Abstract
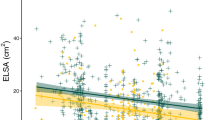
With the increased availability of high-resolution sequence information, genome-wide association (GWA) studies have become feasible in a number of species1,2,3,4,5,6,7,8. The vast majority of these studies are conducted in human populations, where it is difficult to provide strong evidence for the functional involvement of unknown genes that are identified using GWA. Here we used the model organism Arabidopsis thaliana to combine high-throughput confocal microscopy imaging of traits at the cellular level, GWA and expression analyses to identify genomic regions that are associated with developmental cell–type traits. We identify and characterize a new F-box gene, KUK, that regulates meristem and cell length. We further show that polymorphisms in the coding sequence are the major causes of KUK allele–dependent natural variation in root development. This work demonstrates the feasibility of GWA using cellular traits to identify causal genes for basic biological processes such as development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
26 November 2013
In the version of this article initially published online, there was an error in the final paragraph of the main text. Specifically, the phrase "previously known regulators" should have appeared as "previously unknown regulators." The error has been corrected for the print, PDF and HTML versions of this article.
References
Parker, C.C., Sokoloff, G., Cheng, R. & Palmer, A.A. Genome-wide association for fear conditioning in an advanced intercross mouse line. Behav. Genet. 42, 437–448 (2012).
Jordan, K.W. et al. Genome-wide association for sensitivity to chronic oxidative stress in Drosophila melanogaster. PLoS ONE 7, e38722 (2012).
Mu, J. et al. Plasmodium falciparum genome-wide scans for positive selection, recombination hot spots and resistance to antimalarial drugs. Nat. Genet. 42, 268–271 (2010).
Tian, F. et al. Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat. Genet. 43, 159–162 (2011).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Gregersen, V.R. et al. Genome-wide association scan and phased haplotype construction for quantitative trait loci affecting boar taint in three pig breeds. BMC Genomics 13, 22 (2012).
Atwell, S. et al. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465, 627–631 (2010).
Klein, R.J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Li, Y., Huang, Y., Bergelson, J., Nordborg, M. & Borevitz, J.O. Association mapping of local climate-sensitive quantitative trait loci in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 107, 21199–21204 (2010).
Chao, D.Y. et al. Genome-wide association studies identify heavy metal ATPase3 as the primary determinant of natural variation in leaf cadmium in Arabidopsis thaliana. PLoS Genet. 8, e1002923 (2012).
Baxter, I. et al. A coastal cline in sodium accumulation in Arabidopsis thaliana is driven by natural variation of the sodium transporter AtHKT1;1. PLoS Genet. 6, e1001193 (2010).
Ideker, T., Dutkowski, J. & Hood, L. Boosting signal-to-noise in complex biology: prior knowledge is power. Cell 144, 860–863 (2011).
Brady, S.M. et al. A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318, 801–806 (2007).
Busch, W. et al. A microfluidic device and computational platform for high-throughput live imaging of gene expression. Nat. Methods 9, 1101–1106 (2012).
Horton, M.W. et al. Genome-wide patterns of genetic variation in worldwide Arabidopsis thaliana accessions from the RegMap panel. Nat. Genet. 44, 212–216 (2012).
Kang, H.M. et al. Efficient control of population structure in model organism association mapping. Genetics 178, 1709–1723 (2008).
Yu, J. et al. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat. Genet. 38, 203–208 (2006).
Seren, Ü. et al. GWAPP: a web application for genome-wide association mapping in Arabidopsis. Plant Cell 24, 4793–4805 (2012).
Brachi, B. et al. Linkage and association mapping of Arabidopsis thaliana flowering time in nature. PLoS Genet. 6, e1000940 (2010).
Skowyra, D., Craig, K.L., Tyers, M., Elledge, S.J. & Harper, J.W. F-box proteins are receptors that recruit phosphorylated substrates to the SCF ubiquitin-ligase complex. Cell 91, 209–219 (1997).
Bai, C. et al. SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box. Cell 86, 263–274 (1996).
