Abstract
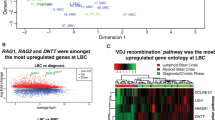
The ETV6-RUNX1 fusion gene, found in 25% of childhood acute lymphoblastic leukemia (ALL) cases, is acquired in utero but requires additional somatic mutations for overt leukemia. We used exome and low-coverage whole-genome sequencing to characterize secondary events associated with leukemic transformation. RAG-mediated deletions emerge as the dominant mutational process, characterized by recombination signal sequence motifs near breakpoints, incorporation of non-templated sequence at junctions, ∼30-fold enrichment at promoters and enhancers of genes actively transcribed in B cell development and an unexpectedly high ratio of recurrent to non-recurrent structural variants. Single-cell tracking shows that this mechanism is active throughout leukemic evolution, with evidence of localized clustering and reiterated deletions. Integration of data on point mutations and rearrangements identifies ATF7IP and MGA as two new tumor-suppressor genes in ALL. Thus, a remarkably parsimonious mutational process transforms ETV6-RUNX1–positive lymphoblasts, targeting the promoters, enhancers and first exons of genes that normally regulate B cell differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bhojwani, D. et al. ETV6-RUNX1–positive childhood acute lymphoblastic leukemia: improved outcome with contemporary therapy. Leukemia 26, 265–270 (2012).
Greaves, M.F. & Wiemels, J. Origins of chromosome translocations in childhood leukaemia. Nat. Rev. Cancer 3, 639–649 (2003).
Mori, H. et al. Chromosome translocations and covert leukemic clones are generated during normal fetal development. Proc. Natl. Acad. Sci. USA 99, 8242–8247 (2002).
Bateman, C.M. et al. Acquisition of genome-wide copy number alterations in monozygotic twins with acute lymphoblastic leukemia. Blood 115, 3553–3558 (2010).
Mullighan, C.G. et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 446, 758–764 (2007).
Zhang, M. & Swanson, P.C. V(D)J recombinase binding and cleavage of cryptic recombination signal sequences identified from lymphoid malignancies. J. Biol. Chem. 283, 6717–6727 (2008).
Mullighan, C.G. et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 322, 1377–1380 (2008).
Raschke, S., Balz, V., Efferth, T., Schulz, W.A. & Florl, A.R. Homozygous deletions of CDKN2A caused by alternative mechanisms in various human cancer cell lines. Genes Chromosom. Cancer 42, 58–67 (2005).
Waanders, E. et al. The origin and nature of tightly clustered BTG1 deletions in precursor B-cell acute lymphoblastic leukemia support a model of multiclonal evolution. PLoS Genet. 8, e1002533 (2012).
Holmfeldt, L. et al. The genomic landscape of hypodiploid acute lymphoblastic leukemia. Nat. Genet. 45, 242–252 (2013).
Hübner, S. et al. High incidence and unique features of antigen receptor gene rearrangements in TEL-AML1–positive leukemias. Leukemia 18, 84–91 (2004).
Schatz, D.G. & Swanson, P.C. V(D)J recombination: mechanisms of initiation. Annu. Rev. Genet. 45, 167–202 (2011).
Fugmann, S.D., Lee, A.I., Shockett, P.E., Villey, I.J. & Schatz, D.G. The RAG proteins and V(D)J recombination: complexes, ends, and transposition. Annu. Rev. Immunol. 18, 495–527 (2000).
Komori, T., Okada, A., Stewart, V. & Alt, F.W. Lack of N regions in antigen receptor variable region genes of TdT-deficient lymphocytes. Science 261, 1171–1175 (1993).
Raghavan, S.C., Swanson, P.C., Ma, Y. & Lieber, M.R. Double-strand break formation by the RAG complex at the Bcl-2 major breakpoint region and at other non-B DNA structures in vitro. Mol. Cell. Biol. 25, 5904–5919 (2005).
Tsai, A.G. et al. Human chromosomal translocations at CpG sites and a theoretical basis for their lineage and stage specificity. Cell 135, 1130–1142 (2008).
