Abstract
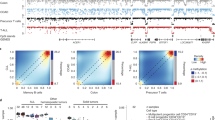
We analyzed the DNA methylome of ten subpopulations spanning the entire B cell differentiation program by whole-genome bisulfite sequencing and high-density microarrays. We observed that non-CpG methylation disappeared upon B cell commitment, whereas CpG methylation changed extensively during B cell maturation, showing an accumulative pattern and affecting around 30% of all measured CpG sites. Early differentiation stages mainly displayed enhancer demethylation, which was associated with upregulation of key B cell transcription factors and affected multiple genes involved in B cell biology. Late differentiation stages, in contrast, showed extensive demethylation of heterochromatin and methylation gain at Polycomb-repressed areas, and genes with apparent functional impact in B cells were not affected. This signature, which has previously been linked to aging and cancer, was particularly widespread in mature cells with an extended lifespan. Comparing B cell neoplasms with their normal counterparts, we determined that they frequently acquire methylation changes in regions already undergoing dynamic methylation during normal B cell differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ziller, M.J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481 (2013).
Bernstein, B.E., Meissner, A. & Lander, E.S. The mammalian epigenome. Cell 128, 669–681 (2007).
Smith, Z.D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Bergman, Y. & Cedar, H. DNA methylation dynamics in health and disease. Nat. Struct. Mol. Biol. 20, 274–281 (2013).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Hovestadt, V. et al. Decoding the regulatory landscape of medulloblastoma using DNA methylation sequencing. Nature 510, 537–541 (2014).
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905 (2013).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Gifford, C.A. et al. Transcriptional and epigenetic dynamics during specification of human embryonic stem cells. Cell 153, 1149–1163 (2013).
Kulis, M. et al. Epigenomic analysis detects widespread gene-body DNA hypomethylation in chronic lymphocytic leukemia. Nat. Genet. 44, 1236–1242 (2012).
Berman, B.P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina–associated domains. Nat. Genet. 44, 40–46 (2012).
Habibi, E. et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13, 360–369 (2013).
Hansen, K.D. et al. Increased methylation variation in epigenetic domains across cancer types. Nat. Genet. 43, 768–775 (2011).
Li, Y. et al. The DNA methylome of human peripheral blood mononuclear cells. PLoS Biol. 8, e1000533 (2010).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Varley, K.E. et al. Dynamic DNA methylation across diverse human cell lines and tissues. Genome Res. 23, 555–567 (2013).
Lee, S.T. et al. A global DNA methylation and gene expression analysis of early human B-cell development reveals a demethylation signature and transcription factor network. Nucleic Acids Res. 40, 11339–11351 (2012).
Lai, A.Y. et al. DNA methylation profiling in human B cells reveals immune regulatory elements and epigenetic plasticity at Alu elements during B cell activation. Genome Res. 23, 2030–2041 (2013).
Shaknovich, R. et al. DNA methyltransferase 1 and DNA methylation patterning contribute to germinal center B-cell differentiation. Blood 118, 3559–3569 (2011).
Matthias, P. & Rolink, A.G. Transcriptional networks in developing and mature B cells. Nat. Rev. Immunol. 5, 497–508 (2005).
Kurosaki, T., Shinohara, H. & Baba, Y. B cell signaling and fate decision. Annu. Rev. Immunol. 28, 21–55 (2010).
Manz, R.A., Thiel, A. & Radbruch, A. Lifetime of plasma cells in the bone marrow. Nature 388, 133–134 (1997).
Slifka, M.K., Antia, R., Whitmire, J.K. & Ahmed, R. Humoral immunity due to long-lived plasma cells. Immunity 8, 363–372 (1998).
Hon, G.C. et al. Epigenetic memory at embryonic enhancers identified in DNA methylation maps from adult mouse tissues. Nat. Genet. 45, 1198–1206 (2013).
Bibikova, M. et al. High density DNA methylation array with single CpG site resolution. Genomics 98, 288–295 (2011).
Ziller, M.J. et al. Genomic distribution and inter-sample variation of non-CpG methylation across human cell types. PLoS Genet. 7, e1002389 (2011).
Arand, J. et al. In vivo control of CpG and non-CpG DNA methylation by DNA methyltransferases. PLoS Genet. 8, e1002750 (2012).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Lefebvre, C. et al. A human B-cell interactome identifies MYB and FOXM1 as master regulators of proliferation in germinal centers. Mol. Syst. Biol. 6, 377 (2010).
Tooze, R.M. A replicative self-renewal model for long-lived plasma cells: questioning irreversible cell cycle exit. Front. Immunol. 4, 460 (2013).
Viré, E. et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439, 871–874 (2006).
Cedar, H. & Bergman, Y. Linking DNA methylation and histone modification: patterns and paradigms. Nat. Rev. Genet. 10, 295–304 (2009).
