Abstract
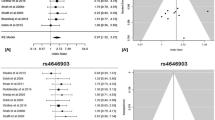
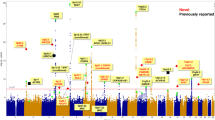
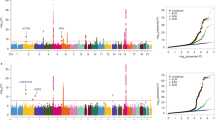
We conducted imputation to the 1000 Genomes Project of four genome-wide association studies of lung cancer in populations of European ancestry (11,348 cases and 15,861 controls) and genotyped an additional 10,246 cases and 38,295 controls for follow-up. We identified large-effect genome-wide associations for squamous lung cancer with the rare variants BRCA2 p.Lys3326X (rs11571833, odds ratio (OR) = 2.47, P = 4.74 × 10−20) and CHEK2 p.Ile157Thr (rs17879961, OR = 0.38, P = 1.27 × 10−13). We also showed an association between common variation at 3q28 (TP63, rs13314271, OR = 1.13, P = 7.22 × 10−10) and lung adenocarcinoma that had been previously reported only in Asians. These findings provide further evidence for inherited genetic susceptibility to lung cancer and its biological basis. Additionally, our analysis demonstrates that imputation can identify rare disease-causing variants with substantive effects on cancer risk from preexisting genome-wide association study data.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
23 January 2017
In the version of this article initially published, the name of author Florence Le Calvez-Kelm appeared incorrectly as Florence LeCalvez-Kelm. The error has been corrected in the HTML and PDF versions of the article.
References
Ferlay, J. et al. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int. J. Cancer 127, 2893–2917 (2010).
Hung, R.J. et al. A susceptibility locus for lung cancer maps to nicotinic acetylcholine receptor subunit genes on 15q25. Nature 452, 633–637 (2008).
Amos, C.I. et al. Genome-wide association scan of tag SNPs identifies a susceptibility locus for lung cancer at 15q25.1. Nat. Genet. 40, 616–622 (2008).
Thorgeirsson, T.E. et al. A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature 452, 638–642 (2008).
McKay, J.D. et al. Lung cancer susceptibility locus at 5p15.33. Nat. Genet. 40, 1404–1406 (2008).
Wang, Y. et al. Common 5p15.33 and 6p21.33 variants influence lung cancer risk. Nat. Genet. 40, 1407–1409 (2008).
Hu, Z. et al. A genome-wide association study identifies two new lung cancer susceptibility loci at 13q12.12 and 22q12.2 in Han Chinese. Nat. Genet. 43, 792–796 (2011).
Miki, D. et al. Variation in TP63 is associated with lung adenocarcinoma susceptibility in Japanese and Korean populations. Nat. Genet. 42, 893–896 (2010).
Lan, Q. et al. Genome-wide association analysis identifies new lung cancer susceptibility loci in never-smoking women in Asia. Nat. Genet. 44, 1330–1335 (2012).
Travis, W.D. et al. International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society: international multidisciplinary classification of lung adenocarcinoma: executive summary. Proc. Am. Thorac. Soc. 8, 381–385 (2011).
Broderick, P. et al. Deciphering the impact of common genetic variation on lung cancer risk: a genome-wide association study. Cancer Res. 69, 6633–6641 (2009).
Landi, M.T. et al. A genome-wide association study of lung cancer identifies a region of chromosome 5p15 associated with risk for adenocarcinoma. Am. J. Hum. Genet. 85, 679–691 (2009).
Timofeeva, M.N. et al. Influence of common genetic variation on lung cancer risk: meta-analysis of 14,900 cases and 29,485 controls. Hum. Mol. Genet. 21, 4980–4995 (2012).
Shi, J. et al. Inherited variation at chromosome 12p13.33, including RAD52, influences the risk of squamous cell lung carcinoma. Cancer Discov. 2, 131–139 (2012).
Huang, Y.T. et al. Cigarette smoking increases copy number alterations in nonsmall-cell lung cancer. Proc. Natl. Acad. Sci. USA 108, 16345–16350 (2011).
Rafnar, T. et al. Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat. Genet. 41, 221–227 (2009).
