Abstract
Although mitochondrial DNA (mtDNA) is prone to mutation and few mtDNA repair mechanisms exist1, crippling mitochondrial mutations are exceedingly rare2. Recent studies have demonstrated strong purifying selection in the mouse female germline3,4. However, the mechanisms underlying positive selection of healthy mitochondria remain to be elucidated. We visualized mtDNA replication during Drosophila melanogaster oogenesis, finding that mtDNA replication commenced before oocyte determination during the late germarium stage and was dependent on mitochondrial fitness. We isolated a temperature-sensitive lethal mtDNA allele, mt:CoIT300I, which resulted in reduced mtDNA replication in the germarium at the restrictive temperature. Additionally, the frequency of the mt:CoIT300I allele in heteroplasmic flies was decreased, both during oogenesis and over multiple generations, at the restrictive temperature. Furthermore, we determined that selection against mt:CoIT300I overlaps with the timing of selective replication of mtDNA in the germarium. These findings establish a previously uncharacterized developmental mechanism for the selective amplification of wild-type mtDNA, which may be evolutionarily conserved to limit the transmission of deleterious mutations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wallace, D.C. A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: a dawn for evolutionary medicine. Annu. Rev. Genet. 39, 359–407 (2005).
Rand, D.M. Mitigating mutational meltdown in mammalian mitochondria. PLoS Biol. 6, e35 (2008).
Fan, W. et al. A mouse model of mitochondrial disease reveals germline selection against severe mtDNA mutations. Science 319, 958–962 (2008).
Stewart, J.B. et al. Strong purifying selection in transmission of mammalian mitochondrial DNA. PLoS Biol. 6, e10 (2008).
de Cuevas, M., Lilly, M.A. & Spradling, A.C. Germline cyst formation in Drosophila. Annu. Rev. Genet. 31, 405–428 (1997).
Horwitz, H.B. & Holt, C.E. Specific inhibition by ethidium bromide of mitochondrial DNA synthesis in physarum polycephalum. J. Cell Biol. 49, 546–553 (1971).
Lin, H., Yue, L. & Spradling, A.C. The Drosophila fusome, a germline-specific organelle, contains membrane skeletal proteins and functions in cyst formation. Development 120, 947–956 (1994).
Fornuskova, D. et al. Novel insights into the assembly and function of human nuclear-encoded cytochrome c oxidase subunits 4, 5a, 6a, 7a and 7b. Biochem. J. 428, 363–374 (2010).
Lou, P.-H. et al. Mitochondrial uncouplers with an extraordinary dynamic range. Biochem. J. 407, 129–140 (2007).
Xu, H., DeLuca, S.Z. & O'Farrell, P.H. Manipulating the animal mitochondrial genome with targeted restriction enzymes. Science 321, 575–577 (2008).
Chance, B. & Williams, G.R. Respiratory enzymes in oxidative phosphorylation. III. The steady state. J. Biol. Chem. 217, 409–427 (1955).
Niki, Y., Chigusa, S.I. & Matsuura, E.T. Complete replacement of mitochondrial DNA in Drosophila. Nature 341, 551–552 (1989).
Spradling, A.C. in The Development of Drosophila melanogaster eds. Bate, M. & Martinez, A. 1–70 (CSHL Press, Cold Spring Harbor, NY, 1993).
Chinnery, P.F. et al. The inheritance of mitochondrial DNA heteroplasmy: random drift, selection or both? Trends Genet. 16, 500–505 (2000).
Freyer, C. et al. Variation in germline mtDNA heteroplasmy is determined prenatally but modified during subsequent transmission. Nat. Genet. 44, 1282–1285 (2012).
Ma, H., Xu, H. & O'Farrell, P.H. Transmission of mitochondrial mutations and action of purifying selection in Drosophila melanogaster. Nat. Genet. 10.1038/ng.2919 (9 March 2014).
Shoubridge, E.A. & Wai, T. Medicine. Sidestepping mutational meltdown. Science 319, 914–915 (2008).
