Abstract
Mycosis fungoides and Sézary syndrome comprise the majority of cutaneous T cell lymphomas (CTCLs), disorders notable for their clinical heterogeneity that can present in skin or peripheral blood. Effective treatment options for CTCL are limited, and the genetic basis of these T cell lymphomas remains incompletely characterized1. Here we report recurrent point mutations and genomic gains of TNFRSF1B, encoding the tumor necrosis factor receptor TNFR2, in 18% of patients with mycosis fungoides and Sézary syndrome. Expression of the recurrent TNFR2 Thr377Ile mutant in T cells leads to enhanced non-canonical NF-κB signaling that is sensitive to the proteasome inhibitor bortezomib. Using an integrative genomic approach, we additionally discovered a recurrent CTLA4-CD28 fusion, as well as mutations in downstream signaling mediators of these receptors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wong, H.K., Mishra, A., Hake, T. & Porcu, P. Evolving insights in the pathogenesis and therapy of cutaneous T-cell lymphoma (mycosis fungoides and Sezary syndrome). Br. J. Haematol. 155, 150–166 (2011).
Kiessling, M.K. et al. High-throughput mutation profiling of CTCL samples reveals KRAS and NRAS mutations sensitizing tumors toward inhibition of the RAS/RAF/MEK signaling cascade. Blood 117, 2433–2440 (2011).
Vaqué, J.P. et al. PLCG1 mutations in cutaneous T-cell lymphomas. Blood 123, 2034–2043 (2014).
So, T. & Croft, M. Regulation of PI-3-kinase and Akt signaling in T lymphocytes and other cells by TNFR family molecules. Front. Immunol. 4, 139 (2013).
Lee, C.S. et al. Transcriptome sequencing in Sezary syndrome identifies Sezary cell and mycosis fungoides–associated lncRNAs and novel transcripts. Blood 120, 3288–3297 (2012).
Newman, A.M. et al. FACTERA: a practical method for the discovery of genomic rearrangements at breakpoint resolution. Bioinformatics 30, 3390–3393 (2014).
Migliazza, A. et al. Heterogeneous chromosomal aberrations generate 3′ truncations of the NFKB2/lyt-10 gene in lymphoid malignancies. Blood 84, 3850–3860 (1994).
Zhang, J., Chang, C.C., Lombardi, L. & Dalla-Favera, R. Rearranged NFKB2 gene in the HUT78 T-lymphoma cell line codes for a constitutively nuclear factor lacking transcriptional repressor functions. Oncogene 9, 1931–1937 (1994).
Vasmatzis, G. et al. Genome-wide analysis reveals recurrent structural abnormalities of TP63 and other p53-related genes in peripheral T-cell lymphomas. Blood 120, 2280–2289 (2012).
Nguyen, L.T. & Ohashi, P.S. Clinical blockade of PD1 and LAG3—potential mechanisms of action. Nat. Rev. Immunol. 15, 45–56 (2015).
Ashkenazi, A. & Dixit, V.M. Death receptors: signaling and modulation. Science 281, 1305–1308 (1998).
Shin, J.H. et al. Positive conversion of negative signaling of CTLA4 potentiates antitumor efficacy of adoptive T-cell therapy in murine tumor models. Blood 119, 5678–5687 (2012).
Mellman, I., Coukos, G. & Dranoff, G. Cancer immunotherapy comes of age. Nature 480, 480–489 (2011).
Sekulic, A. et al. Personalized treatment of Sezary syndrome by targeting a novel CTLA4:CD28 fusion. Mol. Genet. Genomic Med. 3, 130–136 (2015).
Cabal-Hierro, L. & Lazo, P.S. Signal transduction by tumor necrosis factor receptors. Cell. Signal. 24, 1297–1305 (2012).
van Doorn, R. et al. Oncogenomic analysis of mycosis fungoides reveals major differences with Sezary syndrome. Blood 113, 127–136 (2009).
Heemann, C. et al. Circulating levels of TNF receptor II are prognostic for patients with peripheral T-cell non-Hodgkin lymphoma. Clin. Cancer Res. 18, 3637–3647 (2012).
Lin, W.M. et al. Characterization of the DNA copy-number genome in the blood of cutaneous T-cell lymphoma patients. J. Invest. Dermatol. 132, 188–197 (2012).
Rodríguez, M. et al. NF-κB signal triggering and termination by tumor necrosis factor receptor 2. J. Biol. Chem. 286, 22814–22824 (2011).
Zinzani, P.L. et al. Phase II trial of proteasome inhibitor bortezomib in patients with relapsed or refractory cutaneous T-cell lymphoma. J. Clin. Oncol. 25, 4293–4297 (2007).
Acuto, O. & Michel, F. CD28-mediated co-stimulation: a quantitative support for TCR signalling. Nat. Rev. Immunol. 3, 939–951 (2003).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Koboldt, D.C. et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics 25, 2283–2285 (2009).
Deng, X. SeqGene: a comprehensive software solution for mining exome- and transcriptome-sequencing data. BMC Bioinformatics 12, 267 (2011).
Ng, S.B. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, 272–276 (2009).