Mathieson, I. & McVean, G. Differential confounding of rare and common variants in spatially structured populations. Nat. Genet. 44, 243–246 (2012).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
Michailidou, K. et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat. Genet. 45, 353–361 (2013).
Akbari, M.R. et al. Germline BRCA2 mutations and the risk of esophageal squamous cell carcinoma. Oncogene 27, 1290–1296 (2008).
Martin, S.T. et al. Increased prevalence of the BRCA2 polymorphic stop codon K3326X among individuals with familial pancreatic cancer. Oncogene 24, 3652–3656 (2005).
Breast Cancer Linkage Consortium. Cancer risks in BRCA2 mutation carriers. J. Natl. Cancer Inst. 91, 1310–1316 (1999).
van Asperen, C.J. et al. Cancer risks in BRCA2 families: estimates for sites other than breast and ovary. J. Med. Genet. 42, 711–719 (2005).
McAllister, K.A. et al. Cancer susceptibility of mice with a homozygous deletion in the COOH-terminal domain of the Brca2 gene. Cancer Res. 62, 990–994 (2002).
Spain, B.H., Larson, C.J., Shihabuddin, L.S., Gage, F.H. & Verma, I.M. Truncated BRCA2 is cytoplasmic: implications for cancer-linked mutations. Proc. Natl. Acad. Sci. USA 96, 13920–13925 (1999).
Yano, K. et al. Nuclear localization signals of the BRCA2 protein. Biochem. Biophys. Res. Commun. 270, 171–175 (2000).
Bahassi, E.M. et al. The checkpoint kinases Chk1 and Chk2 regulate the functional associations between hBRCA2 and Rad51 in response to DNA damage. Oncogene 27, 3977–3985 (2008).
Mazoyer, S. et al. A polymorphic stop codon in BRCA2. Nat. Genet. 14, 253–254 (1996).
Wu, K. et al. Functional evaluation and cancer risk assessment of BRCA2 unclassified variants. Cancer Res. 65, 417–426 (2005).
Brennan, P. et al. Uncommon CHEK2 mis-sense variant and reduced risk of tobacco-related cancers: case control study. Hum. Mol. Genet. 16, 1794–1801 (2007).
Cybulski, C. et al. Constitutional CHEK2 mutations are associated with a decreased risk of lung and laryngeal cancers. Carcinogenesis 29, 762–765 (2008).
Han, F.F., Guo, C.L. & Liu, L.H. The effect of CHEK2 variant I157T on cancer susceptibility: evidence from a meta-analysis. DNA Cell Biol. 32, 329–335 (2013).
Flores, E.R. The roles of p63 in cancer. Cell Cycle 6, 300–304 (2007).
Katoh, I., Aisaki, K.I., Kurata, S.I., Ikawa, S. & Ikawa, Y. p51A (TAp63γ), a p53 homolog, accumulates in response to DNA damage for cell regulation. Oncogene 19, 3126–3130 (2000).
Petitjean, A. et al. Properties of the six isoforms of p63: p53-like regulation in response to genotoxic stress and cross talk with ΔNp73. Carcinogenesis 29, 273–281 (2008).
Hao, K. et al. Lung eQTLs to help reveal the molecular underpinnings of asthma. PLoS Genet. 8, e1003029 (2012).
Omenn, G.S. et al. The β-carotene and retinol efficacy trial (CARET) for chemoprevention of lung cancer in high risk populations: smokers and asbestos-exposed workers. Cancer Res. 54 (suppl. 7), 2038s–2043s (1994).
Scélo, G. et al. Occupational exposure to vinyl chloride, acrylonitrile and styrene and lung cancer risk (Europe). Cancer Causes Control 15, 445–452 (2004).
Feyler, A. et al. Point: myeloperoxidase −463G→A polymorphism and lung cancer risk. Cancer Epidemiol. Biomarkers Prev. 11, 1550–1554 (2002).
Nelis, M. et al. Genetic structure of Europeans: a view from the North-East. PLoS ONE 4, e5472 (2009).
