Abstract
Heterotaxy results from a failure to establish normal left-right asymmetry early in embryonic development. By whole-exome sequencing, whole-genome sequencing and high-throughput cohort resequencing, we identified recessive mutations in MMP21 (encoding matrix metallopeptidase 21) in nine index cases with heterotaxy. In addition, Mmp21-mutant mice and mmp21-morphant zebrafish displayed heterotaxy and abnormal cardiac looping, respectively, suggesting a new role for extracellular matrix remodeling in the establishment of laterality in vertebrates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Primary accessions
European Nucleotide Archive
NCBI Reference Sequence
Referenced accessions
NCBI Reference Sequence
References
Sutherland, M.J. & Ware, S.M. Am. J. Med. Genet. C. Semin. Med. Genet. 151C, 307–317 (2009).
Jacobs, J.P. et al. Cardiol. Young 17 (suppl. 2), 1–28 (2007).
Lin, A.E. et al. Am. J. Med. Genet. A. 164A, 2581–2591 (2014).
Saunders, C.J. et al. Sci. Transl. Med. 4, 154ra135 (2012).
Brembeck, F.H. et al. Genes Dev. 18, 2225–2230 (2004).
Matsuura, K. et al. Nat. Commun. 2, 548 (2011).
Ahokas, K. et al. Gene 301, 31–41 (2002).
Marchenko, G.N., Marchenko, N.D. & Strongin, A.Y. Biochem. J. 372, 503–515 (2003).
Li, Y. et al. Nature 521, 520–524 (2015).
Akawi, N. et al. Nat. Genet. doi:10.1038/ng.3410 (2015).
Collins, M.M. & Ryan, A.K. Genesis 52, 488–502 (2014).
Yu, Q. & Stamenkovic, I. Genes Dev. 14, 163–176 (2000).
Gros, J., Feistel, K., Viebahn, C., Blum, M. & Tabin, C.J. Science 324, 941–944 (2009).
Soden, S.E. et al. Sci. Transl. Med. 6, 265ra168 (2014).
Fan, X., Abbott, T.E., Larson, D. & Chen, K. Curr. Protoc. Bioinformatics 45, 15.6.1–15.6.11 (2014).
Handsaker, R.E., Korn, J.M., Nemesh, J. & McCarroll, S.A. Nat. Genet. 43, 269–276 (2011).
Bahlo, M. & Bromhead, C.J. Bioinformatics 25, 1961–1962 (2009).
Smith, K.R. et al. Genome Biol. 12, R85 (2011).
Abecasis, G.R., Cherny, S.S., Cookson, W.O. & Cardon, L.R. Nat. Genet. 30, 97–101 (2002).
Gazal, S., Sahbatou, M., Babron, M.C., Genin, E. & Leutenegger, A.L. Bioinformatics 30, 1940–1941 (2014).
Gall, A.L. et al. J. Mol. Biol. 307, 577–586 (2001).
Thisse, C. & Thisse, B. Nat. Protoc. 3, 59–69 (2008).
Huang, C.J., Tu, C.T., Hsiao, C.D., Hsieh, F.J. & Tsai, H.J. Dev. Dyn. 228, 30–40 (2003).
Acknowledgements
We thank the Fédération Française de Cardiologie, the Société Française de Cardiologie, the Centre de Ressources Biologiques (CRB-ADN; BB-033-00065) at Institut Imagine, the zebrafish core facility of the Cardiovascular Development Consortium (CvDC), the Divisions of Clinical Genetics and Neonatology at Children's Mercy–Kansas City, the families for their participation, V. Salle, C. Ollagnier and B. Aime for excellent technical assistance, X. Liu and W. Devine for diagnosis of CHDs in mutant mouse embryos, and H. Roest Crollius and M. Delous for discussions. This work was supported by grants from the Fondation pour la Recherche Médicale (HeartGenomics), the Agence Nationale de la Recherche (ANR-10-IAHU-01), the Programme Hospitalier de Recherche Clinique (PHRC; 2008), the Fondation Renaud Fèbvre, the National Institute of Child Health and Human Development (NICHD) and the National Human Genome Research Institute (NHGRI) (U19HD077693), the National Center for Advancing Translational Sciences (NCATS; CTSA grant TL1TR000120), the National Heart, Lung, and Blood Institute (NHLBI; U01HL098180 and U01HL098188), the National Institute of General Medical Sciences (NIGMS; R01GM104412) and the US National Institutes of Health (S10RR029439).