Hesse, J.E., Lieber, M.R., Mizuuchi, K. & Gellert, M. V(D)J recombination: a functional definition of the joining signals. Genes Dev. 3, 1053–1061 (1989).
Mullighan, C.G. et al. BCR-ABL1 lymphoblastic leukaemia is characterized by the deletion of Ikaros. Nature 453, 110–114 (2008).
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Campbell, P.J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010).
Baca, S.C. et al. Punctuated evolution of prostate cancer genomes. Cell 153, 666–677 (2013).
Bailey, T.L., Williams, N., Misleh, C. & Li, W.W. MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 34, W369–W373 (2006).
Stephens, P.J. et al. Complex landscapes of somatic rearrangement in human breast cancer genomes. Nature 462, 1005–1010 (2009).
Berger, M.F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Stephens, P.J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Shimazaki, N., Tsai, A.G. & Lieber, M.R. H3K4me3 stimulates the V(D)J RAG complex for both nicking and hairpinning in trans in addition to tethering in cis: implications for translocations. Mol. Cell 34, 535–544 (2009).
Anderson, K. et al. Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature 469, 356–361 (2011).
Potter, N.E. et al. Single cell mutational profiling and clonal phylogeny in cancer. Genome Res. 23, 2115–2125 (2013).
Familiades, J. et al. PAX5 mutations occur frequently in adult B-cell progenitor acute lymphoblastic leukemia and PAX5 haploinsufficiency is associated with BCR-ABL1 and TCF3-PBX1 fusion genes: a GRAALL study. Leukemia 23, 1989–1998 (2009).
Kempski, H. et al. An investigation of the t(12;21) rearrangement in children with B-precursor acute lymphoblastic leukaemia using cytogenetic and molecular methods. Br. J. Haematol. 105, 684–689 (1999).
van Delft, F.W. et al. Clonal origins of relapse in ETV6-RUNX1 acute lymphoblastic leukemia. Blood 117, 6247–6254 (2011).
Hurlin, P.J., Steingrimsson, E., Copeland, N.G., Jenkins, N.A. & Eisenman, R.N. Mga, a dual-specificity transcription factor that interacts with Max and contains a T-domain DNA-binding motif. EMBO J. 18, 7019–7028 (1999).
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Chen, S. et al. Novel non-TCR chromosome translocations t(3;11)(q25;p13) and t(X;11)(q25;p13) activating LMO2 by juxtaposition with MBNL1 and STAG2. Leukemia 25, 1632–1635 (2011).
Jaffe, J.D. et al. Global chromatin profiling reveals NSD2 mutations in pediatric acute lymphoblastic leukemia. Nat. Genet. 45, 1386–1391 (2013).
Harris, R.S., Petersen-Mahrt, S.K. & Neuberger, M.S. RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol. Cell 10, 1247–1253 (2002).
Neuberger, M.S. & Rada, C. Somatic hypermutation: activation-induced deaminase for C/G followed by polymerase eta for A/T. J. Exp. Med. 204, 7–10 (2007).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Roberts, S.A. et al. Clustered mutations in yeast and in human cancers can arise from damaged long single-strand DNA regions. Mol. Cell 46, 424–435 (2012).
Roberts, K.G. et al. Genetic alterations activating kinase and cytokine receptor signaling in high-risk acute lymphoblastic leukemia. Cancer Cell 22, 153–166 (2012).
Tsai, A.G. & Lieber, M.R. RAGs found “not guilty”: cleared by DNA evidence. Blood 111, 1750 (2008).
Boubakour-Azzouz, I., Bertrand, P., Claes, A., Lopez, B.S. & Rougeon, F. Terminal deoxynucleotidyl transferase requires KU80 and XRCC4 to promote N-addition at non-V(D)J chromosomal breaks in non-lymphoid cells. Nucleic Acids Res. 40, 8381–8391 (2012).
Hong, D. et al. Initiating and cancer-propagating cells in TEL-AML1–associated childhood leukemia. Science 319, 336–339 (2008).
Tsuzuki, S., Seto, M., Greaves, M. & Enver, T. Modeling first-hit functions of the t(12;21) TEL-AML1 translocation in mice. Proc. Natl. Acad. Sci. USA 101, 8443–8448 (2004).
