Abstract
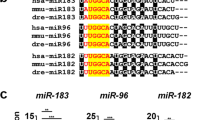
MicroRNAs (miRNAs) bind to complementary sites in their target mRNAs to mediate post-transcriptional repression1,2, with the specificity of target recognition being crucially dependent on the miRNA seed region3. Impaired miRNA target binding resulting from SNPs within mRNA target sites has been shown to lead to pathologies associated with dysregulated gene expression4,5,6,7. However, no pathogenic mutations within the mature sequence of a miRNA have been reported so far. Here we show that point mutations in the seed region of miR-96, a miRNA expressed in hair cells of the inner ear8, result in autosomal dominant, progressive hearing loss. This is the first study implicating a miRNA in a mendelian disorder. The identified mutations have a strong impact on miR-96 biogenesis and result in a significant reduction of mRNA targeting. We propose that these mutations alter the regulatory role of miR-96 in maintaining gene expression profiles in hair cells required for their normal function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Kloosterman, W.P. & Plasterk, R.H. The diverse functions of microRNAs in animal development and disease. Dev. Cell 11, 441–450 (2006).
He, L. & Hannon, G.J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Abelson, J.F. et al. Sequence variants in SLITRK1 are associated with Tourette's syndrome. Science 310, 317–320 (2005).
Clop, A. et al. A mutation creating a potential illegitimate microRNA target site in the myostatin gene affects muscularity in sheep. Nat. Genet. 38, 813–818 (2006).
Sethupathy, P. et al. Human microRNA-155 on chromosome 21 differentially interacts with its polymorphic target in the AGTR1 3′ untranslated region: a mechanism for functional single-nucleotide polymorphisms related to phenotypes. Am. J. Hum. Genet. 81, 405–413 (2007).
Jensen, K.P. et al. A common polymorphism in serotonin receptor 1B mRNA moderates regulation by miR-96 and associates with aggressive human behaviors. Mol. Psychiatry advance online publication, doi:10.1038/mp.2008.15 (19 February 2008).
Weston, M.D., Pierce, M.L., Rocha-Sanchez, S., Beisel, K.W. & Soukup, G.A. MicroRNA gene expression in the mouse inner ear. Brain Res. 1111, 95–104 (2006).
Modamio-Høybjør, S. et al. A novel locus for autosomal dominant nonsyndromic hearing loss, DFNA50, maps to chromosome 7q32 between the DFNB17 and DFNB13 deafness loci. J. Med. Genet. 41, e14 (2004).
Griffiths-Jones, S., Grocock, R.J., van Dongen, S., Bateman, A. & Enright, A.J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 34, D140–D144 (2006).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125, 887–901 (2006).
Duan, R., Pak, C. & Jin, P. Single nucleotide polymorphism associated with mature miR-125a alters the processing of pri-miRNA. Hum. Mol. Genet. 16, 1124–1131 (2007).
Denti, M.A., Rosa, A., Sthandier, O., De Angelis, F.G. & Bozzoni, I. A new vector, based on the PolII promoter of the U1 snRNA gene, for the expression of siRNAs in mammalian cells. Mol. Ther. 10, 191–199 (2004).
Enright, A. et al. MicroRNA targets in Drosophila. Genome Biol. 5, R1 (2003).
John, B. et al. Human microRNA targets. PLoS Biol. 2, 1862–1879 (2004).
Kertesz, M., Iovino, N., Unnerstall, U., Gaul, U. & Segal, E. The role of site accessibility in microRNA target recognition. Nat. Genet. 39, 1278–1284 (2007).
Lewis, M. et al. An ENU-induced mutation of miR96 associated with progressive hearing loss in mice. Nat. Genet. advance online publication, doi:10.1038/ng.369 (12 April 2009).
Xu, S., Witmer, P.D., Lumayag, S., Kovacs, B. & Valle, D. MicroRNA (miRNA) transcriptome of mouse retina and identification of a sensory organ-specific miRNA cluster. J. Biol. Chem. 282, 25053–25066 (2007).
