Abstract
Polycomb/Trithorax response elements (PRE/TREs) can switch their function reversibly between silencing and activation by mechanisms that are poorly understood. Here we show that a switch in forward and reverse noncoding transcription from the Drosophila melanogaster vestigial (vg) PRE/TRE switches the status of the element between silencing (induced by the forward strand) and activation (induced by the reverse strand). In vitro, both noncoding RNAs inhibit PRC2 histone methyltransferase activity, but, in vivo, only the reverse strand binds PRC2. Overexpression of the reverse strand evicts PRC2 from chromatin and inhibits its enzymatic activity. We propose that the interaction of RNAs with PRC2 is differentially regulated in vivo, allowing regulated inhibition of local PRC2 activity. Genome-wide analysis shows that strand switching of noncoding RNAs occurs at several hundred Polycomb-binding sites in fly and vertebrate genomes. This work identifies a previously unreported and potentially widespread class of PRE/TREs that switch function by switching the direction of noncoding RNA transcription.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Di Croce, L. & Helin, K. Transcriptional regulation by Polycomb group proteins. Nat. Struct. Mol. Biol. 20, 1147–1155 (2013).
Steffen, P.A. & Ringrose, L. What are memories made of? How Polycomb and Trithorax proteins mediate epigenetic memory. Nat. Rev. Mol. Cell Biol. 15, 340–356 (2014).
Müller, J. & Kassis, J.A. Polycomb response elements and targeting of Polycomb group proteins in Drosophila. Curr. Opin. Genet. Dev. 16, 476–484 (2006).
Hekimoglu, B. & Ringrose, L. Non-coding RNAs in polycomb/trithorax regulation. RNA Biol. 6, 129–137 (2009).
Spitale, R.C., Tsai, M.C. & Chang, H.Y. RNA templating the epigenome: long noncoding RNAs as molecular scaffolds. Epigenetics 6, 539–543 (2011).
Brockdorff, N. Noncoding RNA and Polycomb recruitment. RNA 19, 429–442 (2013).
Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Zhao, J., Sun, B.K., Erwin, J.A., Song, J.J. & Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, 750–756 (2008).
Kanhere, A. et al. Short RNAs are transcribed from repressed polycomb target genes and interact with polycomb repressive complex-2. Mol. Cell 38, 675–688 (2010).
Zhao, J. et al. Genome-wide identification of polycomb-associated RNAs by RIP-seq. Mol. Cell 40, 939–953 (2010).
Wang, K.C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Guil, S. et al. Intronic RNAs mediate EZH2 regulation of epigenetic targets. Nat. Struct. Mol. Biol. 19, 664–670 (2012).
Davidovich, C., Zheng, L., Goodrich, K.J. & Cech, T.R. Promiscuous RNA binding by Polycomb repressive complex 2. Nat. Struct. Mol. Biol. 20, 1250–1257 (2013).
Cifuentes-Rojas, C., Hernandez, A.J., Sarma, K. & Lee, J.T. Regulatory interactions between RNA and Polycomb repressive complex 2. Mol. Cell 10.1016/j.molcel.2014.05.009 (29 May 2014).
Cohen, B., Simcox, A.A. & Cohen, S.M. Allocation of the thoracic imaginal primordia in the Drosophila embryo. Development 117, 597–608 (1993).
Williams, J.A., Bell, J.B. & Carroll, S.B. Control of Drosophila wing and haltere development by the nuclear vestigial gene product. Genes Dev. 5, 2481–2495 (1991).
Deng, H., Hughes, S.C., Bell, J.B. & Simmonds, A.J. Alternative requirements for Vestigial, Scalloped, and Dmef2 during muscle differentiation in Drosophila melanogaster. Mol. Biol. Cell 20, 256–269 (2009).
Bernard, F. et al. Control of apterous by vestigial drives indirect flight muscle development in Drosophila. Dev. Biol. 260, 391–403 (2003).
Guss, K.A., Mistry, H. & Skeath, J.B. Vestigial expression in the Drosophila embryonic central nervous system. Dev. Dyn. 237, 2483–2489 (2008).