Ross, M.E. et al. Classification of pediatric acute lymphoblastic leukemia by gene expression profiling. Blood 102, 2951–2959 (2003).
Kozarewa, I. et al. Amplification-free Illumina sequencing-library preparation facilitates improved mapping and assembly of (G+C)-biased genomes. Nat. Methods 6, 291–295 (2009).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl. Acad. Sci. USA 107, 16910–16915 (2010).
Alexandrov, L.B., Nik-Zainal, S., Wedge, D.C., Campbell, P.J. & Stratton, M.R. Deciphering signatures of mutational processes operative in human cancer. Cell Rep. 3, 246–259 (2013).
Greenman, C., Wooster, R., Futreal, P.A., Stratton, M.R. & Easton, D.F. Statistical analysis of pathogenicity of somatic mutations in cancer. Genetics 173, 2187–2198 (2006).
Klipper-Aurbach, Y. et al. Mathematical formulae for the prediction of the residual β cell function during the first two years of disease in children and adolescents with insulin-dependent diabetes mellitus. Med. Hypotheses 45, 486–490 (1995).
van Dongen, J.J. et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 17, 2257–2317 (2003).
Acknowledgements
This work was supported by the Kay Kendall Leukaemia Fund (KKLF; grant reference KKL407), Leukemia and Lymphoma Research (grant reference 11021) and the Wellcome Trust (grant reference 077012/Z/05/Z). P.J.C. is personally funded through a Wellcome Trust Senior Clinical Research Fellowship (grant reference WT088340MA). P.V.L. is supported by a postdoctoral research fellowship from Research Foundation–Flanders (FWO). S.N.-Z. is a Wellcome Trust Intermediate Clinical Fellow (grant reference WT100183MA). F.W.v.D. is funded by KKLF (grant reference KKL417). J.Z. is supported by University Hospital Motol (grant MH-CR DRO 00064203).
Author information
Authors and Affiliations
Contributions
E.P., M.G. and P.J.C. designed the study and wrote the manuscript, with assistance from M.R.S. E.P. designed experiments, performed experiments, analyzed sequencing data and performed and reviewed bioinformatics and statistical analyses. I.R. performed sample preparation, validation experiments and evaluation of sequencing data. Y.L. performed bioinformatics and statistical analyses and wrote the manuscript. D.C.W., L.B.A., I.M. and P.V.L. performed statistical analysis. N.E.P., I.T., F.W.v.D. and A.M.F. performed experiments. G.G., J.T., C.L., S.L.C., J.M., J.H., A.M., K.R., S.N.-Z., M.R., L.S., D.R.J., A.P.B., J.G. and J.W.T. support variant calling algorithms and sequencing analysis platforms. L.M., B.R. and S.O. performed sample preparation and experiments. J.Z., H.K., G.C., M.M. and A.B. provided and prepared samples and experimental materials. All authors reviewed the manuscript during its preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 (PDF 13180 kb)
Supplementary Tables 1–19
Supplementary Tables 1–19 (XLSX 2768 kb)
Rights and permissions
About this article
Cite this article
Papaemmanuil, E., Rapado, I., Li, Y. et al. RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nat Genet 46, 116–125 (2014). https://doi.org/10.1038/ng.2874
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2874
This article is cited by
-
RNA binding protein IGF2BP1 synergizes with ETV6-RUNX1 to drive oncogenic signaling in B-cell Acute Lymphoblastic Leukemia
Journal of Experimental & Clinical Cancer Research (2023)
-
Evolutionary determinants of curability in cancer
Nature Ecology & Evolution (2023)
-
Conserved transcription factors promote cell fate stability and restrict reprogramming potential in differentiated cells
Nature Communications (2023)
-
Covert pre-leukaemic clones in healthy co-twins of patients with childhood acute lymphoblastic leukaemia
Leukemia (2023)
-
Clonal origin of KMT2A wild-type lineage-switch leukemia following CAR-T cell and blinatumomab therapy
Nature Cancer (2023)