Välk, K. et al. Gene expression profiles of non-small cell lung cancer: survival prediction and new biomarkers. Oncology 79, 283–292 (2010).
Holmen, J. et al. The Nord-Trondelag Health Study 1995–97 (HUNT2): objectives, contents, methods and participation. Norsk Epidemiologi 13, 1932 (2003).
Landi, M.T. et al. Environment And Genetics in Lung cancer Etiology (EAGLE) study: an integrative population-based case-control study of lung cancer. BMC Public Health 8, 203 (2008).
ATBC Cancer Prevention Study Group. The α-tocopherol, β-carotene lung cancer prevention study: design, methods, participant characteristics, and compliance. Ann. Epidemiol. 4, 1–10 (1994).
Hayes, R.B. et al. Methods for etiologic and early marker investigations in the PLCO trial. Mutat. Res. 592, 147–154 (2005).
Calle, E.E. et al. The American Cancer Society Cancer Prevention Study II Nutrition Cohort: rationale, study design, and baseline characteristics. Cancer 94, 2490–2501 (2002).
Eisen, T., Matakidou, A. & Houlston, R. Identification of low penetrance alleles for lung cancer: the GEnetic Lung CAncer Predisposition Study (GELCAPS). BMC Cancer 8, 244 (2008).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Su, L. et al. Genotypes and haplotypes of matrix metalloproteinase 1, 3 and 12 genes and the risk of lung cancer. Carcinogenesis 27, 1024–1029 (2006).
Kong, A. et al. Parental origin of sequence variants associated with complex diseases. Nature 462, 868–874 (2009).
Styrkarsdottir, U. et al. Nonsense mutation in the LGR4 gene is associated with several human diseases and other traits. Nature 497, 517–520 (2013).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Boeing, H., Wahrendorf, J. & Becker, N. EPIC-Germany—a source for studies into diet and risk of chronic diseases. European Investigation into Cancer and Nutrition. Ann. Nutr. Metab. 43, 195–204 (1999).
Dally, H. et al. The CYP3A4*1B allele increases risk for small cell lung cancer: effect of gender and smoking dose. Pharmacogenetics 13, 607–618 (2003).
Penegar, S. et al. National study of colorectal cancer genetics. Br. J. Cancer 97, 1305–1309 (2007).
Timofeeva, M.N. et al. Genetic polymorphisms in 15q25 and 19q13 loci, cotinine levels, and risk of lung cancer in EPIC. Cancer Epidemiol. Biomarkers Prev. 20, 2250–2261 (2011).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G.R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
Zeggini, E. et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 40, 638–645 (2008).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Aulchenko, Y.S., Struchalin, M.V. & van Duijn, C.M. ProbABEL package for genome-wide association analysis of imputed data. BMC Bioinformatics 11, 134 (2010).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Higgins, J.P., Thompson, S.G., Deeks, J.J. & Altman, D.G. Measuring inconsistency in meta-analyses. Br. Med. J. 327, 557–560 (2003).
Johnson, A.D. et al. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics 24, 2938–2939 (2008).
Gabriel, S.B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Thorgeirsson, T.E. et al. Sequence variants at CHRNB3-CHRNA6 and CYP2A6 affect smoking behavior. Nat. Genet. 42, 448–453 (2010).