Birnbaum, K. et al. A gene expression map of the Arabidopsis root. Science 302, 1956–1960 (2003).
Dello Ioio, R. et al. A genetic framework for the control of cell division and differentiation in the root meristem. Science 322, 1380–1384 (2008).
Mouchel, C.F., Briggs, G.C. & Hardtke, C.S. Natural genetic variation in Arabidopsis identifies BREVIS RADIX, a novel regulator of cell proliferation and elongation in the root. Genes Dev. 18, 700–714 (2004).
Alonso, J.M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Murray, M.G. & Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4325 (1980).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Hu, T.T. et al. The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nat. Genet. 43, 476–481 (2011).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Günther, T. & Schmid, K.J. Improved haplotype-based detection of ongoing selective sweeps towards an application in Arabidopsis thaliana. BMC Res. Notes 4, 232 (2011).
Gautier, M. & Vitalis, R. rehh: an R package to detect footprints of selection in genome-wide SNP data from haplotype structure. Bioinformatics 28, 1176–1177 (2012).
Hellens, R.P., Edwards, E.A., Leyland, N.R., Bean, S. & Mullineaux, P.M. pGreen: a versatile and flexible binary Ti vector for Agrobacterium-mediated plant transformation. Plant Mol. Biol. 42, 819–832 (2000).
Weigel, D. & Glazebrook, J. In planta transformation of Arabidopsis. CSH Protoc. 2006, pii: pdb.prot4668 (2006).
Ruxton, G.D. The unequal variance t-test is an underused alternative to Student's t-test and the Mann-Whitney U test. Behav. Ecol. 17, 688–690 (2006).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J. R. Stat. Soc., B 57, 289–300 (1995).
Acknowledgements
We are grateful for assistance in the GWA analysis and access to software from A. Korte, Ü. Seren, B. Vilhjalmsson and M. Nordborg and for technical assistance from B. Wohlrab and C. Göschl. We thank P. Benfey, T. Greb, M. Nordborg, L. Valledor, A. Korte, D. Filiault and members of the Busch laboratory for valuable discussions and critical reading of the manuscript. We thank V. Nizhynska, P. Korte and M. Nordborg (Gregor Mendel Institute, Vienna, Austria) for donating seeds for natural accession, S. Waidmann, P. Sánchez and J. Agustí (Gregor Mendel Institute, Vienna, Austria) for materials and advice on vectors and cloning and T. Friese for manuscript editing. This research was supported by funds from the Austrian Academy of Science through the Gregor Mendel Institute and a European Molecular Biology Organization (EMBO) long-term fellowship to T.T.
Author information
Authors and Affiliations
Contributions
T.T. carried out the population genetic analyses. S.B.S. performed the experiments with the transgenic KUK lines. M.M. performed the phenotyping, trait quantification, qRT-PCRs and cloning. W.B. conducted KUK-YFP reporter line analyses. W.B. and M.M. conceived the experiments, analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–12 (PDF 11087 kb)
Source data
Rights and permissions
About this article
Cite this article
Meijón, M., Satbhai, S., Tsuchimatsu, T. et al. Genome-wide association study using cellular traits identifies a new regulator of root development in Arabidopsis. Nat Genet 46, 77–81 (2014). https://doi.org/10.1038/ng.2824
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2824
This article is cited by
-
The receptor kinase SRF3 coordinates iron-level and flagellin dependent defense and growth responses in plants
Nature Communications (2022)
-
Association mapping of drought tolerance and agronomic traits in rice (Oryza sativa L.) landraces
BMC Plant Biology (2021)
-
Meta-QTLs, ortho-MQTLs and candidate genes for nitrogen use efficiency and root system architecture in bread wheat (Triticum aestivum L.)
Physiology and Molecular Biology of Plants (2021)
-
Natural variation of BSK3 tunes brassinosteroid signaling to regulate root foraging under low nitrogen
Nature Communications (2019)
-
Natural variation among Arabidopsis thaliana accessions in tolerance to high magnesium supply
Scientific Reports (2018)