Abstract
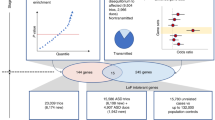
To assess the relative impact of inherited and de novo variants on autism risk, we generated a comprehensive set of exonic single-nucleotide variants (SNVs) and copy number variants (CNVs) from 2,377 families with autism. We find that private, inherited truncating SNVs in conserved genes are enriched in probands (odds ratio = 1.14, P = 0.0002) in comparison to unaffected siblings, an effect involving significant maternal transmission bias to sons. We also observe a bias for inherited CNVs, specifically for small (<100 kb), maternally inherited events (P = 0.01) that are enriched in CHD8 target genes (P = 7.4 × 10−3). Using a logistic regression model, we show that private truncating SNVs and rare, inherited CNVs are statistically independent risk factors for autism, with odds ratios of 1.11 (P = 0.0002) and 1.23 (P = 0.01), respectively. This analysis identifies a second class of candidate genes (for example, RIMS1, CUL7 and LZTR1) where transmitted mutations may create a sensitized background but are unlikely to be completely penetrant.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Autism and Developmental Disabilities Monitoring Network Surveillance Year 2008 Principal Investigators and Centers for Disease Control and Prevention. Prevalence of autism spectrum disorders—Autism and Developmental Disabilities Monitoring Network, 14 sites, United States, 2008. MMWR Surveill. Summ. 61, 1–19 (2012).
Gaugler, T. et al. Most genetic risk for autism resides with common variation. Nat. Genet. 46, 881–885 (2014).
Hallmayer, J. et al. Genetic heritability and shared environmental factors among twin pairs with autism. Arch. Gen. Psychiatry 68, 1095–1102 (2011).
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012).
O'Roak, B.J. et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat. Genet. 43, 585–589 (2011).
O'Roak, B.J. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012).
O'Roak, B.J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012).
Sanders, S.J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
Zhao, X. et al. A unified genetic theory for sporadic and inherited autism. Proc. Natl. Acad. Sci. USA 104, 12831–12836 (2007).
Krumm, N. et al. Transmission disequilibrium of small CNVs in simplex autism. Am. J. Hum. Genet. 93, 595–606 (2013).
Poultney, C.S. et al. Identification of small exonic CNV from whole-exome sequence data and application to autism spectrum disorder. Am. J. Hum. Genet. 93, 607–619 (2013).
Pinto, D. et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am. J. Hum. Genet. 94, 677–694 (2014).
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 466, 368–372 (2010).
Jacquemont, S. et al. A higher mutational burden in females supports a “female protective model” in neurodevelopmental disorders. Am. J. Hum. Genet. 94, 415–425 (2014).
Ronemus, M., Iossifov, I., Levy, D. & Wigler, M. The role of de novo mutations in the genetics of autism spectrum disorders. Nat. Rev. Genet. 15, 133–141 (2014).
Sanders, S.J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863–885 (2011).
Petrovski, S., Wang, Q., Heinzen, E.L., Allen, A.S. & Goldstein, D.B. Genic intolerance to functional variation and the interpretation of personal genomes. PLoS Genet. 9, e1003709 (2013).
Samocha, K.E. et al. A framework for the interpretation of de novo mutation in human disease. Nat. Genet. 46, 944–950 (2014).
He, Z. et al. Rare-variant extensions of the transmission disequilibrium test: application to autism exome sequence data. Am. J. Hum. Genet. 94, 33–46 (2014).
Constantino, J.N. et al. Validation of a brief quantitative measure of autistic traits: comparison of the social responsiveness scale with the autism diagnostic interview-revised. J. Autism Dev. Disord. 33, 427–433 (2003).
Darnell, J.C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Subtil-Rodríguez, A. et al. The chromatin remodeller CHD8 is required for E2F-dependent transcription activation of S-phase genes. Nucleic Acids Res. 42, 2185–2196 (2014).
Bernier, R. et al. Disruptive CHD8 mutations define a subtype of autism early in development. Cell 158, 263–276 (2014).
Taylor, J.W. Simple estimation of population attributable risk from case-control studies. Am. J. Epidemiol. 106, 260 (1977).
