Abstract
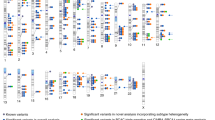
Genome-wide association studies (GWAS) and large-scale replication studies have identified common variants in 79 loci associated with breast cancer, explaining ∼14% of the familial risk of the disease. To identify new susceptibility loci, we performed a meta-analysis of 11 GWAS, comprising 15,748 breast cancer cases and 18,084 controls together with 46,785 cases and 42,892 controls from 41 studies genotyped on a 211,155-marker custom array (iCOGS). Analyses were restricted to women of European ancestry. We generated genotypes for more than 11 million SNPs by imputation using the 1000 Genomes Project reference panel, and we identified 15 new loci associated with breast cancer at P < 5 × 10−8. Combining association analysis with ChIP-seq chromatin binding data in mammary cell lines and ChIA-PET chromatin interaction data from ENCODE, we identified likely target genes in two regions: SETBP1 at 18q12.3 and RNF115 and PDZK1 at 1q21.1. One association appears to be driven by an amino acid substitution encoded in EXO1.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kamangar, F., Dores, G.M. & Anderson, W.F. Patterns of cancer incidence, mortality, and prevalence across five continents: defining priorities to reduce cancer disparities in different geographic regions of the world. J. Clin. Oncol. 24, 2137–2150 (2006).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Hunter, D.J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat. Genet. 39, 870–874 (2007).
Stacey, S.N. et al. Common variants on chromosome 5p12 confer susceptibility to estrogen receptor–positive breast cancer. Nat. Genet. 40, 703–706 (2008).
Stacey, S.N. et al. Common variants on chromosomes 2q35 and 16q12 confer susceptibility to estrogen receptor–positive breast cancer. Nat. Genet. 39, 865–869 (2007).
Ahmed, S. et al. Newly discovered breast cancer susceptibility loci on 3p24 and 17q23.2. Nat. Genet. 41, 585–590 (2009).
Zheng, W. et al. Genome-wide association study identifies a new breast cancer susceptibility locus at 6q25.1. Nat. Genet. 41, 324–328 (2009).
Thomas, G. et al. A multistage genome-wide association study in breast cancer identifies two new risk alleles at 1p11.2 and 14q24.1 (RAD51L1). Nat. Genet. 41, 579–584 (2009).
Turnbull, C. et al. Genome-wide association study identifies five new breast cancer susceptibility loci. Nat. Genet. 42, 504–507 (2010).
Antoniou, A.C. et al. A locus on 19p13 modifies risk of breast cancer in BRCA1 mutation carriers and is associated with hormone receptor–negative breast cancer in the general population. Nat. Genet. 42, 885–892 (2010).
Fletcher, O. et al. Novel breast cancer susceptibility locus at 9q31.2: results of a genome-wide association study. J. Natl. Cancer Inst. 103, 425–435 (2011).
Haiman, C.A. et al. A common variant at the TERT-CLPTM1L locus is associated with estrogen receptor–negative breast cancer. Nat. Genet. 43, 1210–1214 (2011).
Ghoussaini, M. et al. Genome-wide association analysis identifies three new breast cancer susceptibility loci. Nat. Genet. 44, 312–318 (2012).
Siddiq, A. et al. A meta-analysis of genome-wide association studies of breast cancer identifies two novel susceptibility loci at 6q14 and 20q11. Hum. Mol. Genet. 21, 5373–5384 (2012).
Long, J. et al. Genome-wide association study in east Asians identifies novel susceptibility loci for breast cancer. PLoS Genet. 8, e1002532 (2012).
Michailidou, K. et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat. Genet. 45, 353–361 (2013).
Garcia-Closas, M. et al. Genome-wide association studies identify four ER negative–specific breast cancer risk loci. Nat. Genet. 45, 392–398 (2013).
Bojesen, S.E. et al. Multiple independent variants at the TERT locus are associated with telomere length and risks of breast and ovarian cancer. Nat. Genet. 45, 371–384 (2013).
Milne, R.L. et al. Common non-synonymous SNPs associated with breast cancer susceptibility: findings from the Breast Cancer Association Consortium. Hum. Mol. Genet. 23, 6096–6111 (2014).
Cai, Q. et al. Genome-wide association analysis in East Asians identifies breast cancer susceptibility loci at 1q32.1, 5q14.3 and 15q26.1. Nat. Genet. 46, 886–890 (2014).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G.R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Cox, A. et al. A common coding variant in CASP8 is associated with breast cancer risk. Nat. Genet. 39, 352–358 (2007).
Lin, W.Y. et al. Identification and characterisation of novel associations in the CASP8/ALS2CR12 region on chromosome 2 with breast cancer risk. Hum. Mol. Genet. 24, 285–298 (2015).
Meijers-Heijboer, H. et al. Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat. Genet. 31, 55–59 (2002).
CHEK2 Breast Cancer Case-Control Consortium. CHEK2*1100delC and susceptibility to breast cancer: a collaborative analysis involving 10,860 breast cancer cases and 9,065 controls from 10 studies. Am. J. Hum. Genet. 74, 1175–1182 (2004).
