Abstract
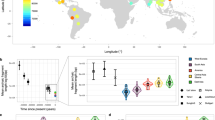
Comparisons of chromosome X and the autosomes can illuminate differences in the histories of males and females as well as shed light on the forces of natural selection. We compared the patterns of variation in these parts of the genome using two datasets that we assembled for this study that are both genomic in scale. Three independent analyses show that around the time of the dispersal of modern humans out of Africa, chromosome X experienced much more genetic drift than is expected from the pattern on the autosomes. This is not predicted by known episodes of demographic history, and we found no similar patterns associated with the dispersals into East Asia and Europe. We conclude that a sex-biased process that reduced the female effective population size, or an episode of natural selection unusually affecting chromosome X, was associated with the founding of non-African populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hammer, M.F. et al. Heterogeneous patterns of variation among multiple human x–linked loci: the possible role of diversity-reducing selection in non-Africans. Genetics 167, 1841–1853 (2004).
Hammer, M.F., Mendez, F.L., Cox, M.P., Woerner, A.E. & Wall, J.D. Sex-biased evolutionary forces shape genomic patterns of human diversity. PLoS Genet. 4, e1000202 (2008).
The International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Li, J.Z. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
Akey, J.M., Zhang, G., Zhang, K., Jin, L. & Shriver, M.D. Interrogating a high-density SNP map for signatures of natural selection. Genome Res. 12, 1805–1814 (2002).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Clark, A.G., Hubisz, M.J., Bustamante, C.D., Williamson, S.H. & Nielsen, R. Ascertainment bias in studies of human genome-wide polymorphism. Genome Res. 15, 1496–1502 (2005).
Keinan, A., Mullikin, J.C., Patterson, N. & Reich, D. Measurement of the human allele frequency spectrum demonstrates greater genetic drift in East Asians than in Europeans. Nat. Genet. 39, 1251–1255 (2007).
Fay, J.C. & Wu, C.I. A human population bottleneck can account for the discordance between patterns of mitochondrial versus nuclear DNA variation. Mol. Biol. Evol. 16, 1003–1005 (1999).
Garrigan, D. & Hammer, M.F. Reconstructing human origins in the genomic era. Nat. Rev. Genet. 7, 669–680 (2006).
Pool, J.E. & Nielsen, R. Population size changes reshape genomic patterns of diversity. Evolution Int. J. Org. Evolution 61, 3001–3006 (2007).
Andolfatto, P. Contrasting patterns of X-linked and autosomal nucleotide variation in Drosophila melanogaster and Drosophila simulans. Mol. Biol. Evol. 18, 279–290 (2001).
Betancourt, A.J., Kim, Y. & Orr, H.A. A pseudohitchhiking model of X vs. autosomal diversity. Genetics 168, 2261–2269 (2004).
Wall, J.D., Andolfatto, P. & Przeworski, M. Testing models of selection and demography in Drosophila simulans. Genetics 162, 203–216 (2002).
Charlesworth, B., Coyne, J.A. & Barton, N.H. The relative rates of evolution of sex chromosomes and autosomes. Am. Nat. 130, 113–146 (1987).
Charlesworth, B., Morgan, M.T. & Charlesworth, D. The effect of deleterious mutations on neutral molecular variation. Genetics 134, 1289–1303 (1993).
Haddrill, P.R., Thornton, K.R., Charlesworth, B. & Andolfatto, P. Multilocus patterns of nucleotide variability and the demographic and selection history of Drosophila melanogaster populations. Genome Res. 15, 790–799 (2005).
Storz, J.F., Payseur, B.A. & Nachman, M.W. Genome scans of DNA variability in humans reveal evidence for selective sweeps outside of Africa. Mol. Biol. Evol. 21, 1800–1811 (2004).
Kayser, M. et al. Reduced Y-chromosome, but not mitochondrial DNA, diversity in human populations from West New Guinea. Am. J. Hum. Genet. 72, 281–302 (2003).
