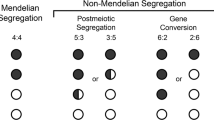
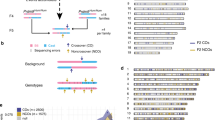
Abstract
The ability to examine all chromatids from a single meiosis in yeast tetrads has been indispensable for defining the mechanisms of homologous recombination initiated by DNA double-strand breaks (DSBs). Using a broadly applicable strategy for the analysis of chromatids from a single meiosis at two recombination hotspots in mouse oocytes and spermatocytes, we demonstrate here the unidirectional transfer of information—gene conversion—in both crossovers and noncrossovers. Whereas gene conversion in crossovers is associated with reciprocal exchange, the unbroken chromatid is not altered in noncrossover gene conversion events, providing strong evidence that noncrossovers arise from a distinct pathway. Gene conversion frequently spares the binding site of the hotspot-specifying protein PRDM9, with the result that erosion of the hotspot is slowed. Thus, mouse tetrad analysis demonstrates how unique aspects of mammalian recombination mechanisms shape hotspot evolutionary dynamics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hunter, N. in Topics in Current Genetics, Molecular Genetics of Recombination Vol. 17/2007 (eds. Aguilera, A. & Rothstein, R.) 381–442 (Springer-Verlag, Heidelberg, Germany, 2007).
Zickler, D. & Kleckner, N. Meiotic chromosomes: integrating structure and function. Annu. Rev. Genet. 33, 603–754 (1999).
de Massy, B. Initiation of meiotic recombination: how and where? Conservation and specificities among eukaryotes. Annu. Rev. Genet. 47, 563–599 (2013).
Winge, O. & Laustsen, O. On two types of spore germination, and on genetic segregations in Saccharomycetes demonstrated through single-spore culture. C. R. Trav. Lab. Carlsberg., Ser. Physiol. 22, 99–117 (1937).
Hurst, D.D., Fogel, S. & Mortimer, R.K. Conversion-associated recombination in yeast. Proc. Natl. Acad. Sci. USA 69, 101–105 (1972).
Szostak, J.W., Orr-Weaver, T.L., Rothstein, R.J. & Stahl, F.W. The double-strand-break repair model for recombination. Cell 33, 25–35 (1983).
Allers, T. & Lichten, M. Differential timing and control of noncrossover and crossover recombination during meiosis. Cell 106, 47–57 (2001).
Börner, G.V., Kleckner, N. & Hunter, N. Crossover/noncrossover differentiation, synaptonemal complex formation, and regulatory surveillance at the leptotene/zygotene transition of meiosis. Cell 117, 29–45 (2004).
Hunter, N. & Kleckner, N. The single-end invasion: an asymmetric intermediate at the double-strand break to double–Holliday junction transition of meiotic recombination. Cell 106, 59–70 (2001).
Martini, E. et al. Genome-wide analysis of heteroduplex DNA in mismatch repair–deficient yeast cells reveals novel properties of meiotic recombination pathways. PLoS Genet. 7, e1002305 (2011).
McMahill, M.S., Sham, C.W. & Bishop, D.K. Synthesis-dependent strand annealing in meiosis. PLoS Biol. 5, e299 (2007).
Arnheim, N., Calabrese, P. & Tiemann-Boege, I. Mammalian meiotic recombination hot spots. Annu. Rev. Genet. 41, 369–399 (2007).
Baudat, F., Imai, Y. & de Massy, B. Meiotic recombination in mammals: localization and regulation. Nat. Rev. Genet. 14, 794–806 (2013).
Paigen, K. & Petkov, P. Mammalian recombination hot spots: properties, control and evolution. Nat. Rev. Genet. 11, 221–233 (2010).
Hou, Y. et al. Genome analyses of single human oocytes. Cell 155, 1492–1506 (2013).
Lu, S. et al. Probing meiotic recombination and aneuploidy of single sperm cells by whole-genome sequencing. Science 338, 1627–1630 (2012).
Wang, J., Fan, H.C., Behr, B. & Quake, S.R. Genome-wide single-cell analysis of recombination activity and de novo mutation rates in human sperm. Cell 150, 402–412 (2012).
Cole, F., Keeney, S. & Jasin, M. Comprehensive, fine-scale dissection of homologous recombination outcomes at a hot spot in mouse meiosis. Mol. Cell 39, 700–710 (2010).
Guillon, H., Baudat, F., Grey, C., Liskay, R.M. & de Massy, B. Crossover and noncrossover pathways in mouse meiosis. Mol. Cell 20, 563–573 (2005).
Guillon, H. & de Massy, B. An initiation site for meiotic crossing-over and gene conversion in the mouse. Nat. Genet. 32, 296–299 (2002).
