Abstract
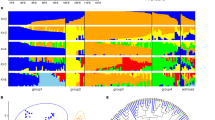
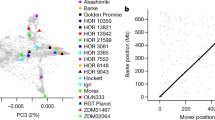
After domestication, during a process of widespread range extension, barley adapted to a broad spectrum of agricultural environments. To explore how the barley genome responded to the environmental challenges it encountered, we sequenced the exomes of a collection of 267 georeferenced landraces and wild accessions. A combination of genome-wide analyses showed that patterns of variation have been strongly shaped by geography and that variant-by-environment associations for individual genes are prominent in our data set. We observed significant correlations of days to heading (flowering) and height with seasonal temperature and dryness variables in common garden experiments, suggesting that these traits were major drivers of environmental adaptation in the sampled germplasm. A detailed analysis of known flowering-associated genes showed that many contain extensive sequence variation and that patterns of single- and multiple-gene haplotypes exhibit strong geographical structuring. This variation appears to have substantially contributed to range-wide ecogeographical adaptation, but many factors key to regional success remain unidentified.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Beddington, J.R. et al. What next for agriculture after Durban? Science 335, 289–290 (2012).
Challinor, A.J. et al. A meta-analysis of crop yield under climate change and adaptation. Nat. Clim. Chang. 4, 287–291 (2014).
McCouch, S. et al. Agriculture: feeding the future. Nature 499, 23–24 (2013).
Zohary, D., Hopf, M. & Weiss, E. Domestication of Plants in the Old World 4th edn. (Oxford University Press, 2013).
Dawson, I.K. et al. Barley: a translational model for adaptation to climate change. New Phytol. 206, 913–931 (2015).
Doebley, J.F., Gaut, B.S. & Smith, B.D. The molecular genetics of crop domestication. Cell 127, 1309–1321 (2006).
Mascher, M. et al. Barley whole exome capture: a tool for genomic research in the genus Hordeum and beyond. Plant J. 76, 494–505 (2013).
International Barley Genome Sequencing Consortium. A physical, genetic and functional sequence assembly of the barley genome. Nature 491, 711–716 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Mascher, M. et al. Anchoring and ordering NGS contig assemblies by population sequencing (POPSEQ). Plant J. 76, 718–727 (2013).
Kilian, B. et al. Haplotype structure at seven barley genes: relevance to gene pool bottlenecks, phylogeny of ear type and site of barley domestication. Mol. Genet. Genomics 276, 230–241 (2006).
Comadran, J. et al. Natural variation in a homolog of Antirrhinum CENTRORADIALIS contributed to spring growth habit and environmental adaptation in cultivated barley. Nat. Genet. 44, 1388–1392 (2012).
Hurst, L.D. Genetics and the understanding of selection. Nat. Rev. Genet. 10, 83–93 (2009).
Russell, J. et al. Analysis of >1000 single nucleotide polymorphisms in geographically matched samples of landrace and wild barley indicates secondary contact and chromosome-level differences in diversity around domestication genes. New Phytol. 191, 564–578 (2011).
Cavanagh, C.R. et al. Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc. Natl. Acad. Sci. USA 110, 8057–8062 (2013).
Li, Y. et al. Resequencing of 200 human exomes identifies an excess of low-frequency non-synonymous coding variants. Nat. Genet. 42, 969–972 (2010).
Meyer, R.S. & Purugganan, M.D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Künzel, G., Korzun, L. & Meister, A. Cytologically integrated physical restriction fragment length polymorphism maps for the barley genome based on translocation breakpoints. Genetics 154, 397–412 (2000).
Künzel, G. & Waugh, R. Integration of microsatellite markers into the translocation-based physical RFLP map of barley chromosome 3H. Theor. Appl. Genet. 105, 660–665 (2002).
Baker, K. et al. The low-recombining pericentromeric region of barley restricts gene diversity and evolution but not gene expression. Plant J. 79, 981–992 (2014).
Morrell, P.L., Gonzales, A.M., Meyer, K.K. & Clegg, M.T. Resequencing data indicate a modest effect of domestication on diversity in barley: a cultigen with multiple origins. J. Hered. 105, 253–264 (2014).
Taketa, S. et al. Barley grain with adhering hulls is controlled by an ERF family transcription factor gene regulating a lipid biosynthesis pathway. Proc. Natl. Acad. Sci. USA 105, 4062–4067 (2008).
Pourkheirandish, M. et al. Evolution of the grain dispersal system in barley. Cell 162, 527–539 (2015).
Begun, D.J. & Aquadro, C.F. Levels of naturally occurring DNA polymorphism correlate with recombination rates in D. melanogaster. Nature 356, 519–520 (1992).