Acknowledgements
We thank all individuals who participated in this study. We are also grateful to the patients, clinicians and allied health care professions. We thank Z. Chen and K. Boyle for sample handling and data management of the Toronto study, and L. Admas and L.R. Zhang for field recruitment. We thank L. Su, Y. Zhao, G. Liu, J. Wain, R. Heist and K. Asomaning for providing computing support at MDACC. We thank G. Thomas and Synergy Lyon Cancer (Lyon France) for high performance computing support and J. Olivier and A. Chabrier for IARC's PGM ion torrent sequencing optimization and TaqMan genotyping, respectively. We thank D. Goldgar for sharing information from The Consortium of Investigators of Modifiers of BRCA1/2 (CIMBA) on sequence variation in BRCA2 from familial breast cancer analysis. We acknowledge the Icelandic Cancer Registry (http://www.krabbameinsskra.is/indexen.jsp?id=summary) for assistance in the ascertainment of the Icelandic patients with lung cancer. The ICR study made use of genotyping data from the Wellcome Trust Case-Control Consortium 2 (WTCCC2); a full list of the investigators who contributed to the generation of the data is available from http://www.wtccc.org.uk. We acknowledge The Cancer Genome Atlas (TCGA) for their contribution of lung cancer genomic data to this study (TCGA Project Number 3230). We also acknowledge support from the National Institute for Health Research Biomedical Research Centre at the Royal Marsden Hospital. This study was supported by the NIH (U19CA148127, R01CA055769, 5R01CA127219, 5R01CA133996 and 5R01CA121197). The work performed at ICR was supported by Cancer Research UK (C1298/A8780 and C1298/A8362), National Cancer Research Network (NCRN), HEAL, Sanofi-Aventis and National Health Service funding to the Royal Marsden Hospital and Institute of Cancer Research, as well as the National Institute for Health Research Biomedical Research Centre. B.K. was the recipient of a Sir John Fisher Foundation PhD studentship. Work at ICR was also supported by NIH GM103534 and the Institute for Quantitative Biomedical Sciences at Dartmouth to C.I.A. The work performed in Toronto was supported by The Canadian Cancer Society Research Institute (020214), Ontario Institute of Cancer and Cancer Care Ontario Chair Award to R.J.H. and G.L. and the Alan Brown Chair and Lusi Wong Programs at the Princess Margaret Hospital Foundation. The work performed at Heidelberg was supported by Deutsche Krebshilfe (70-2387 and 70-2919) and the German Federal Ministry of Education and Research (EPIC-Heidelberg). The work performed at IARC was supported by the Institut National du Cancer, France, the European Community (LSHG-CT-2005-512113), the Norwegian Cancer Association, the Functional Genomics Programme of Research Council of Norway, the European Regional Development Fund and the State Budget of the Czech Republic (RECAMO, CZ.1.05/2.1.00/03.0101), the NIH (R01-CA111703 and UO1-CA63673), the Fred Hutchinson Cancer Research Center, the US NCI (R01 CA092039), an FP7 grant (REGPOT 245536), the Estonian Government (SF0180142s08), the EU European Regional Development Fund in the frame of Centre of Excellence in Genomics and Estonian Research Infrastructure's Roadmap and the University of Tartu (SP1GVARENG) and an IARC Postdoctoral Fellowship (M.N.T.). Work at the NCI was supported by the Intramural Research Program of the NIH, the NCI, US Public Health Service contracts NCI (N01-CN-45165, N01-RC-45035, N01-RC-37004, NO1-CN-25514, NO1-CN-25515, NO1-CN-25512, NO1-CN-25513, NO1-CN-25516, NO1-CN-25511, NO1-CN-25524, NO1-CN-25518, NO1-CN-75022, NO1-CN-25476 and NO1-CN-25404), the American Cancer Society, the NIH Genes, Environment and Health Initiative in part by HG-06-033-NCI-01 and RO1HL091172-01, genotyping at the Johns Hopkins University Center for Inherited Disease Research (U01HG004438 and NIH HHSN268200782096C) and study coordination at the GENEVA Coordination Center (U01 HG004446). Work was also supported by NIH grants (P50 CA70907, R01CA121197, RO1 CA127219, U19 CA148127 and RO1 CA55769) and a Cancer Prevention Research Institute of Texas grant (RP100443). Genotyping was provided by the Center for Inherited Disease Research (CIDR). Work performed at Harvard was supported by the NIH (CA074386, CA092824 and CA090578). The Icelandic study was supported in part by NIH DA17932.
