Abstract
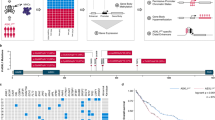
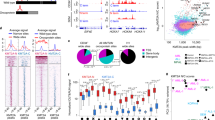
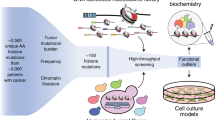
Epigenetic dysregulation is an emerging hallmark of cancers. We developed a high-information-content mass spectrometry approach to profile global histone modifications in human cancers. When applied to 115 lines from the Cancer Cell Line Encyclopedia1, this approach identified distinct molecular chromatin signatures. One signature was characterized by increased histone 3 lysine 36 (H3K36) dimethylation, exhibited by several lines harboring translocations in NSD2, which encodes a methyltransferase. A previously unknown NSD2 p.Glu1099Lys (p.E1099K) variant was identified in nontranslocated acute lymphoblastic leukemia (ALL) cell lines sharing this signature. Ectopic expression of the variant induced a chromatin signature characteristic of NSD2 hyperactivation and promoted transformation. NSD2 knockdown selectively inhibited the proliferation of NSD2-mutant lines and impaired the in vivo growth of an NSD2-mutant ALL xenograft. Sequencing analysis of >1,000 pediatric cancer genomes identified the NSD2 p.E1099K alteration in 14% of t(12;21) ETV6-RUNX1–containing ALLs. These findings identify NSD2 as a potential therapeutic target for pediatric ALL and provide a general framework for the functional annotation of cancer epigenomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Peach, S.E., Rudomin, E.L., Udeshi, N.D., Carr, S.A. & Jaffe, J.D. Quantitative assessment of chromatin immunoprecipitation grade antibodies directed against histone modifications reveals patterns of co-occurring marks on histone protein molecules. Mol. Cell. Proteomics 11, 128–137 (2012).
Ryan, R.J. & Bernstein, B.E. Molecular biology. Genetic events that shape the cancer epigenome. Science 336, 1513–1514 (2012).
Morin, R.D. et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476, 298–303 (2011).
Shih, A.H., Abdel-Wahab, O., Patel, J.P. & Levine, R.L. The role of mutations in epigenetic regulators in myeloid malignancies. Nat. Rev. Cancer 12, 599–612 (2012).
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012).
McCabe, M.T. et al. Mutation of A677 in histone methyltransferase EZH2 in human B-cell lymphoma promotes hypertrimethylation of histone H3 on lysine 27 (H3K27). Proc. Natl. Acad. Sci. USA 109, 2989–2994 (2012).
Sneeringer, C.J. et al. Coordinated activities of wild-type plus mutant EZH2 drive tumor-associated hypertrimethylation of lysine 27 on histone H3 (H3K27) in human B-cell lymphomas. Proc. Natl. Acad. Sci. USA 107, 20980–20985 (2010).
Yap, D.B. et al. Somatic mutations at EZH2 Y641 act dominantly through a mechanism of selectively altered PRC2 catalytic activity, to increase H3K27 trimethylation. Blood 117, 2451–2459 (2011).
Ernst, T. et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat. Genet. 42, 722–726 (2010).
Malgeri, U. et al. Detection of t(4;14)(p16.3;q32) chromosomal translocation in multiple myeloma by reverse transcription-polymerase chain reaction analysis of IGH-MMSET fusion transcripts. Cancer Res. 60, 4058–4061 (2000).
Chesi, M. et al. The t(4;14) translocation in myeloma dysregulates both FGFR3 and a novel gene, MMSET, resulting in IgH/MMSET hybrid transcripts. Blood 92, 3025–3034 (1998).
Martinez-Garcia, E. et al. The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood 117, 211–220 (2011).
Kuo, A.J. et al. NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol. Cell 44, 609–620 (2011).
Li, Y. et al. The target of the NSD family of histone lysine methyltransferases depends on the nature of the substrate. J. Biol. Chem. 284, 34283–34295 (2009).
Qiao, Q. et al. The structure of NSD1 reveals an autoregulatory mechanism underlying histone H3K36 methylation. J. Biol. Chem. 286, 8361–8368 (2011).
Lauring, J. et al. The multiple myeloma associated MMSET gene contributes to cellular adhesion, clonogenic growth, and tumorigenicity. Blood 111, 856–864 (2008).
Mullighan, C.G. et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 446, 758–764 (2007).
Mullighan, C.G. et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 322, 1377–1380 (2008).
Seligson, D.B. et al. Global levels of histone modifications predict prognosis in different cancers. Am. J. Pathol. 174, 1619–1628 (2009).
Kurdistani, S.K. Histone modifications as markers of cancer prognosis: a cellular view. Br. J. Cancer 97, 1–5 (2007).
Chervona, Y. & Costa, M. Histone modifications and cancer: biomarkers of prognosis? Am. J. Cancer Res. 2, 589–597 (2012).