Kloosterman, W.P., Wienholds, E., de Bruijn, E., Kauppinen, S. & Plasterk, R.H. In situ detection of miRNAs in animal embryos using LNA-modified oligonucleotide probes. Nat. Methods 3, 27–29 (2006).
Wienholds, E. et al. MicroRNA expression in zebrafish embryonic development. Science 309, 310–311 (2005).
Pierce, M.L. et al. MicroRNA-183 family conservation and ciliated neurosensory organ expression. Evol. Dev. 10, 106–113 (2008).
Calin, G.A. et al. Frequent deletions and down-regulation of microRNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 99, 15524–15529 (2002).
Soifer, H., Rossi, J.J. & Saetrom, P. MicroRNAs in disease and potential therapeautic applications. Mol. Ther. 15, 2070–2079 (2007).
del Castillo, I. et al. A deletion involving the connexin 30 gene in nonsyndromic hearing impairment. N. Engl. J. Med. 346, 243–249 (2002).
Tuddenham, L. et al. The cartilage specific microRNA-140 targets histone deacetylase 4 in mouse cells. FEBS Lett. 580, 4214–4217 (2006).
Pall, G.S., Codony-Servat, C., Byrne, J., Ritchie, L. & Hamilton, A. Carbodiimide-mediated cross-linking of RNA to nylon membranes improves the detection of siRNA, miRNA and piRNA by northern blot. Nucleic Acids Res. 35, e60 (2007).
Acknowledgements
We thank members of the Spanish families whose participation made this study possible. We also express our gratitude to A. Fatica and I. Bozzoni (Universita di Roma “La Sapienza”) for the donation of the microRNA expression vector and to P. Morales (Hospital 12 de Octubre) for providing control DNAs. S.M.-H. is recipient of a fellowship from L'Oréal-UNESCO “for women in Science”. A.M. is recipient of a contract from the Centre for Biomedical Research on Rare Diseases (CIBERER) and M.M. is recipient of a contract from the EuroHear project. F.M.-M. is a fellow from the Spanish Ministerio de Ciencia y Tecnología. This work was supported by grants from the Spanish Ministerio de Ciencia y Tecnología (SAF2005-06355 to F.M.), Spanish Fondo de Investigaciones anitarias (FIS CP03/00014 and PI08/0045 to M.A.M.-P; PI-051942; G03/203), the European Commission (FP6 Integrated Projects: SIROCCO LSHG-CT-2006-037900 to T.D.; EUROHEAR, LSHG-CT-2004-512063), Deafness Research UK and the Wellcome Trust.
Author information
Authors and Affiliations
Contributions
The exclusion of candidates genes within the DFNA50 genetic interval and the identification of the miR96 mutations were carried out by A.M., S.M.-H., M.M., F.M.-M. and L.O. Screening in autosomal recessive deafness families was designed and performed by I.d.C. and L.A.A. Vector generation for luciferase assays and miR96 expression was carried out by A.M., and the experiments were designed and performed by N.R. and T.D. Experimental design of the study and writing of the manuscript was carried out by M.A.M.-P. K.P.S. shared data and ideas. Scientific discussion of data and critical reading of the paper were performed by K.P.S., T.D., I.d.C., F.M. and M.A.M.-P.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Methods and Supplementary Tables 1–3 (PDF 415 kb)
Rights and permissions
About this article
Cite this article
Mencía, Á., Modamio-Høybjør, S., Redshaw, N. et al. Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss. Nat Genet 41, 609–613 (2009). https://doi.org/10.1038/ng.355
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.355
This article is cited by
-
AGO2 silences mobile transposons in the nucleus of quiescent cells
Nature Structural & Molecular Biology (2023)
-
Comparative expression analysis of the Atoh7 gene regulatory network in the mouse and chicken auditory hindbrain
Cell and Tissue Research (2023)
-
Investigating the characteristics of genes and variants associated with self-reported hearing difficulty in older adults in the UK Biobank
BMC Biology (2022)
-
MicroRNA editing patterns in Huntington’s disease
Scientific Reports (2022)
-
Genomic analysis of childhood hearing loss in the Yoruba population of Nigeria
European Journal of Human Genetics (2022)