Abstract
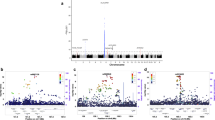
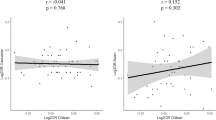
We used a candidate gene approach to identify a set of SNPs, located in a predicted regulatory region on chromosome 1q44 downstream of NLRP3 (previously known as CIAS1 and NALP3) that are associated with Crohn's disease. The associations were consistently replicated in four sample sets from individuals of European descent. In the combined analysis of all samples (710 father-mother-child trios, 239 cases and 107 controls), these SNPs were strongly associated with risk of Crohn's disease (Pcombined = 3.49 × 10−9, odds ratio = 1.78, confidence interval = 1.47–2.16 for rs10733113), reaching a level consistent with the stringent significance thresholds imposed by whole-genome association studies. In addition, we observed significant associations between SNPs in the associated regions and NLRP3 expression and IL-1β production. Mutations in NLRP3 are known to be responsible for three rare autoinflammatory disorders1,2. These results suggest that the NLRP3 region is also implicated in the susceptibility of more common inflammatory diseases such as Crohn's disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Agostini, L. et al. NALP3 forms an IL-1beta-processing inflammasome with increased activity in Muckle-Wells autoinflammatory disorder. Immunity 20, 319–325 (2004).
Mariathasan, S. & Monack, D.M. Inflammasome adaptors and sensors: intracellular regulators of infection and inflammation. Nat. Rev. Immunol. 7, 31–40 (2007).
Podolsky, D.K. Inflammatory bowel disease. N. Engl. J. Med. 347, 417–429 (2002).
Ting, J.P., Kastner, D.L. & Hoffman, H.M. CATERPILLERs, pyrin and hereditary immunological disorders. Nat. Rev. Immunol. 6, 183–195 (2006).
Pétrilli, V., Dostert, C., Muruve, D.A. & Tschopp, J. The inflammasome: a danger sensing complex triggering innate immunity. Curr. Opin. Immunol. 19, 615–622 (2007).
Martinon, F. & Tschopp, J. Inflammatory caspases and inflammasomes: master switches of inflammation. Cell Death Differ. 14, 10–22 (2007).
Taylor, J. et al. ESPERR: learning strong and weak signals in genomic sequence alignments to identify functional elements. Genome Res. 16, 1596–1604 (2006).
de Bakker, P.I. et al. Efficiency and power in genetic association studies. Nat. Genet. 37, 1217–1223 (2005).
Kummer, J.A. et al. Inflammasome components NALP 1 and 3 show distinct but separate expression profiles in human tissues suggesting a site-specific role in the inflammatory response. J. Histochem. Cytochem. 55, 443–452 (2007).
Martinon, F., Agostini, L., Meylan, E. & Tschopp, J. Identification of bacterial muramyl dipeptide as activator of the NALP3/cryopyrin inflammasome. Curr. Biol. 14, 1929–1934 (2004).
Hawkins, P.N., Lachmann, H.J. & McDermott, M.F. Interleukin-1-receptor antagonist in the Muckle-Wells syndrome. N. Engl. J. Med. 348, 2583–2584 (2003).
Hoffman, H.M. et al. Prevention of cold-associated acute inflammation in familial cold autoinflammatory syndrome by interleukin-1 receptor antagonist. Lancet 364, 1779–1785 (2004).
Li, J. et al. Regulation of IL-8 and IL-1beta expression in Crohn's disease associated NOD2/CARD15 mutations. Hum. Mol. Genet. 13, 1715–1725 (2004).
Van Heel, D.A. et al. Muramyl dipeptide and toll-like receptor sensitivity in NOD2-associated Crohn's disease. Lancet 365, 1794–1796 (2005).
Kramer, M., Netea, M.G., de Jong, D.J., Kullberg, B.J. & Adema, G.J. Impaired dendritic cell function in Crohn's disease patients with NOD2 3020insC mutation. J. Leukoc. Biol. 79, 860–866 (2006).
van Beelen, A.J. et al. Stimulation of the intracellular bacterial sensor NOD2 programs dendritic cells to promote interleukin-17 production in human memory T cells. Immunity 27, 660–669 (2007).
Duerr, R.H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006).
Rioux, J.D. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nat. Genet. 39, 596–604 (2007).
Libioulle, C. et al. Novel Crohn disease locus identified by genome-wide association maps to a gene desert on 5p13.1 and modulates expression of PTGER4. PLoS Genet. 3, e58 (2007).
Parkes, M. et al. Sequence variants in the autophagy gene IRGM and multiple other replicating loci contribute to Crohn's disease susceptibility. Nat. Genet. 39, 830–832 (2007).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Barrett, J.C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat. Genet. 40, 955–962 (2008).