Steinman, G. & Mankuta, D. Insulin-like growth factor and the etiology of autism. Med. Hypotheses 80, 475–480 (2013).
Bozdagi, O., Tavassoli, T. & Buxbaum, J.D. Insulin-like growth factor-1 rescues synaptic and motor deficits in a mouse model of autism and developmental delay. Mol. Autism 4, 9 (2013).
Dong, S. et al. De novo insertions and deletions of predominantly paternal origin are associated with autism spectrum disorder. Cell Rep. 9, 16–23 (2014).
Litterman, N. et al. An OBSL1-Cul7Fbxw8 ubiquitin ligase signaling mechanism regulates Golgi morphology and dendrite patterning. PLoS Biol. 9, e1001060 (2011).
Håvik, B. et al. The complement control–related genes CSMD1 and CSMD2 associate to schizophrenia. Biol. Psychiatry 70, 35–42 (2011).
Cukier, H.N. et al. Exome sequencing of extended families with autism reveals genes shared across neurodevelopmental and neuropsychiatric disorders. Mol. Autism 5, 1 (2014).
Kurahashi, H. et al. Isolation and characterization of a novel gene deleted in DiGeorge syndrome. Hum. Mol. Genet. 4, 541–549 (1995).
Kaindl, A.M. et al. Many roads lead to primary autosomal recessive microcephaly. Prog. Neurobiol. 90, 363–383 (2010).
Girirajan, S. et al. A recurrent 16p12.1 microdeletion supports a two-hit model for severe developmental delay. Nat. Genet. 42, 203–209 (2010).
Béna, F. et al. Molecular and clinical characterization of 25 individuals with exonic deletions of NRXN1 and comprehensive review of the literature. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 162B, 388–403 (2013).
Lupski, J.R. Digenic inheritance and Mendelian disease. Nat. Genet. 44, 1291–1292 (2012).
Morita, M. et al. A novel 4EHP-GIGYF2 translational repressor complex is essential for mammalian development. Mol. Cell. Biol. 32, 3585–3593 (2012).
Fischbach, G.D. & Lord, C. The Simons Simplex Collection: a resource for identification of autism genetic risk factors. Neuron 68, 192–195 (2010).
Stessman, H.A., Bernier, R. & Eichler, E.E. A genotype-first approach to defining the subtypes of a complex disease. Cell 156, 872–877 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, B. et al. QPLOT: a quality assessment tool for next generation sequencing data. Biomed. Res. Int. 2013, 865181 (2013).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv http://arxiv.org/abs/1207.3907 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Liu, X., Jian, X. & Boerwinkle, E. dbNSFP v2.0: a database of human non-synonymous SNVs and their functional predictions and annotations. Hum. Mutat. 34, E2393–E2402 (2013).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Krumm, N. et al. Copy number variation detection and genotyping from exome sequence data. Genome Res. 22, 1525–1532 (2012).
Fromer, M. et al. Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth. Am. J. Hum. Genet. 91, 597–607 (2012).
Hach, F. et al. mrsFAST: a cache-oblivious algorithm for short-read mapping. Nat. Methods 7, 576–577 (2010).
Venkatraman, E.S. & Olshen, A.B. A faster circular binary segmentation algorithm for the analysis of array CGH data. Bioinformatics 23, 657–663 (2007).
Krumm, N., O'Roak, B.J., Shendure, J. & Eichler, E.E. A de novo convergence of autism genetics and molecular neuroscience. Trends Neurosci. 37, 95–105 (2014).
Ritchie, M.E., Carvalho, B.S., Hetrick, K.N., Tavare, S. & Irizarry, R.A. R/Bioconductor software for Illumina's Infinium whole-genome genotyping BeadChips. Bioinformatics 25, 2621–2623 (2009).
Scharpf, R.B., Irizarry, R.A., Ritchie, M.E., Carvalho, B. & Ruczinski, I. Using the R package crlmm for genotyping and copy number estimation. J. Stat. Softw. 40, 1–32 (2011).