Gudbjartsson, D.F. et al. Many sequence variants affecting diversity of adult human height. Nat. Genet. 40, 609–615 (2008).
Kuchenbaecker, K.B. et al. Identification of six new susceptibility loci for invasive epithelial ovarian cancer. Nat. Genet. 47, 164–171 (2015).
Udler, M.S., Tyrer, J. & Easton, D.F. Evaluating the power to discriminate between highly correlated SNPs in genetic association studies. Genet. Epidemiol. 34, 463–468 (2010).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Corradin, O. et al. Combinatorial effects of multiple enhancer variants in linkage disequilibrium dictate levels of gene expression to confer susceptibility to common traits. Genome Res. 24, 1–13 (2014).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Wang, Z. et al. RNF115/BCA2 E3 ubiquitin ligase promotes breast cancer cell proliferation through targeting p21Waf1/Cip1 for ubiquitin-mediated degradation. Neoplasia 15, 1028–1035 (2013).
Kim, H. et al. PDZK1 is a novel factor in breast cancer that is indirectly regulated by estrogen through IGF-1R and promotes estrogen-mediated growth. Mol. Med. 19, 253–262 (2013).
Ahsan, H. et al. A genome-wide association study of early-onset breast cancer identifies PFKM as a novel breast cancer gene and supports a common genetic spectrum for breast cancer at any age. Cancer Epidemiol. Biomarkers Prev. 23, 658–669 (2014).
Stevens, K.N. et al. 19p13.1 is a triple-negative-specific breast cancer susceptibility locus. Cancer Res. 72, 1795–1803 (2012).
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Acknowledgements
The authors thank all the individuals who took part in these studies and all the researchers, clinicians, technicians and administrative staff who have enabled this work to be carried out.
BCAC is funded by Cancer Research UK (C1287/A10118, C1287/A12014) and by the European Community's Seventh Framework Programme under grant agreement 223175 (HEALTH-F2-2009-223175) (COGS). Meetings of the BCAC have been funded by the European Union COST programme (BM0606). Genotyping on the iCOGS array was funded by the European Union (HEALTH-F2-2009-223175), Cancer Research UK (C1287/A10710, C8197/A16565), the Canadian Institutes of Health Research (CIHR) for the CIHR Team in Familial Risks of Breast Cancer program and the Ministry of Economic Development, Innovation and Export Trade of Quebec, grant PSR-SIIRI-701. Combination of the GWAS data was supported in part by the US National Institutes of Health (NIH) Cancer Post-Cancer GWAS initiative, grant 1 U19 CA148065-01 (DRIVE, part of the GAME-ON initiative). For a full description of funding and acknowledgments, see the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
K. Michailidou and D.F.E. performed the statistical analysis and drafted the manuscript. D.F.E. conceived and coordinated the synthesis of the iCOGS array and led the BCAC. P.H. coordinated COGS. J. Benitez led the iCOGS genotyping working group. A.G.-N., G.P., M.R.A., N.Á., D.H., J. Benitez, D.V., F.B., D.C.T., J.S., A.M.D., C.L., C. Baynes, S.A., C.S.H. and M.J.M. coordinated genotyping of the iCOGS array. M.G.-C., P.P.D.P.P. and M.K.S. led the BCAC pathology and survival working group. J.C.-C. led the BCAC risk factor working group. A.M.D. and G.C.-T. led the iCOGS quality control working group. J. Beesley, J.D. and M.J.L. provided bioinformatics support. M.K.B. and Q. Wang provided data management support for BCAC. S. Canisius provided analysis of the TCGA expression data. J.L.H., M.C.S., H.T. and C.A. coordinated ABCFS. M.K.S., A.B., S.V. and S. Cornelissen coordinated ABCS. K. Muir, A. Lophatananon, S.S.-B. and P.S. coordinated ACP. P.A.F., A. Hein, M.W.B. and L.H. coordinated BBCC. J.P., I.d.-S.-S., O.F. and L.G. coordinated BBCS. E.J.S., I.T., M.J.K. and N.M. coordinated BIGGS. P.K., D.J.H., S.L., S.M.G., M.M.G., W.R.D., C.A.H., F.S., B.E.H., L.L.M., C.D.B., S.J.C., J.F. and R.N.H. coordinated BPC3. B.B., F.M., H.S. and C. Sohn coordinated BSUCH. N.R. and C. Turnbull coordinated BOCS. P.G., T.T., C. Mulot and M. Sanchez coordinated CECILE. S.E.B., B.G.N., H.F. and S.F.N. coordinated CGPS. A.G.