Abstract
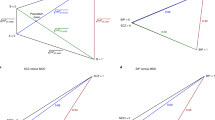
There is growing evidence of shared risk alleles for complex traits (pleiotropy), including autoimmune and neuropsychiatric diseases. This might be due to sharing among all individuals (whole-group pleiotropy) or a subset of individuals in a genetically heterogeneous cohort (subgroup heterogeneity). Here we describe the use of a well-powered statistic, BUHMBOX, to distinguish between those two situations using genotype data. We observed a shared genetic basis for 11 autoimmune diseases and type 1 diabetes (T1D; P < 1 × 10−4) and for 11 autoimmune diseases and rheumatoid arthritis (RA; P < 1 × 10−3). This sharing was not explained by subgroup heterogeneity (corrected PBUHMBOX > 0.2; 6,670 T1D cases and 7,279 RA cases). Genetic sharing between seronegative and seropostive RA (P < 1 × 10−9) had significant evidence of subgroup heterogeneity, suggesting a subgroup of seropositive-like cases within seronegative cases (PBUHMBOX = 0.008; 2,406 seronegative RA cases). We also observed a shared genetic basis for major depressive disorder (MDD) and schizophrenia (P < 1 × 10−4) that was not explained by subgroup heterogeneity (PBUHMBOX = 0.28; 9,238 MDD cases).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Sivakumaran, S. et al. Abundant pleiotropy in human complex diseases and traits. Am. J. Hum. Genet. 89, 607–618 (2011).
Cotsapas, C. et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet. 7, e1002254 (2011).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Fortune, M.D. et al. Statistical colocalization of genetic risk variants for related autoimmune diseases in the context of common controls. Nat. Genet. 47, 839–846 (2015).
Lee, S.H., Yang, J., Goddard, M.E., Visscher, P.M. & Wray, N.R. Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism–derived genomic relationships and restricted maximum likelihood. Bioinformatics 28, 2540–2542 (2012).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Pendergrass, S.A. et al. Phenome-wide association study (PheWAS) for detection of pleiotropy within the Population Architecture using Genomics and Epidemiology (PAGE) Network. PLoS Genet. 9, e1003087 (2013).
Collins, F.S. & Varmus, H. A new initiative on precision medicine. N. Engl. J. Med. 372, 793–795 (2015).
Criswell, L.A. et al. Analysis of families in the Multiple Autoimmune Disease Genetics Consortium (MADGC) collection: the PTPN22 620W allele associates with multiple autoimmune phenotypes. Am. J. Hum. Genet. 76, 561–571 (2005).
Kendler, K.S., Neale, M.C., Kessler, R.C., Heath, A.C. & Eaves, L.J. Major depression and generalized anxiety disorder. Same genes, (partly) different environments? Arch. Gen. Psychiatry 49, 716–722 (1992).
Wray, N.R., Goddard, M.E. & Visscher, P.M. Prediction of individual genetic risk to disease from genome-wide association studies. Genome Res. 17, 1520–1528 (2007).
International Schizophrenia Consortium. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Lee, S.H. et al. New data and an old puzzle: the negative association between schizophrenia and rheumatoid arthritis. Int. J. Epidemiol. 44, 1706–1721 (2015).
Power, R.A. et al. Polygenic risk scores for schizophrenia and bipolar disorder predict creativity. Nat. Neurosci. 18, 953–955 (2015).
Solovieff, N., Cotsapas, C., Lee, P.H., Purcell, S.M. & Smoller, J.W. Pleiotropy in complex traits: challenges and strategies. Nat. Rev. Genet. 14, 483–495 (2013).
Wray, N.R., Lee, S.H. & Kendler, K.S. Impact of diagnostic misclassification on estimation of genetic correlations using genome-wide genotypes. Eur. J. Hum. Genet. 20, 668–674 (2012).
Silverberg, M.S. et al. Diagnostic misclassification reduces the ability to detect linkage in inflammatory bowel disease genetic studies. Gut 49, 773–776 (2001).
van der Linden, M.P. et al. Value of anti–modified citrullinated vimentin and third-generation anti–cyclic citrullinated peptide compared with second-generation anti–cyclic citrullinated peptide and rheumatoid factor in predicting disease outcome in undifferentiated arthritis and rheumatoid arthritis. Arthritis Rheum. 60, 2232–2241 (2009).
Wiik, A.S., van Venrooij, W.J. & Pruijn, G.J. All you wanted to know about anti-CCP but were afraid to ask. Autoimmun. Rev. 10, 90–93 (2010).