Enderle, D. et al. Polycomb preferentially targets stalled promoters of coding and noncoding transcripts. Genome Res. 21, 216–226 (2011).
Kharchenko, P.V. et al. Comprehensive analysis of the chromatin landscape in Drosophila melanogaster. Nature 471, 480–485 (2011).
Lee, N., Maurange, C., Ringrose, L. & Paro, R. Suppression of Polycomb group proteins by JNK signalling induces transdetermination in Drosophila imaginal discs. Nature 438, 234–237 (2005).
Pérez, L. et al. Enhancer-PRE communication contributes to the expansion of gene expression domains in proliferating primordia. Development 138, 3125–3134 (2011).
Oktaba, K. et al. Dynamic regulation by polycomb group protein complexes controls pattern formation and the cell cycle in Drosophila. Dev. Cell 15, 877–889 (2008).
Schwartz, Y.B. et al. Genome-wide analysis of Polycomb targets in Drosophila melanogaster. Nat. Genet. 38, 700–705 (2006).
Okulski, H., Druck, B., Bhalerao, S. & Ringrose, L. Quantitative analysis of polycomb response elements (PREs) at identical genomic locations distinguishes contributions of PRE sequence and genomic environment. Epigenetics Chromatin 4, 4 (2011).
Bate, M. The embryonic development of larval muscles in Drosophila. Development 110, 791–804 (1990).
DeVido, S.K., Kwon, D., Brown, J.L. & Kassis, J.A. The role of Polycomb-group response elements in regulation of engrailed transcription in Drosophila. Development 135, 669–676 (2008).
Bantignies, F. et al. Polycomb-dependent regulatory contacts between distant Hox loci in Drosophila. Cell 144, 214–226 (2011).
Kaneko, S., Son, J., Shen, S.S., Reinberg, D. & Bonasio, R. PRC2 binds active promoters and contacts nascent RNAs in embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1258–1264 (2013).
Hoskins, R.A. et al. Genome-wide analysis of promoter architecture in Drosophila melanogaster. Genome Res. 21, 182–192 (2011).
Katayama, S., Kanamori, M. & Hayashizaki, Y. Integrated analysis of the genome and the transcriptome by FANTOM. Brief. Bioinform. 5, 249–258 (2004).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Peng, J.C. et al. Jarid2/Jumonji coordinates control of PRC2 enzymatic activity and target gene occupancy in pluripotent cells. Cell 139, 1290–1302 (2009).
Ietswaart, R., Wu, Z. & Dean, C. Flowering time control: another window to the connection between antisense RNA and chromatin. Trends Genet. 28, 445–453 (2012).
Chen, H.H., Maeda, T., Mullett, S.J. & Stewart, A.F. Transcription cofactor Vgl-2 is required for skeletal muscle differentiation. Genesis 39, 273–279 (2004).
Pelechano, V. & Steinmetz, L.M. Gene regulation by antisense transcription. Nat. Rev. Genet. 14, 880–893 (2013).
Williams, J.A., Paddock, S.W., Vorwerk, K. & Carroll, S.B. Organization of wing formation and induction of a wing-patterning gene at the dorsal/ventral compartment boundary. Nature 368, 299–305 (1994).
Kim, J. et al. Integration of positional signals and regulation of wing formation and identity by Drosophila vestigial gene. Nature 382, 133–138 (1996).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Gemkow, M.J., Buchenau, P. & Arndt-Jovin, D.J. FISH in whole-mount Drosophila embryos. RNA: activation of a transcriptional locus, DNA: gene architecture and expression. Bioimaging 4, 107–120 (1996).
Milán, M., Campuzano, S. & Garcia-Bellido, A. Cell cycling and patterned cell proliferation in the wing primordium of Drosophila. Proc. Natl. Acad. Sci. USA 93, 640–645 (1996).
Aegerter-Wilmsen, T., Aegerter, C.M., Hafen, E. & Basler, K. Model for the regulation of size in the wing imaginal disc of Drosophila. Mech. Dev. 124, 318–326 (2007).