Brinkman, A.B. et al. Sequential ChIP–bisulfite sequencing enables direct genome-scale investigation of chromatin and DNA methylation cross-talk. Genome Res. 22, 1128–1138 (2012).
Statham, A.L. et al. Bisulfite sequencing of chromatin immunoprecipitated DNA (BisChIP-seq) directly informs methylation status of histone-modified DNA. Genome Res. 22, 1120–1127 (2012).
Martín-Subero, J.I. et al. New insights into the biology and origin of mature aggressive B-cell lymphomas by combined epigenomic, genomic, and transcriptional profiling. Blood 113, 2488–2497 (2009).
Ohm, J.E. et al. A stem cell–like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nat. Genet. 39, 237–242 (2007).
Schlesinger, Y. et al. Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nat. Genet. 39, 232–236 (2007).
Widschwendter, M. et al. Epigenetic stem cell signature in cancer. Nat. Genet. 39, 157–158 (2007).
Busche, S. et al. Integration of high-resolution methylome and transcriptome analyses to dissect epigenomic changes in childhood acute lymphoblastic leukemia. Cancer Res. 73, 4323–4336 (2013).
Agirre, X. et al. Whole-epigenome analysis in multiple myeloma reveals DNA hypermethylation of B cell–specific enhancers. Genome Res. 25, 478–487 (2015).
López-Otín, C., Blasco, M.A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Horvath, S. et al. Aging effects on DNA methylation modules in human brain and blood tissue. Genome Biol. 13, R97 (2012).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367 (2013).
Heyn, H. et al. Distinct DNA methylomes of newborns and centenarians. Proc. Natl. Acad. Sci. USA 109, 10522–10527 (2012).
Maegawa, S. et al. Widespread and tissue specific age-related DNA methylation changes in mice. Genome Res. 20, 332–340 (2010).
Jaffe, A.E. & Irizarry, R.A. Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biol. 15, R31 (2014).
Deaton, A.M. et al. Cell type–specific DNA methylation at intragenic CpG islands in the immune system. Genome Res. 21, 1074–1086 (2011).
Hodges, E. et al. Directional DNA methylation changes and complex intermediate states accompany lineage specificity in the adult hematopoietic compartment. Mol. Cell 44, 17–28 (2011).
Jeong, M. et al. Large conserved domains of low DNA methylation maintained by Dnmt3a. Nat. Genet. 46, 17–23 (2014).
Tsankov, A.M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Stadler, M.B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Schmidl, C. et al. Lineage-specific DNA methylation in T cells correlates with histone methylation and enhancer activity. Genome Res. 19, 1165–1174 (2009).
Tagoh, H. et al. Dynamic reorganization of chromatin structure and selective DNA demethylation prior to stable enhancer complex formation during differentiation of primary hematopoietic cells in vitro. Blood 103, 2950–2955 (2004).
Wiench, M. et al. DNA methylation status predicts cell type–specific enhancer activity. EMBO J. 30, 3028–3039 (2011).
Taberlay, P.C., Statham, A.L., Kelly, T.K., Clark, S.J. & Jones, P.A. Reconfiguration of nucleosome-depleted regions at distal regulatory elements accompanies DNA methylation of enhancers and insulators in cancer. Genome Res. 24, 1421–1432 (2014).
Schlesinger, F., Smith, A.D., Gingeras, T.R., Hannon, G.J. & Hodges, E. De novo DNA demethylation and noncoding transcription define active intergenic regulatory elements. Genome Res. 23, 1601–1614 (2013).
Aran, D., Sabato, S. & Hellman, A. DNA methylation of distal regulatory sites characterizes dysregulation of cancer genes. Genome Biol. 14, R21 (2013).
Aran, D., Toperoff, G., Rosenberg, M. & Hellman, A. Replication timing–related and gene body–specific methylation of active human genes. Hum. Mol. Genet. 20, 670–680 (2011).
Teschendorff, A.E. et al. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 20, 440–446 (2010).
Cruickshanks, H.A. et al. Senescent cells harbour features of the cancer epigenome. Nat. Cell Biol. 15, 1495–1506 (2013).
Rakyan, V.K. et al. Human aging-associated DNA hypermethylation occurs preferentially at bivalent chromatin domains. Genome Res. 20, 434–439 (2010).
Marco-Sola, S., Sammeth, M., Guigo, R. & Ribeca, P. The GEM mapper: fast, accurate and versatile alignment by filtration. Nat. Methods 9, 1185–1188 (2012).
Bibikova, M. et al. Genome-wide DNA methylation profiling using Infinium® assay. Epigenomics 1, 177–200 (2009).
Aryee, M.J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Maksimovic, J., Gordon, L. & Oshlack, A. SWAN: subset-quantile within array normalization for Illumina Infinium HumanMethylation450 BeadChips. Genome Biol. 13, R44 (2012).