Acknowledgements
We thank D. Obenshain, D. Hall, B. Koser and S. Novikova for providing support for usage of the Amazon Cloud and for assistance in the deposition of SNV and CNV call sets into the National Database for Autism Research (NDAR). We are grateful to the laboratories of M. Wigler and M. State for providing early access to exome sequencing data as well as access to SNP microarray data. We also thank T. Brown for assistance in editing this manuscript. Funding for this study was provided, in part, by the US National Institutes of Health (1U01MH100233 to E.E.E.), by the National Institute for Mental Health (R01MH101221 to E.E.E. and R01MH100047 to R.B.) and by the Simons Foundation (SFARI 89368 to R.B. and SFARI 137578 to E.E.E.). E.E.E. is an investigator of the Howard Hughes Medical Institute. We are grateful to all of the families at the participating Simons Simplex Collection (SSC) sites, as well as the principal investigators (A. Beaudet, R. Bernier, J. Constantino, E. Cook, E. Fombonne, D. Geschwind, R. Goin-Kochel, E. Hanson, D. Grice, A. Klin, D. Ledbetter, C. Lord, C. Martin, D. Martin, R. Maxim, J. Miles, O. Ousley, K. Pelphrey, B. Peterson, J. Piggot, C. Saulnier, M. State, W. Stone, J. Sutcliffe, C. Walsh, Z. Warren and E. Wijsman). We appreciate obtaining access to phenotypic data on Simons Foundation Autism Research Initiative (SFARI) Base. Approved researchers can obtain the SSC population data set described in this study by applying at https://base.sfari.org/.
Author information
Authors and Affiliations
Contributions
N.K., T.N.T. and E.E.E. designed experiments and wrote and edited the manuscript. N.K. performed sequence data reanalysis and created and analyzed the SNV call set. T.N.T. created and analyzed the CNV call set, analyzed SNP microarray data, performed statistical analyses for SNV and CNV quality control, and examined epidemiological features for the full data set. C.B., L.V., K.M., K.W. and H.A.S. performed validation experiments and sample handling. A.R. and B.P.C. provided additional computational support. Z.-X.H. and S.M.L. performed the TDT tests and statistical analyses. R.B. provided phenotype data and additional SSC variables where needed.
Corresponding author
Ethics declarations
Competing interests
E.E.E. is on the scientific advisory board (SAB) of DNAnexus, Inc., and is a consultant for the Kunming University of Science and Technology (KUST) as part of the 1000 China Talent Program.
Integrated supplementary information
Supplementary Figure 1 PCA for ancestry in SNV data.
(a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype data with colors indicating ethnicity specified in the Simons Foundation Autism Research Initiative (SFARI) Base. (d) PCA of SNV genotype data with colors indicating calculated ethnicity.
Supplementary Figure 2 Allele balance in probands at identified de novo variant sites.
Histogram of proband’s allele balance at 141 sites that we attempted to validate. Green, Sanger validation was successful (the site is a true de novo variant); black outline, the site failed to validate as a de novo variant.
Supplementary Figure 3 Distribution of IQ for autism, PDD and Asperger’s clinical impressions.
Pearson correlation of IQ and all three categories: r2 = 0.18, P < 1 × 10−10.
Supplementary Figure 4 CNV validation rates for CoNIFER and XHMM.
Shown are Venn diagrams with blue indicating events (deletions/duplications), at varying size ranges, called by CoNIFER and yellow indicating events called by XHMM. These are events validated by SNP microarray data.
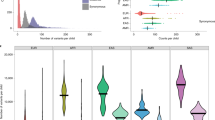
Supplementary Figure 5 Head size distribution for maternal CNV carriers.
The top portion of the diagram shows density distributions of normalized head size per individual for maternally inherited deletions (red) and duplications (blue). Below are jitter plots with each point indicating an individual sample and the respective normalized head size. These are shown for individuals with deletions and duplications carrying a CHD8 target gene and also those who have deletions and duplications that do not contain a CHD8 target gene. A normalized head size of less than or equal to –2 is considered microcephaly and greater than or equal to 2 is macrocephaly.
Supplementary Figure 6 Convergence of de novo and inherited mutations.