Liu, X., Jian, X. & Boerwinkle, E. dbNSFP v2.0: a database of human non-synonymous SNVs and their functional predictions and annotations. Hum. Mutat. 34, E2393–E2402 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Shi, J. & Walker, M.G. Gene set enrichment analysis (GSEA) for interpreting gene expression profiles. Curr. Bioinformatics 2, 133–137 (2007).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Acknowledgements
We thank A. Alizadeh, H. Chang, S. Artandi, L. Boxer, D. Webster and J. Kovalski for presubmission review of the manuscript. We thank all the patients who generously participated in this study. This work was supported by the Office of Research and Development of the US Department of Veterans Affairs and by US National Institutes of Health (NIH) grant R01CA142635 awarded to P.A.K. A.U. was supported by US NIH grant F32CA168091 and by the American Society of Hematology Research Training Award for Fellows. A.B. received support from US NIH grant F310CA180408. E.R. received support from the Dermatology Foundation. The studies were generously supported by the Haas Family Foundation and the Drs. Martin and Dorothy Spatz Charitable Foundation.
Author information
Authors and Affiliations
Contributions
A.U. and A.B. designed and executed experiments, analyzed data and wrote the manuscript. E.R., C.S.L., J.R., A.M., A.Z., R.O., S.K., R.A., W.-K.W. and D.G. helped execute experiments, analyzed data and contributed to the design of experimentation. M.T. performed clinical data analysis. M.S. helped design experiments. A.R. and Y.K. provided clinical data and analysis. P.A.K. designed experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Details of sequencing analysis.
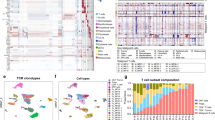
(a) Flow chart showing which patients fall into each category and were used for analysis. (b) Graph showing the average and median coverage for all samples that were resequenced. (c) Graph showing the sensitivity and specificity of each of the mutation callers used in our analysis pipeline as compared to the gold standard of a SNP array. (d) Box plots showing the range, quartiles and mean of allele frequency for each mutation in a patient in stage 1, 2 or 4. Stage 3 was not included due to small sample size. Allele frequencies were obtained from the mutational caller used to make the call as described in Supplementary Table 2.
Supplementary Figure 2 Clinical and genomic annotations of patients with Mycosis fungoides and Sézary syndrome.
(a) Somatically acquired mutations in CTCL samples of genes mutated in at least 5% of patients. Identical recurrent mutations marked in blue were validated by Sanger sequencing. Diagnosis of Mycosis fungoides or Sézary syndrome is marked, as is large-cell transformation (LCT). (b) Pathway diagram of canonical NF-κB1 signaling highlighting multiple pathway members altered in the targeted resequencing data set.
Supplementary Figure 3 Additional fusion validation.
(a) Diagram of TP63 fusion indicating the location of the breakpoint. (b) The genomic breakpoint of the TP63 fusion was validated by Sanger sequencing in Mycosis fungoides 64. (c) Diagram of the TNFRSF10A breakpoint. (d) Sanger validation of the genomic breakpoint in TNFRSF10A in Sézary syndrome 69. (e) Diagram of the CTLA4-CD28 fusion event. (f) Sanger validation of the genomic breakpoint in Mycosis fungoides 59. (g) Sanger validation of the cDNA breakpoint in Mycosis fungoides 59. cDNA was generated with quantitative PCR amplification. (h) Jurkat cells with exogenously expressed empty vector (EV), CTLA4 or CTLA4-CD28 fusion construct stained with CTLA4 antibody or isotype control and analyzed using flow cytometry.
Supplementary Figure 4 TNFR2 genomic gain.
(a) Picture of a group of cells with the gain from a punch biopsy of a Mycosis fungoides tumor; this is the same patient as shown in Figure 3a. (b) Graph of the ratio of TNFRSF1B to the control region in ddPCR and FISH analyses in HH cells.
Supplementary Figure 5 Patient survival stratified by genomic and clinical profile.