-N., J. Benitez, M.P.Z. and J.I.A.P. coordinated CNIO-BCS. H.A.-C. and S.L.N. coordinated CTS. H. Brenner, A.K.D., V.A. and C. Stegmaier coordinated ESTHER. A. Meindl, R.K.S., C. Sutter and R.Y. coordinated GC-HBOC. H. Brauch, U.H. and T.B. coordinated GENICA. H.N., T.A.M., K. Aittomäki, C. Blomqvist, K. Aaltonen and S.K. coordinated HEBCS. K. Matsuo, H. Ito, H. Iwata and K.T. coordinated HERPACC. T.D. and N.V.B. coordinated HMBCS. A. Lindblom and S. Margolin coordinated KARBAC. A. Mannermaa, V.K., V.-M.K. and J.M.H. coordinated KBCP. G.C.-T. and J. Beesley coordinated kConFab/AOCS. A.H.W., C. Tseng, D.V.D.B. and D.O.S. coordinated LAABC. D.L., P.N., H.W. and E.v.L. coordinated LMBC. J.C.-C., D.F.-J., U.E., S.B. and A.R. coordinated MARIE. P.R., P. Peterlongo, S. Manoukian and L. Bernard coordinated MBCSG. F.J.C., J.E.O., E.H. and C.V. coordinated MCBCS. G.G.G., R.L.M. and C. McLean coordinated MCCS. C.A.H., B.E.H., F.S. and L.L.M. coordinated MEC. J.S., M.S.G., F.L. and M.D. coordinated MTLGEBCS. S.H.T., C.H.Y., N.A.M.T. and G.-H.T. coordinated MYBRCA. V.N.K., G.I.G.A. and S.N. coordinated NBCS. W.Z., S.L.H., M. Shrubsole and J. Long coordinated NBHS. R.W., K.P., A.J.-V. and M.G. coordinated OBCS. I.L.A., J.A.K., G.G. and A.M.M. coordinated OFBCR. P.D., R.A.E.M.T., C. Seynaeve and C.J.V.A. coordinated ORIGO. M.G.-C., J.F., S.J.C. and L. Brinton coordinated PBCS. K.C., H.D., M.E. and J.S.B. coordinated pKARMA. M.J.H., A. Hollestelle, J.W.M.M. and J.M.C. coordinated RBCS. P.H., J. Li, J. Liu and K.H. coordinated SASBAC. X.-O.S., W.L., Y.-T.G. and H.C. coordinated SBCGS. A.C., S.S.C. and M.W.R.R. coordinated SBCS. W.B., L.B.S. and Q.C. coordinated SCCS. M. Shah and B.J.P. coordinated SEARCH. D.K., J.-Y.C., S.K.P. and K.-Y.Y. coordinated SEBCS. M.H., H.M., K.S.C. and C.W.C. coordinated SGBCC. U.H., M.K. and D. Torres coordinated SKKDKFZS. A.J., J. Lubinski, K.J. and T.H. coordinated SZBCS. S. Sangrajrang, V.G., P.B. and J.M. coordinated TBCS. F.J.C., S. Slager, A.E.T., C.B.A. and D.Y. coordinated the TNBCC. C.-Y.S., C.-N.H., P.-E.W. and M.-F.H. coordinated TWBCS. A.J.S.,A.A., N.O. and M.J.S. coordinated UKBGS. H.A., M.G.K., A.S.W., E.M.J., K.E.M., M.D.G., R.M.S., G.U., E.M., D.F.S. and G.C. coordinated EBCG GWAS. Q. Waisfisz, H.M.-H., M.A.A. and R.B.v.d.L. coordinated DFBBCS GWAS. D.F.E., N.R. and C. Turnbull coordinated UK2 GWAS. F.C., D. Trichopoulos, P. Peeters, E.L., M. Sund, K.-T.K., M.J.G., D.P., L.D., J.-M.H. and L.M.M. coordinated EPIC. All authors provided critical review of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–9 and Supplementary Note. (PDF 5459 kb)
Supplementary Table 10
Set of all 522 SNPs correlated with 1 of the 15 lead SNPs and that could not be ruled out as potentially causal (based on a likelihood ratio of 100:1). (XLS 92 kb)
Supplementary Table 11
Associations between the 15 new susceptibility variants and expression of neighboring genes in breast tumors and normal breast tissue, from the TCGA data set. (XLS 78 kb)
Rights and permissions
About this article
Cite this article
Michailidou, K., Beesley, J., Lindstrom, S. et al. Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat Genet 47, 373–380 (2015). https://doi.org/10.1038/ng.3242
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3242
This article is cited by
-
Disulfiram ameliorates STING/MITA-dependent inflammation and autoimmunity by targeting RNF115
Cellular & Molecular Immunology (2024)
-
The impact of circulating protein levels identified by affinity proteomics on short-term, overall breast cancer risk
British Journal of Cancer (2024)
-
Mendelian randomization and transcriptomic analysis reveal an inverse causal relationship between Alzheimer’s disease and cancer
Journal of Translational Medicine (2023)
-
Family history and breast cancer risk for Asian women: a systematic review and meta-analysis
BMC Medicine (2023)
-
Automatic block-wise genotype-phenotype association detection based on hidden Markov model
BMC Bioinformatics (2023)