Examples of genes with an excess of disruptive mutations in probands include (a) CSMD1, (b) RIMS1 and (c) CUL7. Displayed for each gene is a RefSeq gene model (larger ticks are exons), validated de novo and private LGD mutations (red arrows) and missense (yellow arrows); disruptive CNV deletions (red) and duplications (blue) are compared for proband (p1) and sibling (s1). CSMD1 and RIMS1 show highly brain-specific expression patterns (adapted from the GTEx Consortium online portal: http://gtexportal.org/). Proband IDs with asterisks are part of trios, not quads.
Supplementary Figure 7 Population attributable risk estimates.
Epidemiological features of variant data were assessed for SNVs in 1,786 quad families (parents plus 1 affected proband and 1 unaffected sibling). We subsequently looked at subsets of these families (1,785 families where both proband and sibling sex were known) stratified by (a) proband sex and ultimately by (b) proband-sibling pairing by sex (male proband/male sibling, male proband/female sibling, female proband/male sibling, female proband/female sibling) in families where the sex was known for both the proband and the sibling.
Supplementary Figure 8 Detailed view of known CNVs affecting the CSMD1 locus.
Of note is the depletion of CNV events at the 3′ end of the gene in the Database of Genomic Variants as compared to the 5′ portion of the gene.
Supplementary Figure 9 Sampling results from overlap analysis of genes enriched for de novo and inherited events.
The blue distribution shows the sum of inherited CNVs and SNVs in probands for random 100-gene samplings from Supplementary Table 8 (10,000 iterations). The red vertical line represents the actual (observed) count of CNVs and SNVs in probands for the 100 top enriched genes by de novo events.
Supplementary Figure 10 Detail of de novo and inherited mutations in interacting proteins found in a single family.
A proband with a compound mutation in synaptic receptor–ligand pair; namely, a de novo non-conservative mutation in neuroligin and an inherited two-exon deletion in neurexin. Phenotypic details of this proband include nonverbal IQ = 57, verbal IQ = 46, adaptive IQ = 77, mixed expressive-receptive language disorder, reported autism severity = 8 (of 10), elevated externalizing symptoms, delays in phrase speech, diagnosed heart murmurs, abnormal EEG, elevated BMI (z = 3.69) and macrocephaly (z = 3.08).
Supplementary Figure 12 CNV calling workflow.
1. BAM data were assembled from all previous SSC studies. 2. All reads were realigned using mrsFAST for CoNIFER and using BWA-mem for XHMM. For calling CNVs as part of CoNIFER, a pipeline using DNAcopy and CGHcall was used for segmentation. 3. Post-calling, CNVs were combined into one data set. 4. To validate exome calls, we utilized both available SNP microarray and array comparative genomic hybridization (aCGH) data. For SNP microarray analysis, probe-level copy number estimates were determined by CRLMM, and validation of events was performed by permutation testing. 5. To identify true de novo events, parents were re-genotyped using CRLMM. 6. Post-validation, there were 291 de novo and 6,319 inherited events.
Supplementary Figure 13 RVIS values in the known autism network.
Histogram of RVIS values for all genes in the autism network described in Krumm et al.53. Also indicated is P value based on 1,000,000 permutations of a randomly selected gene set of the same size from the whole genome.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13. (PDF 967 kb)
Supplementary Tables 1–14
Supplementary Tables 1–14. (XLSX 3483 kb)
Supplementary Data Set
All rare inherited variants. (ZIP 11493 kb)
Rights and permissions
About this article
Cite this article
Krumm, N., Turner, T., Baker, C. et al. Excess of rare, inherited truncating mutations in autism. Nat Genet 47, 582–588 (2015). https://doi.org/10.1038/ng.3303
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3303
This article is cited by
-
DGH-GO: dissecting the genetic heterogeneity of complex diseases using gene ontology
BMC Bioinformatics (2023)
-
Comparison of three bioinformatics tools in the detection of ASD candidate variants from whole exome sequencing data
Scientific Reports (2023)
-
Identifying rare genetic variants in 21 highly multiplex autism families: the role of diagnosis and autistic traits
Molecular Psychiatry (2023)
-
Landscape of mSWI/SNF chromatin remodeling complex perturbations in neurodevelopmental disorders
Nature Genetics (2023)
-
Childhood Academic Performance: A Potential Marker of Genetic Liability to Autism
Journal of Autism and Developmental Disorders (2023)