Abstract
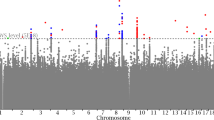
To identify genetic factors influencing quantitative traits of biomedical importance, we conducted a genome-wide association study in 8,842 samples from population-based cohorts recruited in Korea. For height and body mass index, most variants detected overlapped those reported in European samples. For the other traits examined, replication of promising GWAS signals in 7,861 independent Korean samples identified six previously unknown loci. For pulse rate, signals reaching genome-wide significance mapped to chromosomes 1q32 (rs12731740, P = 2.9 × 10−9) and 6q22 (rs12110693, P = 1.6 × 10−9), with the latter ∼400 kb from the coding sequence of GJA1. For systolic blood pressure, the most compelling association involved chromosome 12q21 and variants near the ATP2B1 gene (rs17249754, P = 1.3 × 10−7). For waist-hip ratio, variants on chromosome 12q24 (rs2074356, P = 7.8 × 10−12) showed convincing associations, although no regional transcript has strong biological candidacy. Finally, we identified two loci influencing bone mineral density at multiple sites. On chromosome 7q31, rs7776725 (within the FAM3C gene) was associated with bone density at the radius (P = 1.0 × 10−11), tibia (P = 1.6 × 10−6) and heel (P = 1.9 × 10−10). On chromosome 7p14, rs1721400 (mapping close to SFRP4, a frizzled protein gene) showed consistent associations at the same three sites (P = 2.2 × 10−3, P = 1.4 × 10−7 and P = 6.0 × 10−4, respectively). This large-scale GWA analysis of well-characterized Korean population-based samples highlights previously unknown biological pathways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rabbee, N. & Speed, T. A genotype calling algorithm for affymetrix SNP arrays. Bioinformatics 22, 7–12 (2006).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
WTCCC. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Weedon, M.N. et al. Genome-wide association analysis identifies 20 loci that influence adult height. Nat. Genet. 40, 575–583 (2008).
Hui, L., DelMonte, T. & Ranade, K. Genotyping using the TaqMan assay. Curr. Protoc. Hum. Genet. Chapter 2 Unit 2.10 (2008).
Gunderson, K.L. et al. Decoding randomly ordered DNA arrays. Genome Res. 14, 870–877 (2004).
Frayling, T.M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Loos, R.J. et al. Common variants near MC4R are associated with fat mass, weight and risk of obesity. Nat. Genet. 40, 768–775 (2008).
Liu, Y.J. et al. Genome-wide association scans identified CTNNBL1 as a novel gene for obesity. Hum. Mol. Genet. 17, 1803–1813 (2008).
Carafoli, E. The Ca2+ pump of the plasma membrane. J. Biol. Chem. 267, 2115–2118 (1992).
Okunade, G.W. et al. Targeted ablation of plasma membrane Ca2+-ATPase (PMCA) 1 and 4 indicates a major housekeeping function for PMCA1 and a critical role in hyperactivated sperm motility and male fertility for PMCA4. J. Biol. Chem. 279, 33742–33750 (2004).
Wang, G., Liszewski, M.K., Chan, A.C. & Atkinson, J.P. Membrane cofactor protein (MCP; CD46): isoform-specific tyrosine phosphorylation. J. Immunol. 164, 1839–1846 (2000).
Dooley, D.C., Oppenlander, B.K. & Xiao, M. Analysis of primitive CD34− and CD34+ hematopoietic cells from adults: gain and loss of CD34 antigen by undifferentiated cells are closely linked to proliferative status in culture. Stem Cells 22, 556–569 (2004).
Peters, N.S. New insights into myocardial arrhythmogenesis: distribution of gap-junctional coupling in normal, ischaemic and hypertrophied human hearts. Clin. Sci. (Lond.) 90, 447–452 (1996).
Yang, B. et al. The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat. Med. 13, 486–491 (2007).
Roell, W. et al. Engraftment of connexin 43-expressing cells prevents post-infarct arrhythmia. Nature 450, 819–824 (2007).
Howarth, F.C., Nowotny, N., Zilahi, E., El Haj, M.A. & Lei, M. Altered expression of gap junction connexin proteins may partly underlie heart rhythm disturbances in the streptozotocin-induced diabetic rat heart. Mol. Cell. Biochem. 305, 145–151 (2007).