Jeffreys, A.J. & May, C.A. Intense and highly localized gene conversion activity in human meiotic crossover hot spots. Nat. Genet. 36, 151–156 (2004).
Ng, S.H., Parvanov, E., Petkov, P.M. & Paigen, K. A quantitative assay for crossover and noncrossover molecular events at individual recombination hotspots in both male and female gametes. Genomics 92, 204–209 (2008).
Baudat, F. & de Massy, B. Cis- and trans-acting elements regulate the mouse Psmb9 meiotic recombination hotspot. PLoS Genet. 3, e100 (2007).
Keeney, S. in Recombination and Meiosis (eds. Egel, R. & Lankenau, D.-H.) 81–123 (Springer-Verlag, Berlin and Heidelberg, Germany, 2007).
Cole, F., Keeney, S. & Jasin, M. Evolutionary conservation of meiotic DSB proteins: more than just Spo11. Genes Dev. 24, 1201–1207 (2010).
Kauppi, L., Jeffreys, A.J. & Keeney, S. Where the crossovers are: recombination distributions in mammals. Nat. Rev. Genet. 5, 413–424 (2004).
Baudat, F. et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327, 836–840 (2010).
Brick, K., Smagulova, F., Khil, P., Camerini-Otero, R.D. & Petukhova, G.V. Genetic recombination is directed away from functional genomic elements in mice. Nature 485, 642–645 (2012).
Myers, S. et al. Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science 327, 876–879 (2010).
Parvanov, E.D., Petkov, P.M. & Paigen, K. Prdm9 controls activation of mammalian recombination hotspots. Science 327, 835 (2010).
Billings, T. et al. DNA binding specificities of the long zinc-finger recombination protein PRDM9. Genome Biol. 14, R35 (2013).
Grey, C. et al. Mouse PRDM9 DNA-binding specificity determines sites of histone H3 lysine 4 trimethylation for initiation of meiotic recombination. PLoS Biol. 9, e1001176 (2011).
Jeffreys, A.J. & Neumann, R. Reciprocal crossover asymmetry and meiotic drive in a human recombination hot spot. Nat. Genet. 31, 267–271 (2002).
Buard, J., Barthes, P., Grey, C. & de Massy, B. Distinct histone modifications define initiation and repair of meiotic recombination in the mouse. EMBO J. 28, 2616–2624 (2009).
Bastos, H. et al. Flow cytometric characterization of viable meiotic and postmeiotic cells by Hoechst 33342 in mouse spermatogenesis. Cytometry A 65, 40–49 (2005).
Cole, F. & Jasin, M. Isolation of meiotic recombinants from mouse sperm. Methods Mol. Biol. 745, 251–282 (2011).
Baker, S.M. et al. Involvement of mouse Mlh1 in DNA mismatch repair and meiotic crossing over. Nat. Genet. 13, 336–342 (1996).
Woods, L.M. et al. Chromosomal influence on meiotic spindle assembly: abnormal meiosis I in female Mlh1 mutant mice. J. Cell Biol. 145, 1395–1406 (1999).
Mancera, E., Bourgon, R., Brozzi, A., Huber, W. & Steinmetz, L.M. High-resolution mapping of meiotic crossovers and non-crossovers in yeast. Nature 454, 479–485 (2008).
Bois, P.R. A highly polymorphic meiotic recombination mouse hot spot exhibits incomplete repair. Mol. Cell. Biol. 27, 7053–7062 (2007).
Jeffreys, A.J. & Neumann, R. Factors influencing recombination frequency and distribution in a human meiotic crossover hotspot. Hum. Mol. Genet. 14, 2277–2287 (2005).
Odenthal-Hesse, L., Berg, I.L., Veselis, A., Jeffreys, A.J. & May, C.A. Transmission distortion affecting human noncrossover but not crossover recombination: a hidden source of meiotic drive. PLoS Genet. 10, e1004106 (2014).
Sarbajna, S. et al. A major recombination hotspot in the XqYq pseudoautosomal region gives new insight into processing of human gene conversion events. Hum. Mol. Genet. 21, 2029–2038 (2012).
Yauk, C.L., Bois, P.R. & Jeffreys, A.J. High-resolution sperm typing of meiotic recombination in the mouse MHC Eβ gene. EMBO J. 22, 1389–1397 (2003).
Wu, L. & Hickson, I.D. The Bloom's syndrome helicase suppresses crossing over during homologous recombination. Nature 426, 870–874 (2003).
Chen, S.Y. et al. Global analysis of the meiotic crossover landscape. Dev. Cell 15, 401–415 (2008).
Kauppi, L. et al. Numerical constraints and feedback control of double-strand breaks in mouse meiosis. Genes Dev. 27, 873–886 (2013).