Frichot, E., Mathieu, F., Trouillon, T., Bouchard, G. & François, O. Fast and efficient estimation of individual ancestry coefficients. Genetics 196, 973–983 (2014).
Russell, J. et al. Genetic diversity and ecological niche modelling of wild barley: refugia, large-scale post-LGM range expansion and limited mid-future climate threats? PLoS One 9, e86021 (2014).
Komatsuda, T. et al. Six-rowed barley originated from a mutation in a homeodomain–leucine zipper I–class homeobox gene. Proc. Natl. Acad. Sci. USA 104, 1424–1429 (2007).
Ramsay, L. et al. INTERMEDIUM-C, a modifier of lateral spikelet fertility in barley, is an ortholog of the maize domestication gene TEOSINTE BRANCHED 1. Nat. Genet. 43, 169–172 (2011).
Pavlidis, P., Živkovic, D., Stamatakis, A. & Alachiotis, N. SweeD: likelihood-based detection of selective sweeps in thousands of genomes. Mol. Biol. Evol. 30, 2224–2234 (2013).
Alachiotis, N., Stamatakis, A. & Pavlidis, P. OmegaPlus: a scalable tool for rapid detection of selective sweeps in whole-genome datasets. Bioinformatics 28, 2274–2275 (2012).
Günther, T. & Coop, G. Robust identification of local adaptation from allele frequencies. Genetics 195, 205–220 (2013).
Calixto, C.P.G., Waugh, R. & Brown, J.W.S. Evolutionary relationships among barley and Arabidopsis core circadian clock and clock-associated genes. J. Mol. Evol. 80, 108–119 (2015).
Jones, H. et al. Population-based resequencing reveals that the flowering time adaptation of cultivated barley originated east of the Fertile Crescent. Mol. Biol. Evol. 25, 2211–2219 (2008).
Turner, A., Beales, J., Faure, S., Dunford, R.P. & Laurie, D.A. The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310, 1031–1034 (2005).
Kumar, P., Henikoff, S. & Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Faure, S. et al. Mutation at the circadian clock gene EARLY MATURITY 8 adapts domesticated barley (Hordeum vulgare) to short growing seasons. Proc. Natl. Acad. Sci. USA 109, 8328–8333 (2012).
Casas, A.M. et al. HvFT1 (VrnH3) drives latitudinal adaptation in Spanish barleys. Theor. Appl. Genet. 122, 1293–1304 (2011).
Morrell, P.L., Lundy, K.E. & Clegg, M.T. Distinct geographic patterns of genetic diversity are maintained in wild barley (Hordeum vulgare ssp. spontaneum) despite migration. Proc. Natl. Acad. Sci. USA 100, 10812–10817 (2003).
Poets, A.M., Fang, Z., Clegg, M.T. & Morrell, P.L. Barley landraces are characterized by geographically heterogeneous genomic origins. Genome Biol. 16, 173 (2015).
Arend, D. et al. e!DAL—a framework to store, share and publish research data. BMC Bioinformatics 15, 214 (2014).
Jakob, S.S. et al. Evolutionary history of wild barley (Hordeum vulgare subsp. spontaneum) analyzed using multilocus sequence data and paleodistribution modeling. Genome Biol. Evol. 6, 685–702 (2014).
Pasam, R.K. et al. Genetic diversity and population structure in a legacy collection of spring barley landraces adapted to a wide range of climates. PLoS One 9, e116164 (2014).
Jones, H. et al. Evolutionary history of barley cultivation in Europe revealed by genetic analysis of extant landraces. BMC Evol. Biol. 11, 320 (2011).
Doyle, J.J. & Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 12, 13–15 (1990).
Harlan, J.R. Crops and Man (American Society of Agronomy and Crop Science Society of America, 1975).
Hijmans, R.J., Cameron, S.E., Parra, J.L., Jones, P.G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Clim. 25, 1965–1978 (2005).
Dray, S. & Dufour, A.B. The ade4 package: implementing the duality diagram for ecologists. J. Stat. Softw. 22, 1–20 (2007).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Jakobsson, M. & Rosenberg, N.A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806 (2007).
Thornton, K. Libsequence: a C++ class library for evolutionary genetic analysis. Bioinformatics 19, 2325–2327 (2003).
Hudson, R.R. Two-locus sampling distributions and their application. Genetics 159, 1805–1817 (2001).
Hufford, M.B. et al. Comparative population genomics of maize domestication and improvement. Nat. Genet. 44, 808–811 (2012).