(a) Patients with LCT histology (n = 26) are significantly more mutated in the gene set that was resequenced than patients without LCT histology (n = 47) (Student’s t test, P = 0.0003). (b) Kaplan-Meier survival curve indicating survival of patients from the time of biopsy to death or current survival (marked as censored points with slashes). Patients with an LCT histology (n = 26) have significantly worse survival than patients without LCT histology (n = 47) (P = 0.0232, Gehan-Breslow-Wilcoxon test; P = 0.0472, Mantel-Cox test). (c) Kaplan-Meier survival curve indicating the survival of patients from the time of biopsy to death or current survival (marked as censored points with slashes). Patients with stage IV Mycosis fungoides or Sézary syndrome disease (n = 39) have significantly worse survival than those with stage IIb + III (n = 21) or stage I (n = 13) disease (P = 0.0199, Gehan-Breslow-Wilcoxon test; P = 0.0583, Mantel-Cox test). (d) Kaplan-Meier survival curve indicating survival of patients from the time of biopsy to death or current survival (marked as censored points with slashes). Patients with a lesion in T cell survival and proliferation pathways (n = 28) have poorer survival than those without an alteration in this pathway (n = 45) (P = 0.0306, Gehan-Breslow-Wilcoxon test; P = 0.0644, Mantel-Cox test). (e) Kaplan-Meier survival curve comparing survival of patients with more than the median number of mutations (n = 30) or fewer than or equal to the median number of mutations (n = 43) in our targeted resequencing. There is no significant difference between these groups (P = 0.26, Gehan-Breslow-Wilcoxon test; P = 0.24, Mantel-Cox test). (f) Kaplan-Meier survival curve comparing the survival of patients older than the median age (n = 36) or younger than or the same as the median age (n = 37). There is no significant difference between these groups (P = 0.43, Gehan-Breslow-Wilcoxon test; P = 0.38, Mantel-Cox test). (g) Kaplan-Meier survival curve comparing the survival of patients with Mycosis fungoides (n = 41) or Sézary syndrome (n = 32) as original diagnosis. There is no significant difference between these groups (P = 0.26, Gehan-Breslow-Wilcoxon test; P = 0.24, Mantel-Cox test). (h) Kaplan-Meier survival curve indicating the survival of patients from the time of biopsy to death or current survival (marked as censored points with slashes). There is no survival difference between patients with a TP53 mutation (n = 9) and those without one (n = 64) (P = 0.94, Gehan-Breslow-Wilcoxon test; P = 0.79, Mantel-Cox test). (i) Kaplan-Meier survival curve indicating the survival of patients from the time of biopsy to death or current survival (marked as censored points with slashes). Patients with a lesion in the top 5% of mutated genes (including TNFR2 gain) (n = 35) have no significant difference in survival compared to those without lesions in these genes (n = 37) (P = 0.33, Gehan-Breslow-Wilcoxon test; P = 0.47, Mantel-Cox test).
Supplementary Figure 6 TNFR2 expression in primary T cells and Jurkat cells.
(a) Primary T cells stained with TNFR2 antibody or isotype control and analyzed using flow cytometry. (b) Jurkat cells with exogenously expressed empty vector, wild-type TNFR2 or mutant TNFR2 stained with TNFR2 antibody or isotype control and analyzed using flow cytometry.
Supplementary Figure 7 Tumor and cell line effects of TNFR2 p.Thr377Ile and pathway inhibition.
(a) Graph of the tumor volumes of subcutaneous tumors generated with Jurkat cells expressing empty vector (EV), wild-type TNFR2 (WT) or mutant TNFR2 (MT). The TNFR2 MT tumor volumes are significantly larger over time compared to the EV controls (P = 0.002), as are the TNFR2 WT volumes (P = 0.04); n = 10 mice per group, total of 30 mice, mean ± standard error shown. Asterisks are used to indicate the results of ANOVA analysis: *P < 0.05, **P < 0.01, ***P < 0.0001, ****P < 0.00001. (b) Dose-response curves of Jurkat cells expressing EV, TNFR2 WT or TNFR2 MT upon treatment with bortezomib for 24 h; n = 3 biological replicates. Mean ± standard error shown. (c) Control or TNFR2-knockout (KO) HH cells stained with TNFR2 antibody or isotype control and analyzed by flow cytometry. (d) Dose-response curves of control or TNFR2 KO HH cells upon treatment with bortezomib for 24 h; n = 3 biological replicates. Mean ± standard error shown. (e) Cell viability of CRISPR control and TNFR2 KO cells over 4 d. There was no significant difference observed. (f) Dose-response curves of the HH CTCL cell line and the Hut78 CTCL cell line exposed to bortezomib treatment for 24 h; n = 3 biological replicates. Mean ± standard error shown.
Supplementary Figure 8 Original immunoblots shown in Figure 4d.
(a) Original TNFR2 immunoblot. (b) Original TRAF2 immunoblot. (c) Original NF-κB2 immunoblot. (d) Original tubulin immunoblot.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8. (PDF 1577 kb)
Supplementary Tables 1–12.
Supplementary Tables 1–12. (XLS 1683 kb)
Rights and permissions
About this article
Cite this article
Ungewickell, A., Bhaduri, A., Rios, E. et al. Genomic analysis of mycosis fungoides and Sézary syndrome identifies recurrent alterations in TNFR2. Nat Genet 47, 1056–1060 (2015). https://doi.org/10.1038/ng.3370
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3370
This article is cited by
-
Interplay between proteasome inhibitors and NF-κB pathway in leukemia and lymphoma: a comprehensive review on challenges ahead of proteasome inhibitors
Cell Communication and Signaling (2024)
-
Transcriptomic and proteomic analysis of tumor suppressive effects of GZ17-6.02 against mycosis fungoides
Scientific Reports (2024)
-
RGS5 augments astrocyte activation and facilitates neuroinflammation via TNF signaling
Journal of Neuroinflammation (2023)
-
TNFR2 antagonist and agonist: a potential therapeutics in cancer immunotherapy
Medical Oncology (2022)
-
Neues zu Pathogenese und molekularem Verständnis bei kutanen T-Zell-Lymphomen
Die Dermatologie (2022)