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156 (2007).
Wang, J.W., Beck, E.S. & McCabe, B.D. A modular toolset for recombination transgenesis and neurogenetic analysis of Drosophila. PLoS ONE 7, e42102 (2012).
Rai, A.N. et al. Elements of the polycomb repressor SU(Z)12 needed for histone H3-K27 methylation, the interface with E(Z), and in vivo function. Mol. Cell. Biol. 33, 4844–4856 (2013).
Steffen, P.A. et al. Quantitative in vivo analysis of chromatin binding of Polycomb and Trithorax group proteins reveals retention of ASH1 on mitotic chromatin. Nucleic Acids Res. 41, 5235–5250 (2013).
Keene, J.D., Komisarow, J.M. & Friedersdorf, M.B. RIP-Chip: the isolation and identification of mRNAs, microRNAs and protein components of ribonucleoprotein complexes from cell extracts. Nat. Protoc. 1, 302–307 (2006).
Orlando, V., Jane, E.P., Chinwalla, V., Harte, P.J. & Paro, R. Binding of trithorax and Polycomb proteins to the bithorax complex: dynamic changes during early Drosophila embryogenesis. EMBO J. 17, 5141–5150 (1998).
Carrington, E.A. & Jones, R.S. The Drosophila Enhancer of zeste gene encodes a chromosomal protein: examination of wild-type and mutant protein distribution. Development 122, 4073–4083 (1996).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Aszódi, A. MULTOVL: fast multiple overlaps of genomic regions. Bioinformatics 28, 3318–3319 (2012).
Valouev, A. et al. Genome-wide analysis of transcription factor binding sites based on ChIP-Seq data. Nat. Methods 5, 829–834 (2008).
Gentleman, R.C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Acknowledgements
We thank S. Gasser for discussions and for critical reading of the manuscript. We thank M. Rehmsmeier and members of our laboratories for discussions, P. Pasierbek for advice and training on imaging, P.A. Steffen (IMBA) for the GFP and E(z)∷GFP constructs, C. Ehrhardt and E. Dworschak for technical assistance, B. Dickson (Janelia Farm) for the enGAL4 driver line, J.M. Dura (Institute of Human Genetics, Montpelier, France) for the daGAL4 driver line, I. Tamir (CSF Vienna) for the bioinformatics analysis of ChIP-seq data and sharing the 'Fuge' algorithm, F. Bantignies (Institute of Human Genetics, Montpelier, France) for advice on three-dimensional DNA FISH, R. Jones (Dedman College, Southern Methodist University) for providing antibody to E(Z) and the Vienna Campus Support Facility (CSF) for library preparation, deep sequencing and the purification of Drosophila PRC2. This work was funded by the Austrian Academy of Sciences, by European Community grants European Union Framework Programme 6 Network of Excellence 'The Epigenome' (to L.R.) and the European Union Framework Programme 7 Network of Excellence 'Epigenesys' (to L.R.), and by an FWF Austrian Science Fund grant (P21525-B20 to L.R.).
Author information
Authors and Affiliations
Contributions
A.L. and L.R. initiated the project. L.R., V.A.H. and A.L. designed the experiments. J.T. performed bioinformatics analysis of genome-wide data sets. G.S. and K.A. performed automated image analysis of three-dimensional DNA FISH. V.A.H., A.L., H.O., C.A., F.R., B.B., A.D., M.R., H.B.S. and L.R. conducted the experiments and analyzed the data. S.K. supervised B.B. A.P. supervised M.R. M.L.V. and J.A.S. provided purified Drosophila PRC2 and polynucleosome substrates. L.R. prepared the manuscript with input from V.A.H., J.T., A.L., B.B. and H.O.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Coding and noncoding transcription at the vg locus.
(a,b) qPCR analysis of developmental and tissue-specific vg mRNA (a) and PRE/TRE (b) transcription, shown as percentage of TBP levels. (c) Time course of vg mRNA (diamonds) and PRE/TRE transcription (circles) during embryonic development. Left, scale for mRNA; right, scale for the PRE/TRE transcript, both are shown as the percentage of TBP levels. Error bars show the s.e.m. of four independent cDNAs prepared from at least two independent RNA preparations for each tissue or time point.