McCabe, M.T. et al. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature 492, 108–112 (2012).
Qi, W. et al. Selective inhibition of Ezh2 by a small molecule inhibitor blocks tumor cells proliferation. Proc. Natl. Acad. Sci. USA 109, 21360–21365 (2012).
Kim, J.Y. et al. Multiple-myeloma-related WHSC1/MMSET isoform RE-IIBP is a histone methyltransferase with transcriptional repression activity. Mol. Cell. Biol. 28, 2023–2034 (2008).
Marango, J. et al. The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood 111, 3145–3154 (2008).
Thomas, C.E., Kelleher, N.L. & Mizzen, C.A. Mass spectrometric characterization of human histone H3: a bird's eye view. J. Proteome Res. 5, 240–247 (2006).
Garcia, B.A. et al. Chemical derivatization of histones for facilitated analysis by mass spectrometry. Nat. Protoc. 2, 933–938 (2007).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Wang, J. et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nat. Methods 8, 652–654 (2011).
Kiefer, F., Arnold, K., Kunzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL Repository and associated resources. Nucleic Acids Res. 37, D387–D392 (2009).
Schapira, M. Structural chemistry of human SET domain protein methyltransferases. Curr. Chem. Genomics 5, 85–94 (2011).
Acknowledgements
We thank R. Pagliarini and R. Tiedt for their critical comments on the manuscript, and the NIBR-BROAD CCLE team for their guidance and contributions to the CCLE project. KMS11-TKO cells were generously provided by Horizon Discovery Ltd., UK. We are grateful for the technical assistance of X. Liu. We thank the members of the St. Jude Children's Research Hospital Washington University Pediatric Cancer Genome Project and the American Lebanese Syrian Associated Charities (ALSAC) of St. Jude Children's Research Hospital for funding. The global chromatin profiling project was enabled by a grant from the Novartis Institutes for Biomedical Research. Additional funding support was provided by the National Cancer Institute and the U.S. Department of Defense (L.A.G.), the Pew Scholars Program in the Biomedical Sciences (C.M.) and the St. Baldrick's Foundation Scholar Award (C.M.).
Author information
Authors and Affiliations
Contributions
Y.W., N.P.E., F.Y., H.M.C. and J.L. performed or directed cellular assays data generation. M.H. and Z.W. performed or directed biochemical experiments. E.R.M. III, R.deB., N.P.E. and Y.W. performed or directed nucleic acid extraction and targeted sequencing of NSD2. J.Z., L.W., J.M., J.E. and C.M. identified the NSD2 somatic mutations in tumor samples. J.Z., L.W., J.M., G.V.K. and K.V. performed bioinformatic analysis. J.E.T. performed MS experiments. Z.Y. and R.H. performed structural modeling. J.D.J., Y.W., H.M.C., M.H., R.M., R.deB., J.L., S.A.C. and F.S. designed experiments and analyzed biological data. Y.W., V.G.C., H.C.B. and V.G. performed or directed tumor xenograft studies. J.D.J., Y.W., H.M.C., M.H., E.R.M. III., K.V., J.Z., L.W. and R.H. prepared figures and tables. J.D.J., H.M.C., J.Z., J.R.D., W.R.S., L.A.G. and F.S. wrote and edited the main text and Supplementary Information. R.S., J.B., G.C., W.R.S. and N.K. contributed to overall project oversight and advising; F.S., J.R.D. and L.A.G. provided overall project leadership.
Corresponding authors
Ethics declarations
Competing interests
Y.W., H.M.C., H.C.B., M.H., N.P.E., F.Y., Z.W., E.R.M. III, Z.Y., R.deB., V.G., K.V., R.S., W.R.S., N.K., J.L., G.C., J.B., V.G.C. and F.S. are employees of Novartis, Inc., as noted in the affiliations. L.A.G. consults for and holds equity in Foundation Medicine and consults for and receives research sponsorship from Novartis, Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–7 (PDF 1697 kb)
Rights and permissions
About this article
Cite this article
Jaffe, J., Wang, Y., Chan, H. et al. Global chromatin profiling reveals NSD2 mutations in pediatric acute lymphoblastic leukemia. Nat Genet 45, 1386–1391 (2013). https://doi.org/10.1038/ng.2777
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2777
This article is cited by
-
Investigating pathological epigenetic aberrations by epi-proteomics
Clinical Epigenetics (2022)
-
A chemical probe targeting the PWWP domain alters NSD2 nucleolar localization
Nature Chemical Biology (2022)
-
Identification of histone methyltransferase NSD2 as an important oncogenic gene in colorectal cancer
Cell Death & Disease (2021)
-
Nuclear organization and regulation of the differentiated state
Cellular and Molecular Life Sciences (2021)
-
Proteomic profiling dataset of chemical perturbations in multiple biological backgrounds
Scientific Data (2021)