Author information
Authors and Affiliations
Contributions
R.S.H. and Y. Wang conceived the study and provided overall project management and drafted the paper. In the UK, Y. Wang performed statistics and bioinformatics of UK data and conducted all meta-analyses; additional support was provided by M.H.; P. Broderick oversaw genotyping and sequencing; A.L. and B.K. performed genotyping and Sanger sequencing; A. Matakidou, T.E. and R.S.H. were responsible for the development and operation of the Genetic Lung Cancer Predisposition Study (GELCAPS); and D.C. and P. Broderick performed next-generation sequencing. At IARC, J.D.M. and P. Brennan provided overall project management; M.N.T., M.D.-S., V.G. and M.V. performed statistics and bioinformatics of IARC data and conducted meta-analysis; J.D.M. and F.L.C.-K. oversaw genotyping and sequencing; and G.S., D.Z., N.S.-D., J. Lissowska, P.R., E.F., D.M., V.B., L.F., V.J., H.E.K., M.E.G., F.S., L.V., I.N., C.C., G.G., M. Lathrop, S.B., T.V., K.V., M.N., A. Metspalu, M. Lathrop, J. Lubiński, Mattias Johansson, P.V., A.A., F.C.-C., H.B.-d.-M., D.T., K.-T.K., Mikael Johansson, E.W., A.T., R.K. and E.R. provided samples and data. For the Dartmouth and MDACC component, C.I.A. provided overall project management, obtained support for genotyping and contributed to statistical analyses; W.V.C. performed imputation analysis; Y.H. performed statistical analyses; and M.R.S. oversaw sample collection and development of the epidemiological studies. M.R.S. was also responsible for collecting samples that are a part of this research. X.W. provided ongoing support for the research protocol and supported large laboratory management of samples. Y.Y. and J.G. performed genotyping. At the NCI, M.T.L. was responsible for the overall project and managed the Environment and Genetics in Lung Cancer Etiology (EAGLE) study; N.E.C. managed the Prostate, Lung, Colon, Ovary Screening Trial (PLCO) study; D.A. managed the α-Tocopherol, β-Carotene Cancer Prevention Study (ATBC); S.M.G. and V.L.S. managed the Cancer Prevention Study II Nutrition Cohort (CPS-II) study; N.C. and W.W. performed statistical analyses; Z.W. performed genotyping and imputation analysis; and S.J.C. oversaw genotyping and imputation analysis. At decode, T.R. and K.S. were responsible for the development and operation of deCODE's lung cancer study; and G.T. and P.S. performed the imputations and statistical analysis of the Icelandic data. At Harvard, D.C.C. was responsible for the overall conduct of the project; L.S. was responsible for sample management, genotyping and laboratory quality control; and Y. Wei performed data management and statistical analyses. For the Heidelberg-EPIC replication, M. Laplana managed DNA samples and performed genotyping; A. Rosenberger managed genotype and phenotype information; A. Risch supervised genotyping and data analysis; and R.K., A. Risch and H.D. conceived and managed studies that contributed samples. For the Toronto replication, R.J.H. and G.L. provided overall supervision of the study conduct, including study design, field recruitment, genotyping and statistical analysis; and X.Z. performed the statistical analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 and Supplementary Tables 1, 3-7 (PDF 2142 kb)
Supplementary Table 2
(a) Details of study participants; (b) quality control of GWAS datasets; (c) details of imputation applied to each GWAS dataset. (XLSX 110 kb)
Source data
Rights and permissions
About this article
Cite this article
Wang, Y., McKay, J., Rafnar, T. et al. Rare variants of large effect in BRCA2 and CHEK2 affect risk of lung cancer. Nat Genet 46, 736–741 (2014). https://doi.org/10.1038/ng.3002
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3002
This article is cited by
-
The UK BiLEVE and Mendelian randomisation: using multivariable instrumental variables to address “damned if you, damned if you don’t” adjustment problems
BMC Research Notes (2023)
-
Causal association between inflammatory bowel disease and 32 site-specific extracolonic cancers: a Mendelian randomization study
BMC Medicine (2023)
-
Prevalence of BRCA1 and BRCA2 germline variants in an unselected pancreatic cancer patient cohort in Pakistan
Hereditary Cancer in Clinical Practice (2023)
-
Causal relationship between gut microbiota and cancers: a two-sample Mendelian randomisation study
BMC Medicine (2023)
-
Association between psoriasis and lung cancer: two-sample Mendelian randomization analyses
BMC Pulmonary Medicine (2023)