Supplementary Figure 2 Mapping of vg PRE/TRE transcripts.
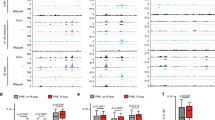
(a) Transcript summary reproduced from Figure 1b. The length of each transcript is given. Colored bars indicate PRE/TRE DNA motifs. Gray box: the 1.6-kb core PRE/TRE26 used for the transgenes (Figs. 2–4 and 6g). (b) Comparison of PRE/TRE prediction score and ChIP enrichments across the PRE/TRE and flanking regions. Genomic coordinates according to Drosophila release Dm3/R5 are shown below. The sequence included in the transgene is highlighted in gray. Top, PRE/TRE prediction score plot generated by jPREdictor (Nucleic Acids Res. 34, W546–W550, 2006). Below are shown ChIP tracks for PC, E(Z) and H3K27me3. Dark gray highlights ChIP-seq results (this study) from 0- to 16-h wild-type embryos. Experiments were performed at least three times for each antibody, and representative tracks from one experiment are shown. Light gray highlights ChIP-chip results from the Sg4 derivative of Drosophila S2 cultured cells25. (c) Key to the annotation in d of 5′ ends mapped by 5′ RACE for the transcripts shown in a. (d) Sequence of the 3.1-kb vg PRE/TRE region shown in a, with 5′ ends shown for each transcript according to the color code in c. Each transcript has several 5′ ends. The diagram in a shows the transcript with the most 5′ start site (i.e., the longest transcript) in each case. These start sites are marked with the symbols outlined in black in d. 5′ RACE mapping was performed at least twice on two independent cDNAs for each tissue. The integrity of each transcript was confirmed by additional PCR analysis (see Supplementary Fig. 3). The 1.6-kb core PRE/TRE used for the transgenes is shown in bold type and outlined in gray. Transgenic transcripts were mapped in homozygous embryos carrying pKC27vg.fwd without induced transcription by GAL4, thus defining the transcripts that are generated by the PRE/TRE promoters carried on the transgene. Mapping of PRE/TRE transcripts in early (0- to 5-h) embryos in which no transcription from the endogenous vg PRE/TRE locus is detectable confirmed that the transgenes generate the two embryonic transcripts whose promoters are within the 1.6-kb transgene (embryo 798 and embryo 1194; see also Supplementary Fig. 3). Mapping was performed at least twice on two independent cDNAs. However, in all other tissues, the background levels of endogenous transcripts precluded unambiguous mapping of the transgenic transcripts. Owing to their highly cell type–specific expression, the transcripts were not detectable by genome-wide profiling methods such as RNA-seq, even in staged embryos (data not shown). They were also not detectable in published genome-wide profiling data sets, presumably for the same reasons. However, four independent methods agree on the identity and presence of transcripts: in situ hybridization (Fig. 1), 5′ RACE (Supplementary Fig. 2d), primer walking (Supplementary Fig. 3b,c) and cDNA cloning (Supplementary Fig. 3d).
Supplementary Figure 3 Primer walking analysis of vg PRE/TRE transcripts.