Seielstad, M.T., Minch, E. & Cavalli-Sforza, L.L. Genetic evidence for a higher female migration rate in humans. Nat. Genet. 20, 278–280 (1998).
Marlowe, F.W. Hunter-gatherers and human evolution. Evol. Anthropol. 14, 54–67 (2005).
Bedoya, G. et al. Admixture dynamics in Hispanics: a shift in the nuclear genetic ancestry of a South American population isolate. Proc. Natl. Acad. Sci. USA 103, 7234–7239 (2006).
Hammer, M.F. et al. Hierarchical patterns of global human Y-chromosome diversity. Mol. Biol. Evol. 18, 1189–1203 (2001).
Helgason, A. et al. Estimating Scandinavian and Gaelic ancestry in the male settlers of Iceland. Am. J. Hum. Genet. 67, 697–717 (2000).
Parra, E.J. et al. Estimating African American admixture proportions by use of population-specific alleles. Am. J. Hum. Genet. 63, 1839–1851 (1998).
Caballero, A. On the effective size of populations with separate sexes, with particular reference to sex-linked genes. Genetics 139, 1007–1011 (1995).
Charlesworth, B. The effect of life-history and mode of inheritance on neutral genetic variability. Genet. Res. 77, 153–166 (2001).
Patterson, N. et al. Methods for high-density admixture mapping of disease genes. Am. J. Hum. Genet. 74, 979–1000 (2004).
Ning, Z., Cox, A.J. & Mullikin, J.C. SSAHA: a fast search method for large DNA databases. Genome Res. 11, 1725–1729 (2001).
Tang, K. et al. Chip-based genotyping by mass spectrometry. Proc. Natl. Acad. Sci. USA 96, 10016–10020 (1999).
Acknowledgements
We thank C. Aquadro, O. Bar-Yosef, M. Bernstein, B. Charlesworth, A. Helgason, E. Lander, D. Lieberman, K. Lohmueller, S. Myers, S. Pääbo, A. Price, S. Schaffner and C. Stringer for comments, and J. Neubauer and A. Waliszewska for genotyping SNPs discovered in two West African X chromosomes. The orangutan sequence reads were generated by the Washington University genome sequencing center (ftp.ncbi.nih.gov/pub/TraceDB/pongo_pygmaeus_abelii); we thank R. Wilson for permission to use these data. J.C.M. was supported by the Intramural Research Program of the US National Human Genome Research Institutes (NHGRI), N.P. by a K-01 career transition award from the US National Institutes of Health (NIH) and D.R. by a Burroughs Wellcome Career Development Award in the Biomedical Sciences. A.K., N.P. and D.R. were also supported by NIH grant U01 HG004168.
Author information
Authors and Affiliations
Contributions
A.K. and D.R. designed the study; J.C.M. and A.K. assembled datasets; D.R., A.K. and J.C.M. conducted genotyping; A.K., J.C.M. and D.R. performed analyses; N.P. provided guidance on statistical analyses; A.K. and D.R. interpreted results and wrote the manuscript, which was edited by all co-authors.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–4, Supplementary Figures 1 and 2, Supplementary Methods and Supplementary Note (PDF 530 kb)
Rights and permissions
About this article
Cite this article
Keinan, A., Mullikin, J., Patterson, N. et al. Accelerated genetic drift on chromosome X during the human dispersal out of Africa. Nat Genet 41, 66–70 (2009). https://doi.org/10.1038/ng.303
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.303
This article is cited by
-
Incomplete lineage sorting rather than hybridization explains the inconsistent phylogeny of the wisent
Communications Biology (2018)
-
Fast diffusion of domesticated maize to temperate zones
Scientific Reports (2017)
-
The genetic history of Cochin Jews from India
Human Genetics (2016)
-
Genome-culture coevolution promotes rapid divergence of killer whale ecotypes
Nature Communications (2016)
-
The Simons Genome Diversity Project: 300 genomes from 142 diverse populations
Nature (2016)