Abstract
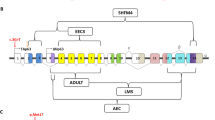
Pierre Robin sequence (PRS) is an important subgroup of cleft palate. We report several lines of evidence for the existence of a 17q24 locus underlying PRS, including linkage analysis results, a clustering of translocation breakpoints 1.06–1.23 Mb upstream of SOX9, and microdeletions both ∼1.5 Mb centromeric and ∼1.5 Mb telomeric of SOX9. We have also identified a heterozygous point mutation in an evolutionarily conserved region of DNA with in vitro and in vivo features of a developmental enhancer. This enhancer is centromeric to the breakpoint cluster and maps within one of the microdeletion regions. The mutation abrogates the in vitro enhancer function and alters binding of the transcription factor MSX1 as compared to the wild-type sequence. In the developing mouse mandible, the 3-Mb region bounded by the microdeletions shows a regionally specific chromatin decompaction in cells expressing Sox9. Some cases of PRS may thus result from developmental misexpression of SOX9 due to disruption of very-long-range cis-regulatory elements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Accessions
GenBank/EMBL/DDBJ
NCBI Reference Sequence
References
Robin, P. A fall of the base of the tongue considered as a new cause of nasopharyngeal respiratory impairment: Pierre Robin sequence, a translation. 1923. Plast. Reconstr. Surg. 93, 1301–1303 (1994).
Etchevers, H.C., Amiel, J. & Lyonnet, S. Molecular bases of human neurocristopathies. Adv. Exp. Med. Biol. 589, 213–234 (2006).
Vintiner, G.M., Temple, I.K., Middleton-Price, H.R., Baraitser, M. & Malcolm, S. Genetic and clinical heterogeneity of Stickler syndrome. Am. J. Med. Genet. 41, 44–48 (1991).
Jamshidi, N. et al. Isolated Robin sequence associated with a balanced t(2;17) chromosomal translocation. J. Med. Genet. 41, e1 (2004).
Jakobsen, L.P. et al. Pierre Robin sequence may be caused by dysregulation of SOX9 and KCNJ2. J. Med. Genet. 44, 381–386 (2007).
Ahituv, N., Prabhakar, S., Poulin, F., Rubin, E.M. & Couronne, O. Mapping cis-regulatory domains in the human genome using multi-species conservation of synteny. Hum. Mol. Genet. 14, 3057–3063 (2005).
Dermitzakis, E.T., Reymond, A. & Antonarakis, S.E. Conserved non-genic sequences—an unexpected feature of mammalian genomes. Nat. Rev. Genet. 6, 151–157 (2005).
Woolfe, A. et al. Highly conserved non-coding sequences are associated with vertebrate development. PLoS Biol. 3, e7 (2005).
Velagaleti, G.V. et al. Position effects due to chromosome breakpoints that map approximately 900 Kb upstream and approximately 1.3 Mb downstream of SOX9 in two patients with campomelic dysplasia. Am. J. Hum. Genet. 76, 652–662 (2005).
Cai, J. et al. Gene expression in pharyngeal arch 1 during human embryonic development. Hum. Mol. Genet. 14, 903–912 (2005).
Satokata, I. & Maas, R. Msx1 deficient mice exhibit cleft palate and abnormalities of craniofacial and tooth development. Nat. Genet. 6, 348–356 (1994).
Cirulli, E.T. & Goldstein, D.B. In vitro assays fail to predict in vivo effects of regulatory polymorphisms. Hum. Mol. Genet. 16, 1931–1939 (2007).
Morey, C., Da Silva, N.R., Perry, P. & Bickmore, W.A. Nuclear reorganisation and chromatin decondensation are conserved, but distinct, mechanisms linked to Hox gene activation. Development 134, 909–919 (2007).
Bagheri-Fam, S., Ferraz, C., Demaille, J., Scherer, G. & Pfeifer, D. Comparative genomics of the SOX9 region in human and Fugu rubripes: conservation of short regulatory sequence elements within large intergenic regions. Genomics 78, 73–82 (2001).
Bagheri-Fam, S. et al. Long-range upstream and downstream enhancers control distinct subsets of the complex spatiotemporal Sox9 expression pattern. Dev. Biol. 291, 382–397 (2006).
Bien-Willner, G.A., Stankiewicz, P. & Lupski, J.R. SOX9cre1, a cis-acting regulatory element located 1.1 Mb upstream of SOX9, mediates its enhancement through the SHH pathway. Hum. Mol. Genet. 16, 1143–1156 (2007).
Suzuki, T., Sakai, D., Osumi, N., Wada, H. & Wakamatsu, Y. Sox genes regulate type 2 collagen expression in avian neural crest cells. Dev. Growth Differ. 48, 477–486 (2006).
Akiyama, H. et al. Interactions between Sox9 and beta-catenin control chondrocyte differentiation. Genes Dev. 18, 1072–1087 (2004).