(a) Transcript summary reproduced from Figure 1b. The length of each transcript is given. Colored bars indicate PRE/TRE DNA motifs. The gray box highlights the 1.6-kb core PRE/TRE26 used for the transgenes (Figs. 2–4 and 6g). (b) The positions and names of the primers used for primer walking analysis. See Supplementary Table 1 for primer sequences. (c) Primer walking example. Left, mapping the 5′ end of the endogenous embryo 798 transcript. RNA was prepared from 10- to 14-h wild-type embryos, and cDNA was primed with primer 10R, to specifically amplify forward-strand transcripts. PCR was performed with 10R in combination with different reverse primers, as indicated and, as a control, on wild-type genomic DNA. Products are seen on cDNA with forward primers 5F and 6F, indicating that the transcript starts downstream of primer 4F. Right, mapping the 5′ end of the endogenous embryo 796 transcript. RNA was prepared from 10- to 14-h wild-type embryos, and cDNA was primed with primer 12F, to specifically amplify reverse-strand transcripts. PCR was performed with primer 12F in combination with different forward primers, as indicated, and, as a control, on wild-type genomic DNA. Products are seen on cDNA with forward primers 14R and 15R, indicating that the transcript starts upstream of primer 16R. 5′ RACE analysis confirmed the results of primer walking and revealed several TSSs for each transcript (see Supplementary Fig. 2). The 3′ ends were defined by performing the above analysis using different primers to prime the first cDNA strand for each transcript. Mapping was performed at least twice on two independent cDNAs. (d) The primers giving the longest transcript defined in each tissue on each strand are shown. In wing and brain, no transcripts were detected from the reverse strand. The transgenic transcripts were mapped in homozygous pKC27vg.fwd transgenic embryos in the absence of the GAL4 driver, as described in the legend to Supplementary Figure 2.
Supplementary Figure 4 Coding and noncoding vg transcription in late embryos.
Double in situ hybridization on embryos showing vg mRNA (red), PRE/TRE reverse-strand transcript (green) and DAPI (blue). Anterior is to the right, posterior is to the left. Scale bar, 20 μm. (a–f) Single optical slices at high magnification through muscle precursors (m), wing (w) and haltere (h) primordia at 12 h (stage 15; a–c) and at 14 h (stage 16; d–f). vg mRNA accumulates in all cells of the wing and haltere disc primordia (a,d)15,16, while the PRE/TRE transcript is absent from wing and haltere discs but highly expressed in segment T2 and T3 muscles (m). (g–l) Single optical slices at high magnification through abdominal body wall muscles in segments A1–A3, at 12 h (stage 15; g–i) and at 14 h (stage 16; j–l). At 12 h, vg mRNA (g) and the PRE/TRE transcript (h) colocalize in nerves (arrows). The PRE/TRE is expressed strongly in muscles (m) at 12 h and 14 h (h,k), while vg mRNA becomes strongly expressed in muscles at 14 h (j). Embryo in situ hybridizations were performed at least three times, and 10–20 embryos were analyzed.
Supplementary Figure 5 vg mRNA and PRE/TRE forward-strand transcription in embryos.
Double in situ hybridization on 10-h (stage 13) embryo showing (a,c) vg mRNA (red), (b,c) PRE/TRE forward-strand transcript (green) and (a–c) DAPI (blue). Anterior is to the right, posterior is to the left. Lateral view, dorsal at top, ventral at bottom. Wing (w) and haltere (h) primordial, muscle precursors (m) and central nervous system (CNS) are indicated in c. vg mRNA is expressed in specific cells (see also Fig. 1); the PRE/TRE forward strand is strongly detected in the head segments and in a diffuse pattern throughout the rest of the embryo. Embryo in situ hybridizations were performed at least three times, and 10–20 embryos were analyzed.
Supplementary Figure 6 Coding and noncoding vg transcription in larval brain.
Double in situ hybridization on 3rd instar larval brains, showing vg mRNA (red), PRE/TRE forward-strand transcript (green) and DAPI (blue) (see Fig. 1b for PRE/TRE transcript orientation and in situ probe). (a–c) Single optical slices showing that vg mRNA and the PRE/TRE transcript are expressed in a complementary pattern. (d–f) High magnification of the boxes in a–c. (e) The PRE/TRE transcript is visible both in the cytoplasm and as nuclear dots (arrows). In situ hybridizations were performed at least three times, and at least five brains were analyzed.
Supplementary Figure 7 Three-dimensional DNA FISH in 3rd instar larval wing discs.