Author information
Authors and Affiliations
Contributions
A.G. analyzed the whole-exome sequencing data, performed mutation validation, segregation studies and zebrafish experiments, and wrote the manuscript. G.C.G., M.S., N.T.K. and H.Y. performed and analyzed CRISPR/Cas9 and ENU experiments. F.B., L.D.S., S.D.F., P.S., S.L., L.d.P. and D.B. recruited patients. M.T. performed zebrafish experiments. H.L., R.E.M., G.J. and A.M.d.B. performed HaloPlex sequencing, mutation validation and segregation studies. A.N., N.A.M., C.J.S., I.T., L.D.C., E.G.F. and S.F.K. performed and analyzed whole-genome sequencing and performed deletion validation studies of family 2. M.O. performed mutation validation and segregation studies. C.N.B., K.A.P. and S.A.M. performed and analyzed CRISPR/Cas9 experiments. C.M., C.B.-F. and P.N. performed whole-exome sequencing. J.-F.D. performed linkage analysis. J.A. recruited patients, supervised genetic studies and wrote the manuscript. P.B. recruited patients, supervised HaloPlex studies, performed linkage analysis and homozygosity mapping, and wrote the manuscript. C.W.L. supervised CRISPR/Cas9 and ENU experiments and wrote the manuscript. C.T.G. analyzed whole-exome sequencing and CRISPR/Cas9 data, performed homology modeling, evolutionary analysis and zebrafish experiments, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Identification of a partial deletion of MMP21 in family 2 by WGS.
IGV screenshot of WGS reads aligned to the MMP21 locus on chromosome 10 in individuals F2-II:4 (proband), F2-II:3 (affected brother), F2-I:2 (unaffected mother) and F2-I:1 (unaffected father). Red stretched paired reads flank the deleted region chr10:127,460,915–127,466,819 in individuals F2-II:4, F2-II:3 and F2-I:2. The position of a frameshift mutation (c.365delT, p.Met122Serfs*55) is boxed in F2-II:4, F2-II:3 and F2-I:1 in exon 2 of MMP21. Genome coordinates are GRCh37/hg19.
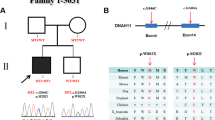
Supplementary Figure 3 Pedigrees of families 10 and 11.
In families 10 and 11, biallelic MMP21 variants were identified by NGS in the proband (numbered in red), but in each case one variant has been reported in the homozygous state in the ExAC browser.
Supplementary Figure 4 Parametric analysis of genome-wide SNP detection in families 6 and 7.
Genotyped individuals are numbered in red. The panels depict autosomal LOD score curves following parametric analysis of families 6 and 7 analyzed simultaneously. Parametric parameters were the following: mutation prevalence, 0.0001; phenocopy rate, 0.0; penetrance for homozygosity or compound heterozygosity, 1.0; penetrance for heterozygosity, 0.0. The intervals with LOD score above 3, on chromosomes 10 (containing MMP21) and 13, are indicated with a red bar. The chromosome 10 peak LOD score remained above 3 when penetrance was decreased to 50%. Thresholds of –2 and +3 are indicated by blue lines below and above the line at 0, respectively.
Supplementary Figure 5 Homozygosity mapping in family 7.
Regions of homozygosity in the two affected individuals of family 7 (III:3 and III:5) are indicated by red bars above the chromosome ideograms. The location of MMP21 is indicated by a red arrow on chromosome 10 in a region where both patients are homozygous.
Supplementary Figure 6 Homology modeling of the MMP21 peptidase domain and amino acids substituted in patients with heterotaxy.
The MMP21 model is based on the MMP11 crystal structure. The upper and lower panels are views of the model rotated approximately 180° around the vertical axis with respect to one another. The peptidase domain of MMPs forms a spherical topology consisting of three α helices and a five-stranded β sheet (Biochim. Biophys. Acta 1803, 20–28, 2010), colored here in blue (helices) and orange (β strands). Residues affected in patients with heterotaxy (Glu215, Ile226 and Ala321) are colored magenta. Predicted hydrogen bonds involving these three residues are indicated as green lines. The side chain of Glu215 is predicted to form a hydrogen bond with the side chain of Arg195 (the latter is in blue).
Supplementary Figure 7 Induction of aberrant splicing by a splice-blocking mmp21 morpholino.
(a) Schematic of the mmp21 zebrafish gene showing the location of MO1 targeting the exon 4/intron 4 splice donor and MO2 targeting the intron 4/exon 5 acceptor. 5′ and 3′ ISH probes are indicated by violet bars. (b) Electrophoresis gel after PCR performed on cDNA (using primers P3 and P6) from wild-type (wt) embryos and from embryos injected with MO1 or MO2. PCR product from primers P3/P6 runs at 584 bp in wild-type embryos. MO1 causes aberrant splicing, indicated by the black asterisk (new aberrant splice product). The red asterisk indicates that wild-type transcript levels are reduced by approximately 75% by MO1. MO2 does not cause aberrant splicing (white asterisk).
Supplementary Figure 8 Sequence confirmation of CRISPR-induced lesions in Mmp21.
Chromatograms showing Mmp21 mutations found in yolk sac DNA or in cloned PCR products amplified from yolk sac DNA, from F0 CRISPR-targeted mouse embryos. (a) Mutant embryo I226T-7 exhibits a homozygous 7-bp deletion, and embryo I226T-1 is compound heterozygous for a deletion in one allele (4/11 bacterial clones) and a knock-in of the desired A-to-G mutation causing the p.Ile226Thr substitution (in conjunction with three other synonymous single-base mutations; only two are shown here) on the other allele (7/11 bacterial colonies). (b) The A321P-12 mutant embryo has biallelic deletions predicted to cause frameshifts. Guides used in these experiments are shown in blue with the PAM sequence in green, and the corresponding orthologous human mutation site is shown in yellow.