Bi, W., Deng, J.M., Zhang, Z., Behringer, R.R. & de Crombrugghe, B. Sox9 is required for cartilage formation. Nat. Genet. 22, 85–89 (1999).
Holder-Espinasse, M. et al. Pierre Robin sequence: a series of 117 consecutive cases. J. Pediatr. 139, 588–590 (2001).
Mori-Akiyama, Y., Akiyama, H., Rowitch, D.H. & de Crombrugghe, B. Sox9 is required for determination of the chondrogenic cell lineage in the cranial neural crest. Proc. Natl. Acad. Sci. USA 100, 9360–9365 (2003).
Andelfinger, G. et al. KCNJ2 mutation results in Andersen syndrome with sex-specific cardiac and skeletal muscle phenotypes. Am. J. Hum. Genet. 71, 663–668 (2002).
Wagner, T. et al. Autosomal sex reversal and campomelic dysplasia are caused by mutations in and around the SRY-related gene SOX9. Cell 79, 1111–1120 (1994).
Kleinjan, D.A. & van Heyningen, V. Long-range control of gene expression: emerging mechanisms and disruption in disease. Am. J. Hum. Genet. 76, 8–32 (2005).
Lettice, L.A. et al. A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 12, 1725–1735 (2003).
Hong, C.S. & Saint-Jeannet, J.P. Sox proteins and neural crest development. Semin. Cell Dev. Biol. 16, 694–703 (2005).
McCauley, D.W. & Bronner-Fraser, M. Importance of SoxE in neural crest development and the evolution of the pharynx. Nature 441, 750–752 (2006).
Pheasant, M. & Mattick, J.S. Raising the estimate of functional human sequences. Genome Res. 17, 1245–1253 (2007).
Sharpe, J. et al. Optical projection tomography as a tool for 3D microscopy and gene expression studies. Science 296, 541–545 (2002).
Essafi, A. et al. Direct transcriptional regulation of Bim by FoxO3a mediates STI571-induced apoptosis in Bcr-Abl-expressing cells. Oncogene 24, 2317–2329 (2005).
Acknowledgements
We are grateful to the affected individuals and their families who participated in this study, to the Associations Françaises du Syndrome de Robin, to the Centres de Références Anomalies Cranio-Faciales Rares (AP-HP, Necker and Trousseau hospitals), and to C. Ozilou, G. Staub and G. Guédu-Molina for assistance. We thank T. Attié-Bitach, G. Couly and L. Legeai-Mallet (Necker) and V. van Heyningen and R. Hill (MRC HGU) for useful discussion. This study was underwritten by grants from the Agence Nationale de la Recherche (ERARE grant CraniRare), EUROCRAN FP5, the Fondation pour la Recherche Médicale (FRM), the MRC (UK) and the National Health and Medical Research Council (Australia). S.T. was supported in part by grant NS039818 from the US National Institutes of Health and S.B. by the FRM.
Author information
Authors and Affiliations
Contributions
S.B., J.A.F. and A.P. performed molecular genetics studies. J.A.F., C.T.G. and N.J. performed chromosomal studies. S.B. and J.R. performed the in vitro enhancer activity experiments. S.B., S.T., C.G., M.V. and H.C.E. performed human expression studies. J.R., S.B. and A.E. performed immunoprecipitation experiments. D.-J.K. performed transgenic assays. J.A.F., S.H., P.P. and D.B. performed the in vivo chromatin compaction studies. M.F. did the OPT image analysis. S.B. and H.R.C. performed the comparative genomic analysis. J.A., V.A., C.A., M.H.-E., N.K., M.M.L., A.P., I.K.T., M.V., P.T., M.-P.V., D.R.F. and S.L. recruited individuals and families affected with PRS. S.L., D.R.F., P.G.F. and J.A. contributed to the concept, strategy, study design and project management. S.B., H.C.E., S.T., P.G.F., D.R.F. and S.L. contributed to the writing of the manuscript. All authors discussed the results.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Note, Supplementary Methods and Supplementary Table 1 (PDF 2655 kb)
Rights and permissions
About this article
Cite this article
Benko, S., Fantes, J., Amiel, J. et al. Highly conserved non-coding elements on either side of SOX9 associated with Pierre Robin sequence. Nat Genet 41, 359–364 (2009). https://doi.org/10.1038/ng.329
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.329
This article is cited by
-
SOX9 gene shows association with adolescent idiopathic scoliosis predisposition in Northwest Indians
European Journal of Medical Research (2024)
-
Multi-omics analysis in human retina uncovers ultraconserved cis-regulatory elements at rare eye disease loci
Nature Communications (2024)
-
Shaping faces: genetic and epigenetic control of craniofacial morphogenesis
Nature Reviews Genetics (2023)
-
Precise modulation of transcription factor levels identifies features underlying dosage sensitivity
Nature Genetics (2023)
-
Current challenges in understanding the role of enhancers in disease
Nature Structural & Molecular Biology (2022)