINO in fibroblasts treated with indicated DNA damaging agents (16 h) or serum starvation (24 h). Below: western blot of p53 and β-actin after 6 h of treatment. Mean +/- s.d. plotted, n = 3. (h) qRT-PCR of DINO across a time course of DNA damage in U2OS cells. Mean +/- s.d. are shown for all bar graphs, n = 3. (i) DINO expression in TP53 -/- H1299 cells transfected with indicated p53 expression constructs +/- Dox (16 h). Mean +/- s.d. plotted, n = 3. (j) qRT-PCR of DINO induction at the indicated hours of doxorubicin treatment in cancer cell lines with known Li-Fraumeni p53 mutations. MDA-MB 468 (breast adenocarcinoma) and SW 636 (ovarian adenocarcinoma). Mean +/- s.d. plotted for all bar graphs, n = 3.
Supplementary Figure 2 Evolutionary conservation of a DNA damage-inducible lncRNA
(a) Location of Dino (green bar) is syntenic to human DINO. Grey bars indicate location of PCR primers tiling the Cdkn1a transcription start site. RNA transcripts were identified for forward strand, but not for the reverse strand. 3’ RACE reveals a poly(A) sequence at 3’ of Dino, indicating this transcript is not a splice variant of Cdkn1a. Below left: Genome browser snapshot of 3-seq in wild-type or Trp53-null MEFs +/- doxorubicin. Biological triplicates shown. Red box indicates mouse Dino reads. Location of mouse Dino (green bar) is shown above (3-seq dataset from ref. 18). Below right: Dino tracks of strand specific 3-seq demonstrated that Dino reads map to the forward strand. (b) qRT-PCR of Trp53, Cdkn1a and Dino expression in mouse ES V6.5 cells 6 h after doxorubicin. Mean +/- s.d. plotted, n = 3. (c) Northern blot with total RNA from ES V6.5 cells +/- doxorubicin (24 h) shows that mDino is an unspliced transcript. The 28S ribosome demonstrates equal total RNA loading in each lane. (d) Genomic context of zebrafish CDKN1A. Grey bars indicate PCR primer amplicons. (e) qRT-PCR of TP53 and CDKN1A in zebrafish embryos treated with UV at 18 hpf and harvested at 24 hpf. Letters correspond to grey bars in d. Mean +/- s.d. plotted, n = 3. (f) qRT-PCR of regions along the zebrafish CDKN1A promoter +/- UV. Mean +/- s.d. plotted, n = 3. (g) Map of conserved regions of DINO microhomology across eight species. All DINO orthologs are oriented with 5’ end of DINO on the left and 3’ end on the right. Direction of DINO ortholog transcription is established by qRT-PCR or RNA-seq in human, mouse, and zebrafish, and predicted based on nearest orthologs in marmoset, rat, elephant, horse, and rabbit. Ten highly conserved, strand-specific motifs were identified across eight species. Five motifs (*) map to domains of human DINO that interact with p53 on CLIP (Fig. 3c). All DINO orthologs contain each of these five motifs, with the exception of marmoset and zebrafish, which each contain four of the five motifs. Above: DINO diagram (from Fig. 3c), blue box indicating major p53 binding region on p53 CLIP. (h) Conserved motifs associated with regions of microhomology diagramed in g, motifs mapping to p53 interacting domains of human DINO (*). Motifs labeled 1 correspond to p53 CLIP domain “B” and the motif labeled 2 maps near p53 CLIP domain “D” (Fig. 3c). (i) Predicted DNA strand encoding DINO and CDKN1A in eight species.
Supplementary Figure 3 DINO regulates the DNA damage response
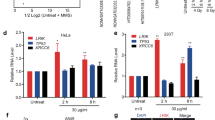
(a) DINO qRT-PCR in DNA damaged diploid human fibroblasts transfected with control or each of three DINO ASOs. Mean +/- s.d. plotted for all bar graphs, n = 3. (b) Heatmap of ATAC-seq signal at p53 response elements in human fibroblasts with indicated treatment conditions (corresponds to Fig. 2c). (c) qRT-PCR of DINO in U2OS cells stably expressing the indicated shRNAs. Mean +/- s.d. plotted, n = 3. (d) Doxorubicin (12 h) DNA damage-induced p53 occupancy of the indicated promoters in U2OS cells harboring control or shRNAs targeting p53 or DINO, as measured by ChIP-qPCR of p53. Mean +/- s.d. plotted, n = 3. (e) FACS plots of BrdU/PI labelled U2OS cells stably expressing indicated shRNA after 26 h of treatment with doxorubicin or untreated control. Black boxes indicate gates for quantification of each phase of the cell cycle and with percentage of cells in each phase displayed. (f) FACS plots of BrdU/PI labelled human fibroblasts treated with siDINO or siControl follow by doxorubicin for 24 h or left untreated. Black boxes indicate gates for quantification of each phase of the cell cycle and with percentage of cells in each phase displayed.