Zeileis, A. & Grothendieck, G. zoo: S3 infrastructure for regular and irregular time series. J. Stat. Softw. 14, 6 (2005).
Bhatia, G., Patterson, N., Sankararaman, S. & Price, A.L. Estimating and interpreting FST: the impact of rare variants. Genome Res. 23, 1514–1521 (2013).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Nordborg, M. & Donnelly, P. The coalescent process with selfing. Genetics 146, 1185–1195 (1997).
Besag, J. & Clifford, P. Sequential Monte Carlo p-values. Biometrika 78, 301–304 (1991).
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979).
Nielsen, R. et al. Genomic scans for selective sweeps using SNP data. Genome Res. 15, 1566–1575 (2005).
Falush, D., Stephens, M. & Pritchard, J.K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Hardy, O.J. & Vekemans, X. SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol. Ecol. Notes 2, 618–620 (2002).
Tange, O. GNU parallel: the command-line power tool. USENIX Magazine 36, 42–47 (2011).
Nakamichi, N. Adaptation to the local environment by modifications of the photoperiod response in crops. Plant Cell Physiol. 56, 594–604 (2015).
Bandelt, H.J., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Micallef, L. & Rodgers, P. eulerAPE: drawing area-proportional 3-Venn diagrams using ellipses. PLoS One 9, e101717 (2014).
Smouse, P.E., Long, J.C. & Sokal, R.R. Multiple regression and correlation extensions of the Mantel test of matrix correspondence. Syst. Zool. 35, 627–632 (1986).
Acknowledgements
We specifically thank B. Thomas and A. Booth for helpful discussions around the common garden experiments. We would like to acknowledge funding from the Scottish Government Research Program to R.W., J.R., I.K.D., M.B. and I.M. and from the European Union Framework Programme 7 WHEALBI project to J.R., R.W., N.S., I.K.D., S.K. and B.K. M.v.Z. has been supported by the CGIAR Climate Change, Agriculture and Food Security (CCAFS) program. The work would not have been possible without funding from BBSRC grant BB/I00663X/1 to R.W., German Science Foundation (DFG) SPP1530 grant KI1465/6-1 to B.K. and BMBF TRITEX 0315954 A to N.S. Funding to G.J.M. was provided by the US Department of Agriculture–National Institute of Food and Agriculture (USDA-NIFA) as part of the Triticeae Coordinated Agricultural Project (TCAP), grant 2011-68002-30029. Analysis carried out by F.F. and K.S. was mainly performed on the computational resource bwUniCluster funded by the Ministry of Science, Research and the Arts Baden-Württemberg and the Universities of the State of Baden-Württemberg, Germany, within the framework program bwHPC.
Author information
Authors and Affiliations
Contributions
R.W., G.J.M., N.S. and B.K. conceived the study. B.K. selected the germplasm for inclusion and purified lines by two rounds of SSD. B.K., R.S., G.J.M., S.H., A. Hofstad and J.R. conducted the common garden experiments and analyzed the data. M.K., A. Himmelbach and N.S. generated the exome sequence data. J.R., I.K.D., K.S., F.F., M.M., S.K., C.C., M.B., J.W.S.B., M.v.Z., T.M.-G. and I.M. analyzed the data and provided information included in the supplementary files. R.W., M.M., I.K.D., J.R., F.F., N.S. and G.J.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Table 2–4 and 6, and Supplementary Figures 1–31. (PDF 7631 kb)
Supplementary Table 1
Information on 267 exome-captured barley accessions. (XLSX 146 kb)
Supplementary Table 5
Polymorphisms in flowering-associated genes. (XLSX 92 kb)
Rights and permissions
About this article
Cite this article
Russell, J., Mascher, M., Dawson, I. et al. Exome sequencing of geographically diverse barley landraces and wild relatives gives insights into environmental adaptation. Nat Genet 48, 1024–1030 (2016). https://doi.org/10.1038/ng.3612
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3612
This article is cited by
-
High-resolution mapping of Ryd4Hb, a major resistance gene to Barley yellow dwarf virus from Hordeum bulbosum
Theoretical and Applied Genetics (2024)
-
Genomic insights into positive selection during barley domestication
BMC Plant Biology (2022)
-
CCCH Zinc finger genes in Barley: genome-wide identification, evolution, expression and haplotype analysis
BMC Plant Biology (2022)
-
Accurate recombination estimation from pooled genotyping and sequencing: a case study on barley
BMC Genomics (2022)
-
Physical geography, isolation by distance and environmental variables shape genomic variation of wild barley (Hordeum vulgare L. ssp. spontaneum) in the Southern Levant
Heredity (2022)