Baker, C.L., Walker, M., Kajita, S., Petkov, P.M. & Paigen, K. PRDM9 binding organizes hotspot nucleosomes and limits Holliday junction migration. Genome Res. 24, 724–732 (2014).
Pan, J. et al. A hierarchical combination of factors shapes the genome-wide topography of yeast meiotic recombination initiation. Cell 144, 719–731 (2011).
Radford, S.J., Sabourin, M.M., McMahan, S. & Sekelsky, J. Meiotic recombination in Drosophila Msh6 mutants yields discontinuous gene conversion tracts. Genetics 176, 53–62 (2007).
Oh, S.D. et al. BLM ortholog, Sgs1, prevents aberrant crossing-over by suppressing formation of multichromatid joint molecules. Cell 130, 259–272 (2007).
Oh, S.D., Lao, J.P., Taylor, A.F., Smith, G.R. & Hunter, N. RecQ helicase, Sgs1, and XPF family endonuclease, Mus81-Mms4, resolve aberrant joint molecules during meiotic recombination. Mol. Cell 31, 324–336 (2008).
Goldfarb, T. & Lichten, M. Frequent and efficient use of the sister chromatid for DNA double-strand break repair during budding yeast meiosis. PLoS Biol. 8, e1000520 (2010).
Boulton, A., Myers, R.S. & Redfield, R.J. The hotspot conversion paradox and the evolution of meiotic recombination. Proc. Natl. Acad. Sci. USA 94, 8058–8063 (1997).
Coop, G. & Myers, S.R. Live hot, die young: transmission distortion in recombination hotspots. PLoS Genet. 3, e35 (2007).
Calabrese, P. A population genetics model with recombination hotspots that are heterogeneous across the population. Proc. Natl. Acad. Sci. USA 104, 4748–4752 (2007).
Pineda-Krch, M. & Redfield, R.J. Persistence and loss of meiotic recombination hotspots. Genetics 169, 2319–2333 (2005).
Anderson, L.K., Reeves, A., Webb, L.M. & Ashley, T. Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151, 1569–1579 (1999).
Cole, F. et al. Homeostatic control of recombination is implemented progressively in mouse meiosis. Nat. Cell Biol. 14, 424–430 (2012).
Shiroishi, T., Sagai, T., Hanzawa, N., Gotoh, H. & Moriwaki, K. Genetic control of sex-dependent meiotic recombination in the major histocompatibility complex of the mouse. EMBO J. 10, 681–686 (1991).
Hanneman, W.H., Schimenti, K.J. & Schimenti, J.C. Molecular analysis of gene conversion in spermatids from transgenic mice. Gene 200, 185–192 (1997).
Jeffreys, A.J., Neumann, R. & Wilson, V. Repeat unit sequence variation in minisatellites: a novel source of DNA polymorphism for studying variation and mutation by single molecule analysis. Cell 60, 473–485 (1990).
Belloc, F. et al. A flow cytometric method using Hoechst 33342 and propidium iodide for simultaneous cell cycle analysis and apoptosis determination in unfixed cells. Cytometry 17, 59–65 (1994).
Barchi, M. et al. ATM promotes the obligate XY crossover and both crossover control and chromosome axis integrity on autosomes. PLoS Genet. 4, e1000076 (2008).
Acknowledgements
We thank members of the Jasin, Keeney and de Massy laboratories, especially J. Lange, E. de Boer, S. Tischfield, S. Yamada and S. Peterson. We also thank P. Hunt (Washington State University) for encouraging the oocyte experiments, A. Gabow (Memorial Sloan Kettering Cancer Center) and Y. Lu (University of Texas MD Anderson Cancer Center) for fixation calculations, and the Réseau des Animaleries de Montpellier (RAM) and D. Haddou for management of mice. This work was supported by R.L. Kirschstein National Research Service Award F32HD51392 (F.C.) and Cancer Prevention and Research Institute of Texas award R1213 (F.C.), the Centre National de la Recherche Scientifique (F.B., C.G. and B.d.M.), the Agence Nationale pour la Recherche (program ANR-09-BLAN-0269-01 to B.d.M.), the Association pour la Recherche contre le Cancer (B.d.M.) and US National Institutes of Health grants R01GM105421 (M.J. and S.K.) and R01HD53855 (S.K. and M.J.). This paper is dedicated to the memory of our colleague Jérôme Buard.
Author information
Authors and Affiliations
Contributions
F.C., F.B., S.K., B.d.M. and M.J. conceived the study, interpreted the data and wrote the manuscript. F.C. and F.B. performed the spermatocyte and oocyte (and associated) recombination experiments, respectively. F.B. and C.G. performed the southwestern blotting. F.C., M.J. and S.K. estimated the fixation rates.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Noncrossovers at the Psmb9 hotspot in B10.A × SGR hybrids determined by analysis of DNA isolated from ovaries and sperm, respectively.