The figure shows data from the individual discs that are shown as merged data in Figure 3a,b as the distribution of three-dimensional distances between the transgenic locus at site 1 and the endogenous vg locus. Transgenic flies (a) carrying a control construct (pKC27mw), (b) expressing the reverse strand (pKC27vg.rev′) or (c) expressing the forward strand (pKC27vg.fwd′) of the vg PRE/TRE were crossed to the line with the enGAL4 driver. enGAL4 is expressed in the posterior half of the wing disc (P). The anterior half of the wing disc (A) serves as a control (see Fig. 3g,l, for expression patterns). For each genotype, data were normalized between discs. Note that, in Figure 3a,b, the data were further normalized between genotypes (for normalization procedures, see additional methods). The y axis shows the three-dimensional distance between the transgenic locus and the endogenous vg locus. Average nuclear radius: 3.6 µm. For two of three discs, induced transcription of the forward strand but not the reverse strand leads to a significant decrease in distances between the transgene and the endogenous locus in the posterior compartment in comparison to the anterior compartment (NS, P value ≥ 0.1, Wilcoxon rank-sum test). Note that all three discs from c are combined in Fig. 3a,b,).
Supplementary Figure 8 In situ hybridization showing the effects of vg RNAi on vg mRNA.
In situ hybridization to detect vg mRNA in the wing discs of transgenic 3rd instar larvae expressing UAS RNAi against vg under the control of the enGAL4 driver. Posterior is to the left, anterior is to the right. Scale bar, 100 µm. enGAL4 is expressed in the posterior half of each disc; see also Figure 3g,l,. (a,b) Control UAS transgene without RNAi. (c–f) Two examples of UAS RNAi. Red, vg mRNA; blue, DAPI. Arrows indicate the borders of the domain of downregulation of vg mRNA. vg mRNA is only downregulated in the central domain of the disc (presumably the domain of highest expression of enGAL4), and the size of the domain in which maximum vg mRNA downregulation occurs is different between discs. The experiment was performed once, and 10 discs were analyzed. Two representative discs are shown to illustrate the variability in vg downregulation.
Supplementary Figure 9 DELFIA assay for HMTase activity.
(a) Detection of H3 peptide mono-, di-, tri- or unmethylated at lysine 27 (500 nM peptide) by DELFIA assay. The y axis shows raw fluorescence measurements. The addition of 380 nM RNA (control RNA 1 in b) leads to a slight decrease in fluorescence (columns 4 and 6) but cannot account for the decrease observed in the presence of enzyme and RNA in b. (b) Comparison of inhibition of human PRC2 HMTase activity by different molecules by DELFIA assay. Unmodified H3 21-40 peptide (500 nM) was incubated with 155 nM human PRC2 complex and either 10 mM of the EZH2 inhibitor GSK126 or 380 nM of different nucleic acids as indicated. RNA fwd, rev: a 1.6-kb RNA in vitro transcribed from the forward or reverse strand of the PRE/TRE shown in Figures 1b (gray box) and 2a or control bacterial RNAs of the same length (ctr1, ctr2; see additional methods). DNA vg: a 1.6-kb double-stranded DNA corresponding to the vg PRE/TRE as shown in Figures 1b (gray box) and 2a. Ctr1 and 2 DNAs: double-stranded DNAs containing the same sequence as the ctr1 and 2 RNAs. (a,b) The mean and s.e.m. of three replicates are shown.
Supplementary Figure 10 ChIP-seq on the endogenous vg locus in the presence of overexpression of PRE/TRE transcripts.
ChIP-seq analysis of E(Z) and H3K27me3 in 0- to 16-h embryos overexpressing the forward- (top; pKC27vg.fwd′) or reverse- (bottom; pKC27vg.rev′) strand from the transgene at site 1. No difference in E(Z) or H3K27 profiles was detectable at the endogenous vg locus. The experiment was repeated three times for H3K27me3 on pKC27vg.fwd′, once for H3K27me3 on pKC27vg.rev′ and once for E(Z) on both pKC27vg.fwd′ and pKC27vg.rev′. Tracks from one representative ChIP experiment are shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1 and 3. (PDF 3324 kb)
PcG-bound regions in fly and mouse that show convergent or divergent transcripts.