Lennard-Jones, J.E. Classification of inflammatory bowel disease. Scand. J. Gastroenterol. Suppl. 170, 2–6 (1989).
Silverberg, M.S. et al. Toward an integrated clinical, molecular and serological classification of inflammatory bowel disease: Report of a Working Party of the 2005 Montreal World Congress of Gastroenterology. Can. J. Gastroenterol. (Suppl A), 19, 5–36 (2005).
Bell, P.A. et al. SNPstream UHT: ultra-high throughput SNP genotyping for pharmacogenomics and drug discovery. Biotechniques Suppl 70–72, 74, 76–77 (2002).
van den Boom, D. & Ehrich, M. Discovery and identification of sequence polymorphisms and mutations with MALDI-TOF MS. Methods Mol. Biol. 366, 287–306 (2007).
Barrett, J.C., Fry, B., Maller, J. & Daly, M.J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Dudbridge, F. Likelihood-based association analysis for nuclear families and unrelated subjects with missing genotype data. Hum. Hered. 66, 87–98 (2008).
Spielman, R.S., McGinnis, R.E. & Ewens, W.J. Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am. J. Hum. Genet. 52, 506–516 (1993).
Holland, P.M., Abramson, R.D., Watson, R. & Gelfand, D.H. Detection of specific polymerase chain reaction product by utilizing the 5′→3′ exonuclease activity of Thermus aquaticus DNA polymerase. Proc. Natl. Acad. Sci. USA 88, 7276–7280 (1991).
Acknowledgements
We thank the study participants and their families for their collaboration; A. Montpetit and the genotyping and sequencing platforms of the McGill University and Genome Québec Innovation Centre for their technical assistance; T. McKenzie, G. Bonventi and C. Landolt-Maticorena for their help with acquisition and preparation of the PBCs; F. Martinon and J. Tschopp for their technical assistance in designing the functional studies; and T. Kwan for helpful discussion. This research was supported by grants to D.F. from the Canada Foundation for Innovation, the Canada Research Chair program, the Research Institute of the McGill University Health Centre and the Crohn's & Colitis Foundation of Canada (CCFC). A.-C.V. is supported by a PhD training award from the Fonds de la recherche en santé du Québec. M.S.S. is funded by the National Institute of Diabetes and Digestive and Kidney Diseases (DK624230), the CCFC and the Mount Sinai Hospital Gale and Graham Wright Research Chair in Digestive Diseases. J.D.R. is funded by grants from the National Institutes of Allergy and Infectious Diseases (AI065687 and AI067152), the National Institute of Diabetes and Digestive and Kidney Diseases (DK064869 and DK062432) and the Crohn's and Colitis Foundation of America (SRA512). E.L. is a senior research associate at the Belgium Foundation for Scientific Research (FNRS). S.V. is a clinical investigator for the Funds for Scientific Research of Flanders. T.J.H. is the recipient of an Investigator Award from the Ontario Institute for Cancer Research through funding from the Government of Ontario's Ministry of Research and Innovation. D.F. is a senior clinical scientist at the Belgium FNRS. None of the funding agencies had any role in the design and conduct of the study; in the collection, analysis and interpretation of the data; or in the preparation, review or approval of the manuscript.
Author information
Authors and Affiliations
Contributions
A.-C.V., G.F. and D.F. conceived and designed the experiments. A.-C.V., G.F., C.C. and N.B. did the experiments. A.-C.V. and M.L. analyzed the data. E.L., M.S.S., C.L., J.B., A.B., D.G., A.C., D.L., P.R.F., J.E.W., M.S., P.R., J.D.R., S.V., T.J.H. and D.F. provided study samples. A.-C.V., M.L. and D.F. wrote the paper, with contributions from all authors.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Tables 1–6 and Supplementary Figures 1 and 2 (PDF 3222 kb)
Rights and permissions
About this article
Cite this article
Villani, AC., Lemire, M., Fortin, G. et al. Common variants in the NLRP3 region contribute to Crohn's disease susceptibility. Nat Genet 41, 71–76 (2009). https://doi.org/10.1038/ng.285
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.285
This article is cited by
-
ERAP1 is a critical regulator of inflammasome-mediated proinflammatory and ER stress responses
BMC Immunology (2022)
-
Systems biology approach highlights mechanistic differences between Crohn’s disease and ulcerative colitis
Scientific Reports (2021)
-
Epithelial sensing of microbiota-derived signals
Genes & Immunity (2021)
-
A case of cryopyrin-associated periodic fever syndrome during canakinumab administration complicated by inflammatory bowel disease
Clinical Rheumatology (2021)
-
Genotype–phenotype associations of polymorphisms within the gene locus of NOD-like receptor pyrin domain containing 3 in Swiss inflammatory bowel disease patients
BMC Gastroenterology (2021)