Bromet, E.J. et al. Diagnostic shifts during the decade following first admission for psychosis. Am. J. Psychiatry 168, 1186–1194 (2011).
Gibson, P. et al. Subtypes of medulloblastoma have distinct developmental origins. Nature 468, 1095–1099 (2010).
Smoller, J.W., Lunetta, K.L. & Robins, J. Implications of comorbidity and ascertainment bias for identifying disease genes. Am. J. Med. Genet. 96, 817–822 (2000).
Burrell, R.A., McGranahan, N., Bartek, J. & Swanton, C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 501, 338–345 (2013).
Jeste, S.S. & Geschwind, D.H. Disentangling the heterogeneity of autism spectrum disorder through genetic findings. Nat. Rev. Neurol. 10, 74–81 (2014).
Flint, J. & Kendler, K.S. The genetics of major depression. Neuron 81, 484–503 (2014).
Cho, J.H. & Feldman, M. Heterogeneity of autoimmune diseases: pathophysiologic insights from genetics and implications for new therapies. Nat. Med. 21, 730–738 (2015).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP–trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Raychaudhuri, S. et al. Genetic variants at CD28, PRDM1 and CD2/CD58 are associated with rheumatoid arthritis risk. Nat. Genet. 41, 1313–1318 (2009).
Eyre, S. et al. High-density genetic mapping identifies new susceptibility loci for rheumatoid arthritis. Nat. Genet. 44, 1336–1340 (2012).
International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003).
Smyth, D.J. et al. Shared and distinct genetic variants in type 1 diabetes and celiac disease. N. Engl. J. Med. 359, 2767–2777 (2008).
Festen, E.A. et al. A meta-analysis of genome-wide association scans identifies IL18RAP, PTPN2, TAGAP, and PUS10 as shared risk loci for Crohn's disease and celiac disease. PLoS Genet. 7, e1001283 (2011).
Zhernakova, A. et al. Meta-analysis of genome-wide association studies in celiac disease and rheumatoid arthritis identifies fourteen non-HLA shared loci. PLoS Genet. 7, e1002004 (2011).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Cotsapas, C. & Hafler, D.A. Immune-mediated disease genetics: the shared basis of pathogenesis. Trends Immunol. 34, 22–26 (2013).
Onengut-Gumuscu, S. et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat. Genet. 47, 381–386 (2015).
Han, B. et al. Fine mapping seronegative and seropositive rheumatoid arthritis to shared and distinct HLA alleles by adjusting for the effects of heterogeneity. Am. J. Hum. Genet. 94, 522–532 (2014).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Ripke, S. et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Wray, N.R. & Maier, R. Genetic basis of complex genetic disease: the contribution of disease heterogeneity to missing heritability. Curr. Epidemiol. Rep. 1, 220–227 (2014).
Jennrich, R.I. An asymptotic χ2 test for the equality of two correlation matrices. J. Am. Stat. Assoc. 65, 904–912 (1970).
Wei, L.J., Lin, D.Y. & Weissfeld, L. Regression analysis of multivariate incomplete failure time data by modeling marginal distributions. J. Am. Stat. Assoc. 84, 1065–1073 (1989).
Lin, D.Y. & Sullivan, P.F. Meta-analysis of genome-wide association studies with overlapping subjects. Am. J. Hum. Genet. 85, 862–872 (2009).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Acknowledgements
This work was supported in part by funding from the US National Institutes of Health (NIH) (1R01AR063759 (S.R.), 1R01AR062886 (S.R.), 1UH2AR067677-01 (S.R.), and U19AI111224-01 (S.R.)) and Doris Duke Charitable Foundation grant 2013097. B.H. is supported by the Asan Institute for Life Sciences, Asan Medical Center, Seoul, Republic of Korea (2016-0717) and the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (HI14C1731). J.G.P. is supported by Fulbright Canada, the Weston Foundation, and Brain Canada through the Canada Brain Research Fund. K.S. is supported by an NIH training grant (T32HG002295). N.R.W. is supported by the Australian National Health and Medical Research Council (1087889 and 1078901). This research uses resources provided by the Type 1 Diabetes Genetics Consortium, a collaborative clinical study sponsored by the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), National Institute of Allergy and Infectious Diseases (NIAID), National Human Genome Research Institute (NHGRI), National Institute of Child Health and Human Development (NICHD), and Juvenile Diabetes Research Foundation International (JDRF) and supported by grant U01DK062418.