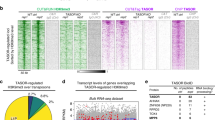
Genomic coordinates of PcG-bound regions with the identity of the closest gene. See the Online Methods and legend in the table for details. (a) mouse_CAGE. Mouse ES cell PcG peaks from ref. 34 that overlap with convergent or divergent CAGE tags from the FANTOM3 CAGE dataset32. In mouse, the CAGE data provide tags from multiple developmental stages and tissues, and the majority of the PcG-bound peaks have multiple tags. Individual tags are not listed in the table, and the coordinates of these are available on request. (b) Fly_modENCODE_MACE. Fly PcG peaks from ref. 21 that overlap with convergent or divergent MACE tags from ref. 20. (c) Fly_modENCODE_MACE_tags. Fly PcG peaks from ref. 21 that overlap with convergent or divergent MACE tags from ref. 20 showing all MACE tags overlapping with each peak. (d) Fly_modENCODE_CAGE. Fly PcG peaks from ref. 21 that overlap with convergent or divergent CAGE tags from ref. 31. (e) Fly_modENCODE_CAGE_tags. Fly PcG peaks from ref. 21 that overlap with convergent or divergent CAGE tags from ref. 31 showing all CAGE tags overlapping with each peak. The source of tag data is listed as 'peaks' (indicating data from CAGE peaks in Supplemental Data File 4 of ref. 31), or 'file 3' (indicating RACE and other annotated transcript start sites from Supplemental Data File 3 of ref. 31). (f) Fly_Enderle_MACE. Fly PcG peaks from ref. 20 that overlap with convergent or divergent MACE tags from the same study. (g) Fly_Enderle_MACE_tags. Fly PcG peaks from ref. 20 that overlap with convergent or divergent MACE tags from the same study showing all MACE tags overlapping with each peak. (h) Fly_Enderle_CAGE. Fly PcG peaks from ref. 20 that overlap with convergent or divergent CAGE tags from ref. 31. (i) Fly_Enderle_CAGE_tags. Fly PcG peaks from ref. 20 that overlap with convergent or divergent CAGE tags from ref. 31 showing all CAGE tags overlapping with each peak. (XLSX 6353 kb)
Three-dimensional animation of vg mRNA showing downregulation upon noncoding RNA expression.
The animation shows the three-dimensional reconstruction of the z stack from the double in situ hybridizations shown in Figure 3l-n. The animation shows a transgenic 3rd instar larval wing disc overexpressing the forward strand from pKC27vg.fwd′ under the control of the enGAL4 driver, which expresses in the posterior half of the disc. At the start of the animation, posterior is to the left, anterior is to the right. The domain of enGAL4 expression was defined in three dimensions using the noncoding RNA signal and is shown in green at the start of the animation, with the remaining ones shown in blue. As the animation plays, first the blue mask and then the green mask is removed, showing the in situ signals: green, forward noncoding RNA; red, vg mRNA; blue, DAPI. Subsequently, the noncoding RNA and DAPI channels are removed, leaving the vg mRNA signal in the red channel. The image is then rotated to show the downregulation of vg mRNA in the posterior part of the disc. See also Figure 3l-p. (MPG 11962 kb)
Rights and permissions
About this article
Cite this article
Herzog, V., Lempradl, A., Trupke, J. et al. A strand-specific switch in noncoding transcription switches the function of a Polycomb/Trithorax response element. Nat Genet 46, 973–981 (2014). https://doi.org/10.1038/ng.3058
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3058
This article is cited by
-
Genomic and functional conservation of lncRNAs: lessons from flies
Mammalian Genome (2022)
-
A theoretical model of Polycomb/Trithorax action unites stable epigenetic memory and dynamic regulation
Nature Communications (2020)
-
RNA-DNA strand exchange by the Drosophila Polycomb complex PRC2
Nature Communications (2020)
-
RNA exploits an exposed regulatory site to inhibit the enzymatic activity of PRC2
Nature Structural & Molecular Biology (2019)
-
G-tract RNA removes Polycomb repressive complex 2 from genes
Nature Structural & Molecular Biology (2019)