Supplementary Figure 4 DINO specifically interacts with p53
(a) RNA chromatography with DINO retrieves p53, but not SUZ12, from whole cell lysate of DNA damaged fibroblasts while LacZ mRNA does not; 2, 6 and 12 pmols of RNA were used. (b) RNA chromatography with DINO reveals no interaction with selected nuclear proteins. (c) qRT-PCR of RNA retrieval with indicated ChIRP probe pools from doxorubicin treated MCF10A cells. Mean +/- s.d. plotted, n = 3. (d) DINO ChIRP-qPCR in doxorubicin treated MCF10A cells demonstrates both RNA-dependent and RNA-independent retrieval of the DINO locus. Mean +/- s.d. plotted. *P < 0.05, n = 4. (e) DINO ChIRP-qPCR at ten p53-regulated (left) and five p53-independent loci (right) using two probe pools (even and odd). RNase pretreated chromatin (+RNase) for each probe pool serves as a negative control. Mean +/- s.d. plotted, n = 4.
Supplementary Figure 5 Structure and function of DINO’s p53 binding domain
(a) Normalized SHAPE reactivity of segment spanning the region of DINO enriched in the UV-CLIP-qPCR. (b) Secondary structure prediction of the enriched region of the DINO RNA. (c) Overexpression of DINOΔ320-450 fails to increase p53 protein stability above that of vector control in a cycloheximide chase analysis.
Supplementary Figure 6 Mouse Dino regulates DNA damage-induced apoptosis
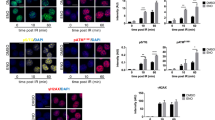
(a) qRT-PCR of multiple genes in thymocytes from Dino+/+ and Dinogfp/gfp mice 6 h after indicated treatment. Mean +/- s.d. plotted for each graph, n = 3. (b) Fraction of Dino+/+ and Dinogfp/gfp thymocytes in early apoptosis 6 h after indicated dose of radiation. Mean +/- s.d. plotted, n = 3. (c) Early apoptotic fraction of CD4+/CD8+ thymocytes 6 h after indicated dose or radiation, *P < 0.05 by Student’s t test. Mean +/- s.d. plotted, n = 3.
Supplementary Figure 7 Mouse Dino binds and stabilizes p53 to regulate the cell cycle
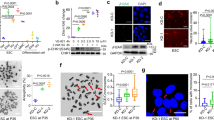
(a) Trp53 expression in early passage MEFs of the indicated genotypes. Mean +/- s.d. plotted for each graph, n = 3. (b) Gene expression heatmap of p53-responsive gene in the DNA damage response in early passage Dino+/+ and Dinolox/lox MEFs. (c) qRT-PCR of p53-dependent genes in early passage p21+/+ and p21-/- MEFs. Mean +/- s.d. plotted, n = 3. (d) qRT-PCR of p53-dependent genes in early passage MEFs of indicated genotypes culture in 0.1% serum. Mean +/- s.d. plotted, n = 3. (e) Quantification of p53 protein accumulation in early passage MEFs of the indicated genotype treated with increasing doses of doxorubicin. (f) Quantitative p53 protein abundance in Dino+/+ and Dinogfp/gfp MEFs with and without proteasome inhibitor MG-132 (10 μM). Mean +/- s.d. plotted, n = 3. (g) p53 UV-CLIP in MEFs of indicate genotypes following treatment with 0.2 μg/mL doxorubicin. Mean +/- s.d. plotted, n = 3.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1347 kb)
Supplementary Table 1
GO terms associated with DINO-dependent gene sets (XLSX 45 kb)
Supplementary Table 2
List of DINO-dependent genes identified in microarray analysis (XLSX 162 kb)
Supplementary Table 3
PCR primer pairs, siRNA and ASO sequences (XLSX 49 kb)
Rights and permissions
About this article
Cite this article
Schmitt, A., Garcia, J., Hung, T. et al. An inducible long noncoding RNA amplifies DNA damage signaling. Nat Genet 48, 1370–1376 (2016). https://doi.org/10.1038/ng.3673
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3673
This article is cited by
-
Microarray analysis identifies coding and non-coding RNA markers of liver injury in whole body irradiated mice
Scientific Reports (2023)
-
Long noncoding RNAs as versatile molecular regulators of cellular stress response and homeostasis
Human Genetics (2023)
-
LINC02126 is a potential diagnostic, prognostic and immunotherapeutic target for lung adenocarcinoma
BMC Pulmonary Medicine (2022)
-
Analysis of lncRNA-miRNA-mRNA expression pattern in heart tissue after total body radiation in a mouse model
Journal of Translational Medicine (2021)
-
Linc-SCRG1 accelerates progression of hepatocellular carcinoma as a ceRNA of miR26a to derepress SKP2
Journal of Experimental & Clinical Cancer Research (2021)