Cox, R.T. & Spradling, A.C. A Balbiani body and the fusome mediate mitochondrial inheritance during Drosophila oogenesis. Development 130, 1579–1590 (2003).
Wai, T., Teoli, D. & Shoubridge, E.A. The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat. Genet. 40, 1484–1488 (2008).
Cox, R.T. & Spradling, A.C. Milton controls the early acquisition of mitochondria by Drosophila oocytes. Development 133, 3371–3377 (2006).
Kloc, M., Bilinski, S. & Etkin, L.D. The Balbiani body and germ cell determinants: 150 years later. Curr. Top. Dev. Biol. 59, 1–36 (2004).
Pepling, M.E., Wilhelm, J.E., O'Hara, A.L., Gephardt, G.W. & Spradling, A.C. Mouse oocytes within germ cell cysts and primordial follicles contain a Balbiani body. Proc. Natl. Acad. Sci. USA 104, 187–192 (2007).
Zhou, R.R., Wang, B., Wang, J., Schatten, H. & Zhang, Y.Z. Is the mitochondrial cloud the selection machinery for preferentially transmitting wild-type mtDNA between generations? Rewinding Müller's ratchet efficiently. Curr. Genet. 56, 101–107 (2010).
Yoneda, M., Tanno, Y., Tsuji, S. & Attardi, G. Detection and quantification of point mutations in mitochondrial DNA by PCR. Methods Enzymol. 264, 432–441 (1996).
Koch, E.A. & Spitzer, R.H. Multiple effects of colchicine on oogenesis in Drosophila: induced sterility and switch of potential oocyte to nurse-cell developmental pathway. Cell Tissue Res. 228, 21–32 (1983).
Calleja, M. et al. Mitochondrial DNA remains intact during Drosophila aging, but the levels of mitochondrial transcripts are significantly reduced. J. Biol. Chem. 268, 18891–18897 (1993).
Acknowledgements
We are indebted to P. O'Farrell, in whose laboratory the mt:CoIT300I fly was isolated. We thank R. Balaban, T. Finkel, A. Michelson, H. Parthasarathy, N. Rusan and R. Youle for comments on the manuscript; P. O'Farrell and A. Spradling for their insightful discussions; P. Hwang for technical assistance; the Spradling laboratory (Carnegie Institute) and the Bloomington and Vienna stock centers for Drosophila stocks; Genetic Services, Inc., for embryo injection; and the Developmental Studies Hybridoma Bank for antibodies. This work was supported by the National, Heart, Lung, and Blood Institute (NHLBI) Intramural Program.
Author information
Authors and Affiliations
Contributions
H.X. conceived and designed the experiments. J.H.H., Z.C. and H.X. performed the experiments, analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 mtDNA replication in the Drosophila germarium visualized by EdU incorporation.
(a) Representative confocal section of a wild-type germarium showing EdU incorporation (green), mitochondria marked by α-ATP synthase α subunit staining (red) and nuclei labeled with DAPI staining (blue) in the absence of the DNA polymerase-α inhibitor aphidicolin or (b) with preincubation with aphidicolin. Without aphidicolin, intense EdU signals label nuclei (arrows) (a). In the presence of aphidicolin, nuclear incorporation is reduced and many puncta were localized within mitochondria (arrowheads) (b). Scale bars, 10 μm.
Supplementary Figure 2 The mtDNA replication inhibitor ethidium bromide inhibits EdU incorporation in the Drosophila germarium.
Representative z projections of confocal sections of wild-type germaria incubated without (control) or with ethidium bromide during EdU incorporation. Arrowhead, mtDNA. Scale bars, 10 μm.
Supplementary Figure 3 mtDNA replication occurs around the fusome in region 2B cysts of the germarium.
Representative z-stack projections of confocal images that span 8 μm in depth of a germarium, showing EdU incorporation around the fusome (arrowhead), labeled by the α-Hts 1B1 antibody (red). Scale bar, 5 μm.
Supplementary Figure 4 General respiratory chain dysfunction disrupts mtDNA replication in the germarium.