Bastian, M., Heymann, S. & Jacomy, M. in Proc. 3rd Int. AAAI Conf. Weblogs and Social Media 361–362 (2009).
Blondel, V.D., Guillaume, J.L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory Exp. 10, P10008 (2008).
Mammana, A., Vingron, M. & Chung, H.R. Inferring nucleosome positions with their histone mark annotation from ChIP data. Bioinformatics 29, 2547–2554 (2013).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Gautier, L., Cope, L., Bolstad, B.M. & Irizarry, R.A. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Fang, J. et al. Epigenetic changes mediated by microRNA miR29 activate cyclooxygenase 2 and λ-1 interferon production during viral infection. J. Virol. 86, 1010–1020 (2012).
Acknowledgements
We thank C. López-Otín for critical reading of this manuscript, M. Dabad Castellà for his assistance with the WGBS data analysis and M.A. Peinado (Institute of Predictive and Personalized Medicine of Cancer, Barcelona) for providing RNA from HCT116 DKO cells. This work was funded by the European Union's Seventh Framework Programme through the Blueprint Consortium (grant agreement 282510) and the Spanish Ministry of Economy and Competitivity (MINECO; project SAF2009-08663). Methylation microarrays were outsourced to the Spanish Centro Nacional de Genotipado (CEGEN-ISCIII). We are indebted to the Genomics core facility of the Institut d'Investigacions Biomédiques August Pi i Sunyer (IDIBAPS) for technical help. This work was partially developed at the Centro Esther Koplowitz (CEK; Barcelona, Spain). M.K. is supported by Agéncia de Gestió d'Ajuts Universitaris i de Recerca (AGAUR; Generalitat de Catalunya), E.C. is an Academia Researcher of the Institució Catalana de Recerca i Estudis Avançats and J.I.M.-S. is a Ramón y Cajal researcher of MINECO.
Author information
Authors and Affiliations
Contributions
M.K., A.C.Q., N.R., M.P., X.A., F.P., D.A., B.P., G. Caron, T.F., M.O.M., M.E.F., S.-T.L. and J.L.W. provided samples from healthy donors and/or purified B cell subpopulations. M.K., A.C.Q., G. Castellano, R.B. and G. Clot analyzed DNA methylation and gene expression arrays. L.A., J.B. and M.G. performed WGBS library preparation and sequencing. A.M., S.H., R.P.S., E.R., A.E. and M.D.-F. processed and analyzed WGBS data. M.K., N.V.-D. and R.V.-B. performed validation experiments. M.K., G. Castellano. S.E., V.P., D. Rico and A.V. functionally characterized dynamically methylated genes. D. Richardson, L.C., A.D. and P.F. were in charge of data management. I.G.G. and H.G.S. coordinated sequencing efforts and performed primary data analysis. H.G.S., R.S., R.K. and E.C. participated in the study design and data interpretation. J.I.M.-S. conceived the study. J.I.M.-S. led the experiments and wrote the manuscript with predominant assistance from M.K. and R.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–26 and Supplementary Tables 1–3. (PDF 2463 kb)
Supplementary Data Set 1
Annotation of CpGs belonging to each of the methylation modules detected by microarrays. (XLSX 7664 kb)
Supplementary Data Set 2
Enrichment analysis of transcription factor binding sites in the 20 DNA methylation modules. (XLSX 144 kb)
Supplementary Data Set 3
Enrichment analysis of transcription factor binding sites using differentially methylated CpGs identified by WGBS. (XLSX 61 kb)
Supplementary Data Set 4
Gene Ontology analysis of dynamic CpGs belonging to 20 main modules. (XLSX 46 kb)
Supplementary Data Set 5
Differential methylation analysis of B cell neoplasms and their normal cell counterparts. (XLSX 7982 kb)
Rights and permissions
About this article
Cite this article
Kulis, M., Merkel, A., Heath, S. et al. Whole-genome fingerprint of the DNA methylome during human B cell differentiation. Nat Genet 47, 746–756 (2015). https://doi.org/10.1038/ng.3291
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3291
This article is cited by
-
SOX11/PRDX2 axis modulates redox homeostasis and chemoresistance in aggressive mantle cell lymphoma
Scientific Reports (2024)
-
DNA methylation analysis explores the molecular basis of plasma cell-free DNA fragmentation
Nature Communications (2023)
-
Identification of intergenerational epigenetic inheritance by whole genome DNA methylation analysis in trios
Scientific Reports (2023)
-
Molecular characterization of Richter syndrome identifies de novo diffuse large B-cell lymphomas with poor prognosis
Nature Communications (2023)
-
Comparative analysis of the DNA methylation landscape in CD4, CD8, and B memory lineages
Clinical Epigenetics (2022)