Author information
Authors and Affiliations
Consortia
Contributions
B.H. and S.R. conceived the statistical approach and organized the project. B.H., J.G.P., and S.R. led and coordinated analyses and wrote the initial manuscript. E.S. and N.R.W. provided guidance on the statistical approach. K.S., C.H.L., D.D., X.H., Y.R.P., and E.K. contributed to the implementation of specific analyses and offered feedback on the statistical methodologies. P.K.G., S.R.D., J.W., J.M., S.E., L.K., S.R., and T.H. contributed RA samples and insight on the clinical implications to RA. W.-M.C., S.O.-G., and S.S.R. contributed T1D samples and insight on clinical implications to T1D. The Major Depressive Disorder Working Group contributed MDD samples and insight on the clinical implications to MDD. All authors contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
Integrated supplementary information
Supplementary Figure 1 Power and false positive rate of the BUHMBOX and GRS approaches for different situations.
Under true pleiotropy without subgroup heterogeneity, the GRS approach appropriately demonstrated 64.8% power to detect shared genetic structure and BUHMBOX demonstrated an appropriate false positive rate of 4.3%. Under true subgroup heterogeneity without pleiotropy, the GRS approach demonstrated 100% power to detect shared genetic structure and BUHMBOX demonstrated 81.7% power to detect subgroup heterogeneity at α = 0.05. The GRS approach has power for both situations and therefore cannot distinguish subgroup heterogeneity from pleiotropy.
Supplementary Figure 2 Expected correlations with respect to heterogeneity proportion, risk allele frequencies, and odds ratios.
The plots show the expected correlations between pairs of alleles for different risk allele frequencies (RAFs) and odds ratios (ORs). (a) We assumed OR = 1.5 for both alleles. (b) We assumed RAF = 0.5 for both alleles.
Supplementary Figure 3 Power of BUHMBOX as a function of effect size.
This plot shows the power of BUHMBOX for detecting heterogeneity in terms of the mean effect size and the proportion of samples that comprise a hidden subgroup (heterogeneity proportion, π). We assume that we have 2,000 cases, 2,000 controls, and 50 associated loci. We sampled OR and RAF values from the GWAS catalog and scaled all log-transformed OR values to have a desired mean value. White lines denote 20%, 40%, 60%, and 80% power.
Supplementary Figure 4 Power of BUHMBOX in polygenic modeling.
We simulated GWAS and examined the power of BUHMBOX by including moderately significant loci up to P < 0.01. We modeled the genetic architecture of disease using rheumatoid arthritis, on the basis of results from Stahl et al. (Nat. Genet. 44, 483–489, 2012). To simulate 2,231 causal variants, we combined 71 independent known rheumatoid arthritis risk loci with an additional 2,160 loci sampled from the joint posterior distribution of RAF and OR values presented in Stahl et al. For null loci, we also used the null RAF distribution presented in Stahl et al. Given this disease model, we simulated a GWAS with 3,964 cases and 12,052 controls (sample sizes from Stahl et al.), assuming disease prevalence of 0.01. Given these GWAS results, we used only the top k GWAS loci defined by P-value threshold t and their observed OR estimates for BUHMBOX power simulations. We assumed N = 5,000 and π = 0.5 for power evaluation and tried different P-value thresholds t from 5 × 10−8 to 0.01.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1, 2, 5 and 6, and Supplementary Note. (PDF 4068 kb)
Supplementary Table 3
Detailed SNP information used for GRS and BUHMBOX analyses. (XLSX 136 kb)
Supplementary Table 4
GRS and BUHMBOX results. (XLSX 54 kb)
Rights and permissions
About this article
Cite this article
Han, B., Pouget, J., Slowikowski, K. et al. A method to decipher pleiotropy by detecting underlying heterogeneity driven by hidden subgroups applied to autoimmune and neuropsychiatric diseases. Nat Genet 48, 803–810 (2016). https://doi.org/10.1038/ng.3572
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3572
This article is cited by
-
An autoimmune pleiotropic SNP modulates IRF5 alternative promoter usage through ZBTB3-mediated chromatin looping
Nature Communications (2023)
-
Genetics of leprosy: today and beyond
Human Genetics (2020)
-
Shared genetic background between children and adults with attention deficit/hyperactivity disorder
Neuropsychopharmacology (2020)
-
Genetic correlations of polygenic disease traits: from theory to practice
Nature Reviews Genetics (2019)
-
Genetic pleiotropy between mood disorders, metabolic, and endocrine traits in a multigenerational pedigree
Translational Psychiatry (2018)