(a,b) EdU incorporation in the presence (a) or absence (b) of RNAi targeted against the nuclear-encoded mitochondrial protein cytochrome c oxidase subunit Va (CoVa). RNAi-mediated knockdown of CoVa in the germ line (UAS-dicer-2, w1118/+; nanos-Gal4/+; CoVa-IR/+) abolishes mtDNA replication in the germarium, while expression of Dicer-2 alone (UAS-dicer-2, w1118/+; nanos-Gal4/+) has no effect. Representative images from >10 germaria are shown. (c,d) z-stack projections of wild-type germaria subjected to treatment with DMSO or the mitochondria uncoupler DNP during incorporation of EdU. Note the lack of EdU incorporation into mtDNA at region 2B with DNP (box). Arrows, nuclear incorporation; arrowheads, mtDNA replication. Scale bars, 10 μm. (e) Quantification of mtDNA replication (mean ± s.d.) in indicated regions of the germarium treated with DNP and control (DMSO). (DMSO n = 4, DNP n = 5). The number of puncta at region 2B was significantly reduced in germaria subjected to DNP treatment compared to control, P < 0.001.
Supplementary Figure 5 Impaired respiration of mt:CoIT300I mitochondria and reduced activity of mt:CoIT300I COX at the non-permissive temperature.
(a) Average RCRs of three biological replicates of wild-type and mutant mitochondria purified from flies that eclosed at 18 °C and were cultured at 29 °C for 0, 2 or 4 d. Value, mean ± s.d. Mutant mitochondria have significant reduction in the RCR compared to the wild type after shifting to 29 °C, *P < 0.01, **P < 0.001. (b) COX activities of extracts from wild-type and mt:CoIT300I flies that eclosed and were maintained at 18 °C for 1 d. Extracts were incubated at 29 °C or 18 °C for 40 min, and COX activity was examined every 10 min. Data represent three biological replicates. Values were normalized with the average COX activity of wild type at 18 °C and 0 min, mean ± s.d.
Supplementary Figure 6 mtDNA replication in wild-type and homoplasmic mt:CoIT300I ovarioles at 29 °C.
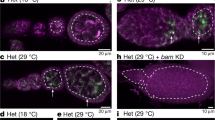
(a) Projections of z stacks showing EdU incorporation in the ovariole germaria of wild-type (top) and mt:CoIT300I (bottom) at 29 °C. Germarium regions 1, 2B and 3 are outlined in boxes from left to right. Note mtDNA replication in a late-stage egg chamber from mt:CoIT300I (outlined). Arrow, nucleus; arrowheads, mtDNA. Scale bars, 10 μm. (b) Quantification of mtDNA replication indicated by the number of EdU puncta in region 1 (wild-type n = 3, mt:CoIT300I n = 5), 2B (wild-type n = 7, mt:CoIT300I n = 7) and 3 (wild-type n = 5, mt:CoIT300I n = 6). Value, mean ± s.d. The number of EdU puncta in region 2B was significantly reduced in mutant germaria compared to wild type, P < 0.0005.
Supplementary Figure 7 Cyst cell death does not contribute to selection against mutant mitochondrial genomes in the germarium of heteroplasmic flies.
(a) Terminal deoxynucleotidyl transferase dUTP nick-end labeling (TUNEL; green) of germaria from wild-type and heteroplasmic mt:CoIT300I ovaries at 18 °C or 29 °C and counterstained with DAPI (pseudocolored with red for better illustration) to label nuclei. Arrowheads, TUNEL-positive nuclei. Scale bars, 10 μm. (b) Quantification of the number of TUNEL-positive nuclei per germarium at 18 °C (wild-type n = 9, mt:CoIT300In = 8) or 29 °C (wild-type n = 14, mt:CoIT300I n = 15). Value, mean ± s.d.
Supplementary Figure 8 Fusome-mediated selective replication of the wild-type genome limits the transmission of mtDNA mutations.