Waerner, T. et al. ILEI: a cytokine essential for EMT, tumor formation, and late events in metastasis in epithelial cells. Cancer Cell 10, 227–239 (2006).
Cho, S.W. et al. Differential effects of secreted frizzled-related proteins (sFRPs) on osteoblastic differentiation of mouse mesenchymal cells and apoptosis of osteoblasts. Biochem. Biophys. Res. Commun. 367, 399–405 (2008).
Nakanishi, R. et al. Secreted frizzled-related protein 4 is a negative regulator of peak BMD in SAMP6 mice. J. Bone Miner. Res. 21, 1713–1721 (2006).
Nakanishi, R. et al. Osteoblast-targeted expression of sfrp4 in mice results in low bone mass. J. Bone Miner. Res. 23, 271–277 (2008).
Lettre, G. et al. Identification of ten loci associated with height highlights new biological pathways in human growth. Nat. Genet. 40, 584–591 (2008).
Gudbjartsson, D.F. et al. Many sequence variants affecting diversity of adult human height. Nat. Genet. 40, 609–615 (2008).
Pati, N., Schowinsky, V., Kokanovic, O., Magnuson, V. & Ghosh, S. A comparison between SNaPshot, pyrosequencing, and biplex invader SNP genotyping methods: accuracy, cost, and throughput. J. Biochem. Biophys. Methods 60, 1–12 (2004).
Hyndman, R.J. & Fan, Y. Sample quantiles in statistical packages. Am. Stat. 50, 361–365 (1996).
Devlin, B., Roeder, K. & Wasserman, L. Genomic control, a new approach to genetic-based association studies. Theor. Popul. Biol. 60, 155–166 (2001).
Price, A.L. et al. Principal component analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Ioannidis, J.P., Patsopoulos, N.A. & Evangelou, E. Heterogeneity in meta-analyses of genome-wide association investigations. PLoS ONE 2, e841 (2007).
Acknowledgements
This work was supported by a grant from the Ministry for Health, Welfare and Family Affairs, Republic of Korea (4845-301-430-260-00), and an intramural grant from the Korea National Institute of Health, Korea Center for Disease Control, Republic of Korea (4845-301-430-210-13).
Author information
Authors and Affiliations
Contributions
The study was designed by H-.L.K., B.O., J-.K.L. and J-.Y.L. Genotyping experiments were performed by J-.E.L., J.H.O., D-.J.K., M.P., S-.H.C., H-.Y.J. and E.Y.C. DNA sample preparation was carried out by M.H.L., J-.W.K. and B-.G.H. Phenotype information was collected by H.M., Y.A., M.S.P., N.H.C. and C.S. Statistical analysis was performed by M.J.G., D.Y., H.R.H., T.P., G.C. and Y.S.C. Bioinformatic analysis was conducted by Y.J.K., J.Y.H., H-.J.B., L.C. and Y.S.C. The manuscript was written by Y.S.C., B.O., J.W.P., J-.K.L., M.I.M. and H-.L.K. All authors reviewed the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–9, Supplementary Methods (PDF 3008 kb)
Rights and permissions
About this article
Cite this article
Cho, Y., Go, M., Kim, Y. et al. A large-scale genome-wide association study of Asian populations uncovers genetic factors influencing eight quantitative traits. Nat Genet 41, 527–534 (2009). https://doi.org/10.1038/ng.357
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.357
This article is cited by
-
Urinary microbiome-based metagenomic signature for the noninvasive diagnosis of hepatocellular carcinoma
British Journal of Cancer (2024)
-
UBAP2 plays a role in bone homeostasis through the regulation of osteoblastogenesis and osteoclastogenesis
Nature Communications (2023)
-
Zebrafish as a Model for Osteoporosis: Functional Validations of Genome-Wide Association Studies
Current Osteoporosis Reports (2023)
-
Multiple Mechanisms Explain Genetic Effects at the CPED1-WNT16 Bone Mineral Density Locus
Current Osteoporosis Reports (2023)
-
The KL genetic polymorphisms Associated with type 2 diabetes Mellitus
Indian Journal of Clinical Biochemistry (2023)