Supplementary Figure 9 Multiz multiple-sequence alignment of MMP21 orthologs.
An alignment of MMP21 orthologs from 100 vertebrate species, via the Multiz Alignments track in the UCSC Genome Browser (human hg19 assembly). Exons are numbered at the top. Several vertebrate clades are listed to the right. Clades in which MMP21 orthologs are absent or incompletely aligned are in red (cetartiodactyla, birds, reptiles). In combination with the ORF decay observed in the alignable portions of cetartiodactyl MMP21 sequences (Supplementary Fig. 10), this distribution is suggestive of convergent gene loss in these three clades. For the cetartiodactyl, bird and reptile species for which a genome assembly is available in the UCSC Genome Browser, we confirmed that the region aligned in Multiz fell within the MMP21 syntenic region, i.e., between EDRF1 and UROS. We also performed BLAST analysis (tblastn) at the NCBI server using the human MMP21 protein sequence and were unable to identify an MMP21 ortholog in birds and reptiles. Note that, although exon 1 for some fish and mammals is not aligned in Multiz, this appears more likely to be due to genome assembly gaps or functional sequence divergence rather than ORF decay.
Supplementary Figure 10 MMP21 ORF degradation in cetartiodactyla.
(a,b) The MMP21 ORFs of selected vertebrates are aligned with regions of exon 6 (a) and exon 7 (b) of human MMP21, via the Multiz Alignments track at the UCSC Genome Browser (hg19). MMP21 sequences from all nine available cetartiodactyl genomes are depicted in each panel, plus a selection of other vertebrates. For the first five exons of MMP21, not all nine cetartiodactyl sequences are alignable in Multiz (Supplementary Fig. 9). The regions depicted in exons 6 and 7 show examples of ORF degradation in all nine cetartiodactyls; stops (asterisks), insertions and deletions.
Supplementary Figure 11 Pattern of loss of MMP21 from vertebrate genomes may correlate with absence of nodal cilia-generated asymmetry.
On the basis of the absence of evidence for an intact MMP21 ORF in the cetartiodactyl, bird and reptile clades and the previous suggestion that in chick and pig left-right asymmetry is generated not by leftward fluid flow but rather by rotational cell movements, we hypothesize that the presence of MMP21 correlates with the presence of cilia-driven and/or cilia-sensing nodal flow. The vertebrate phylogenetic tree is reproduced from the UCSC Genome Browser page “Vertebrate Multiz Alignment & Conservation (100 Species)," hg19 assembly.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Tables 1, 3, 6–10 and 12. (PDF 3459 kb)
Supplementary Table 2
Patient phenotypes and MMP21 genotypes for families 1 to 9. na, not available. (XLSX 14 kb)
Supplementary Table 4
Phenotypic and variant data for families 10 and 11. (XLSX 11 kb)
Supplementary Table 5
Genes identified in linkage intervals following parametric analysis of genome-wide SNP detection in families 6 and 7. rs2461224 and rs670243 were used as boundaries for the interval with LOD score above 3 on chromosome 10 (containing MMP21). rs544400 and rs9316663 were used as boundaries for the interval with LOD score above 3 on chromosome 13. (XLSX 34 kb)
Supplementary Table 11
Sequence of CRISPR/Cas9-treated Mmp21 alleles obtained by TOPO cloning of yolk sac DNA PCR products from 20 F0 embryos. In the column “sequence,” the position of the knock-in designed to model human mutation p.Ile226Thr is in red. For the mutation designed to model p.Ala321Pro, the position of the targeted mouse nucleotide falls outside of the sequence depicted (no embryos harbored this mutation). In the column “Allele Frequency,” ratios in brackets indicate the proportions of bacterial colonies obtained for a given allele type in each of the 20 embryos. In the protein sequence column, altered amino acids are indicated in red. In the protein summary column, coordinates refer to mouse RefSeq sequence NP_694423.1. (XLSX 14 kb)
Rights and permissions
About this article
Cite this article
Guimier, A., Gabriel, G., Bajolle, F. et al. MMP21 is mutated in human heterotaxy and is required for normal left-right asymmetry in vertebrates. Nat Genet 47, 1260–1263 (2015). https://doi.org/10.1038/ng.3376
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3376
This article is cited by
-
De novo disruptive heterozygous MMP21 variants are potential predisposing genetic risk factors in Chinese Han heterotaxy children
Human Genomics (2022)
-
Discovery of a genetic module essential for assigning left–right asymmetry in humans and ancestral vertebrates
Nature Genetics (2022)
-
Bicc1 and Dicer regulate left-right patterning through post-transcriptional control of the Nodal inhibitor Dand5
Nature Communications (2021)
-
Broad spectrum of CRISPR-induced edits in an embryonic lethal gene
Scientific Reports (2021)
-
Whole-exome sequencing reveals a combination of extremely rare single-nucleotide polymorphism of DNAH9 and RSPH1 genes in a Japanese fetus with situs viscerum inversus
Medical Molecular Morphology (2021)