(a) Model for the selective amplification of healthy mitochondria in region 2B of the germarium. See the main text for the description. (b) The impact of overall mutation frequency (f) and mtDNA copy number per mitochondrion (c) on selection. If only the unhealthy mitochondria containing 100% mutant genomes were selected against, the efficacy of selection would be determined by the probability of a given mitochondrion containing 100% mutant genomes can be deduced as
(see Online Methods for detail). The graph shows P as a function of f, with c set at 1, 5 or 10, indicating that the lower the mtDNA copy number per an organelle, the more efficient the selection will be.
Supplementary Figure 9 Mito-GFP labels mitochondria.
(a) An ovariole expressing GFP fusion protein containing the mitochondrial-targeting sequence of citrate synthase (Mito-GFP)11 in germline cells, stained with an antibody against the ATP synthase α subunit (ATP-S; red) that labels mitochondria in both germ cells (arrowheads) and surrounding somatic cells (arrows). A representative image from >10 germaria is shown. Scale bar, 10 μm. (b) Enlarged view of the boxed region in a showing Mito-GFP and ATP-S colocalized at the fusome in region 2B of the germarium. Genotype: w1118/+;; nanos-Gal4, UAS-Mito-GFP/+. Scale bar, 10 μm.
Supplementary Figure 10 The microtubule network and mitochondrial fitness are essential for mitochondria localization at the fusome.
Mito-GFP (green) localization and α-Hts 1B1 staining (red) of wild-type germaria subjected to colchicine or CCCP treatment. Images are z-stack projections of multiple confocal sections of region 2B of the germarium spanning 8 μm in depth. Genotype: w1118/+;; nanos-Gal4, UAS-Mito-GFP/+. Representative images from >10 germaria for each condition are shown. Scale bars, 5 μm.
Supplementary Figure 11 Mitochondria localization to the fusome is disrupted in homoplasmic mt:CoIT300I flies.
Germaria from flies expressing Mito-GFP driven by nanos-Gal4 (w1118;; nanos-Gal4, UAS-Mito-GFP/+) with wild-type or mt:CoIT300I mtDNA at 29 °C, stained with α-Hts 1B1 antibody (red) to illustrate the fusome (arrows). Images are z projections of confocal sections spanning 8 μm in depth. Note Mito-GFP signal at the periphery of cells (arrowhead) in mt:CoIT300I at 29 °C. Representative images from >10 germaria for each genotype are shown. Scale bars, 5 μm.
Supplementary Figure 12 Quantification of heteroplasmy.
(a) Quantification of wild-type mtDNA product amplified by PCR from mixed template (wild type and mt:CoIT300I) and digested with XhoI compared to the amount of wild-type DNA in a digest containing a mix of wild-type and mt:CoIT300I PCR products. Quantified levels were plotted against the actual levels in the mix, and overlap of the two standard curves indicates a lack of impact of the wild type/mt:CoIT300I heteroduplex during PCR on quantification. (b) Quantification of the mt:CoIT300I level throughout fly development. E, embryo; 1L, first instar larvae; 2L, second instar larvae; 3L, third instar larvae; EP, early pupae; LP, late pupae; A, adult. Value, mean ± s.d of 3 biological replicates, each containing 50 flies.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 (PDF 10263 kb)
Rights and permissions
About this article
Cite this article
Hill, J., Chen, Z. & Xu, H. Selective propagation of functional mitochondrial DNA during oogenesis restricts the transmission of a deleterious mitochondrial variant. Nat Genet 46, 389–392 (2014). https://doi.org/10.1038/ng.2920
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2920
This article is cited by
-
Two mitochondrial DNA polymorphisms modulate cardiolipin binding and lead to synthetic lethality
Nature Communications (2024)
-
Effects of adverse fertility-related factors on mitochondrial DNA in the oocyte: a comprehensive review
Reproductive Biology and Endocrinology (2023)
-
Maternal aging increases offspring adult body size via transmission of donut-shaped mitochondria
Cell Research (2023)
-
Sex-linked deubiquitinase establishes uniparental transmission of chloroplast DNA
Nature Communications (2022)
-
Molecular mechanisms of coronary microvascular endothelial dysfunction in diabetes mellitus: focus on mitochondrial quality surveillance
Angiogenesis (2022)