Noncrossovers were identified on the B10.A chromosome at the seven queried polymorphisms (asterisks) in pooled ovaries (a) and sperm (b) as described23. See Supplementary Table 1 for detailed results. Once a noncrossover was identified at the queried polymorphism, the extent of gene conversion was assessed by sequencing the PCR product. For pooled ovaries, DNA was extracted from the ovaries of 2 newborn litters (6 and 5 females, respectively), and an estimated ∼4,739 genomes from oocytes were tested. For sperm, ∼8,981 genomes from a single 4-month-old mouse were tested for the –87 polymorphism and ∼15,739 genomes were tested for the other 6 polymorphisms. The polymorphisms known to affect PRDM9 binding are 70 and, to a lesser extent, 87, which are located 17 bp apart (red arrowhead32). The mean minimal and maximal gene conversion tracts are as indicated. In these particular experiments, the overall frequencies of detected noncrossovers (on the B10.A chromosome; representing >90% of noncrossover events in this hybrid) and crossovers (SGR-B10.A orientation only), respectively, were 0.89% and 0.40% in oocytes and 0.45% and 0.16% in sperm.
Supplementary Figure 2 Noncrossovers from sperm DNA at the A3 hotspot in A/J × DBA/2J hybrids.
(a) Noncrossovers were identified using established methods36 on the DBA/2J chromosome in sperm DNA isolated from the same mice used for tetrad analysis. The mean minimal and maximal gene conversion tracts are as indicated. A nearly identical frequency (Table 1) and similar distribution of noncrossovers were obtained. Co-conversions were observed at a lower frequency in spermatocytes (6 of 111 noncrossovers) than in sperm (13 of 78 noncrossovers; P = 0.0142), although whether this difference reflects a biological distinction between the cell types is unclear. (b) For each polymorphism, the frequency and distribution of noncrossovers on the DBA/2J chromosome was compared between tetrad analysis (top; the same data as shown in Fig. 2e) to sperm (bottom). Polymorphisms analyzed with oligonucleotide probes new to this study are indicated by dots.
Supplementary Figure 3 Single-chromatid analysis cannot distinguish recombination mechanisms because gene conversion tracts are not identified.
(a) Examples of two possible crossovers initiated by a DSB on the frequently cleaved chromosome. In the example on the left, the crossover breakpoints flank the site of the DSB such that the gene conversion tract encompasses the center of the hotspot. In the example on the right, both crossover breakpoints occur to one side of the DSB, such that the conversion tract does not overlap the center of the hotspot. (b) Example of a crossover initiated by a DSB on the infrequently cleaved chromosome in which the crossover breakpoints flank the site of the DSB. Note that the breakpoint on the lower chromosome (breakpoint 3b) is similar to that from the off-center conversion in a (breakpoint 2b). (c) Distribution of crossover breakpoints at the A3 hotspot from A/J × DBA/2J mice previously determined from sperm analysis18. The A/J to DBA/2J (top) and DBA/2J to A/J (bottom) breakpoints are shifted relative to each other (asymmetric), consistent with substantially more frequent initiation on the DBA/2J chromosome. Vertical lines represent the midpoint for each orientation. The intervals in which the crossover breakpoints from a and b occur are indicated. (d) Southwestern analysis of the PRDM9wm7 protein for the A3 hotspot. The horizontal bars represent the positions of the eight overlapping ∼250-bp DNA probes generated for the A/J and DBA/2J genotype. Representative southwesterns for each probe against His-tagged PRDM9wm7 are shown along with a loading control (probed with antibody to His) on the far right.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Tables 1–3. (PDF 1801 kb)
Source data
Rights and permissions
About this article
Cite this article
Cole, F., Baudat, F., Grey, C. et al. Mouse tetrad analysis provides insights into recombination mechanisms and hotspot evolutionary dynamics. Nat Genet 46, 1072–1080 (2014). https://doi.org/10.1038/ng.3068
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3068
This article is cited by
-
Evolution of the recombination regulator PRDM9 in minke whales
BMC Genomics (2022)
-
Gene conversion: a non-Mendelian process integral to meiotic recombination
Heredity (2022)
-
Stage-resolved Hi-C analyses reveal meiotic chromosome organizational features influencing homolog alignment
Nature Communications (2021)
-
ATM and PRDM9 regulate SPO11-bound recombination intermediates during meiosis
Nature Communications (2020)
-
The evolutionarily conserved piRNA-producing locus pi6 is required for male mouse fertility
Nature Genetics (2020)