Abstract
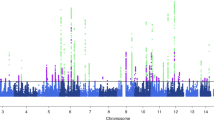
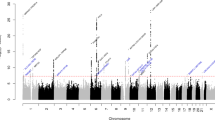
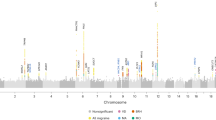
Migraine is the most common brain disorder, affecting approximately 14% of the adult population, but its molecular mechanisms are poorly understood. We report the results of a meta-analysis across 29 genome-wide association studies, including a total of 23,285 individuals with migraine (cases) and 95,425 population-matched controls. We identified 12 loci associated with migraine susceptibility (P < 5 × 10−8). Five loci are new: near AJAP1 at 1p36, near TSPAN2 at 1p13, within FHL5 at 6q16, within C7orf10 at 7p14 and near MMP16 at 8q21. Three of these loci were identified in disease subgroup analyses. Brain tissue expression quantitative trait locus analysis suggests potential functional candidate genes at four loci: APOA1BP, TBC1D7, FUT9, STAT6 and ATP5B.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Anttila, V. et al. Genome-wide association study of migraine implicates a common susceptibility variant on 8q22.1. Nat. Genet. 42, 869–873 (2010).
Chasman, D.I. et al. Genome-wide association study reveals three susceptibility loci for common migraine in the general population. Nat. Genet. 43, 695–698 (2011).
Ligthart, L. et al. Meta-analysis of genome-wide association for migraine in six population-based European cohorts. Eur. J. Hum. Genet. 19, 901–907 (2011).
Freilinger, T. et al. Genome-wide association analysis identifies susceptibility loci for migraine without aura. Nat. Genet. 44, 777–782 (2012).
International Headache Society. The International Classification of Headache Disorders: 2nd edition. Cephalalgia 24 (suppl. 1), 9–160 (2004).
Fimia, G.M., De Cesare, D. & Sassone-Corsi, P. A family of LIM-only transcriptional coactivators: tissue-specific expression and selective activation of CREB and CREM. Mol. Cell. Biol. 20, 8613–8622 (2000).
Dash, P.K., Hochner, B. & Kandel, E.R. Injection of the cAMP-responsive element into the nucleus of Aplysia sensory neurons blocks long-term facilitation. Nature 345, 718–721 (1990).
Lee, Y.S. & Silva, A.J. The molecular and cellular biology of enhanced cognition. Nat. Rev. Neurosci. 10, 126–140 (2009).
Sherman, E.A. et al. Genetic mapping of glutaric aciduria, type 3, to chromosome 7 and identification of mutations in c7orf10. Am. J. Hum. Genet. 83, 604–609 (2008).
Schreiner, A. et al. Junction protein shrew-1 influences cell invasion and interacts with invasion-promoting protein CD147. Mol. Biol. Cell 18, 1272–1281 (2007).
Lafleur, M.A., Xu, D. & Hemler, M.E. Tetraspanin proteins regulate membrane type-1 matrix metalloproteinase–dependent pericellular proteolysis. Mol. Biol. Cell 20, 2030–2040 (2009).
Rozanov, D.V., Hahn-Dantona, E., Strickland, D.K. & Strongin, A.Y. The low density lipoprotein receptor–related protein LRP is regulated by membrane type-1 matrix metalloproteinase (MT1-MMP) proteolysis in malignant cells. J. Biol. Chem. 279, 4260–4268 (2004).
Borrie, S.C., Baeumer, B.E. & Bandtlow, C.E. The Nogo-66 receptor family in the intact and diseased CNS. Cell Tissue Res. 349, 105–117 (2012).
Schürks, M. et al. Migraine and cardiovascular disease: systematic review and meta-analysis. Br. Med. J. 339, b3914 (2009).
Mizuguchi, T. et al. Heterozygous TGFBR2 mutations in Marfan syndrome. Nat. Genet. 36, 855–860 (2004).
Biros, E., Walker, P.J., Nataatmadja, M., West, M. & Golledge, J. Downregulation of transforming growth factor, β receptor 2 and Notch signaling pathway in human abdominal aortic aneurysm. Atherosclerosis 221, 383–386 (2012).
Kathiresan, S. et al. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat. Genet. 41, 334–341 (2009).
Terada, N. et al. The tetraspanin protein, CD9, is expressed by progenitor cells committed to oligodendrogenesis and is linked to β1 integrin, CD81, and Tspan-2. Glia 40, 350–359 (2002).
Flavell, S.W. et al. Genome-wide analysis of MEF2 transcriptional program reveals synaptic target genes and neuronal activity–dependent polyadenylation site selection. Neuron 60, 1022–1038 (2008).
Tsuzuki, K., Xing, H., Ling, J. & Gu, J.G. Menthol-induced Ca2+ release from presynaptic Ca2+ stores potentiates sensory synaptic transmission. J. Neurosci. 24, 762–771 (2004).
Diniz, L.P. et al. Astrocyte-induced synaptogenesis is mediated by transforming growth factor β signaling through modulation of D-serine levels in cerebral cortex neurons. J. Biol. Chem. 287, 41432–41445 (2012).
Allen, P.B., Greenfield, A.T., Svenningsson, P., Haspeslagh, D.C. & Greengard, P. Phactrs 1–4: s family of protein phosphatase 1 and actin regulatory proteins. Proc. Natl. Acad. Sci. USA 101, 7187–7192 (2004).
Ferraro, G.B., Morrison, C.J., Overall, C.M., Strittmatter, S.M. & Fournier, A.E. Membrane-type matrix metalloproteinase-3 regulates neuronal responsiveness to myelin through Nogo-66 receptor 1 cleavage. J. Biol. Chem. 286, 31418–31424 (2011).
Wilson, P.M., Fryer, R.H., Fang, Y. & Hatten, M.E. Astn2, a novel member of the astrotactin gene family, regulates the trafficking of ASTN1 during glial-guided neuronal migration. J. Neurosci. 30, 8529–8540 (2010).
May, P. et al. Neuronal LRP1 functionally associates with postsynaptic proteins and is required for normal motor function in mice. Mol. Cell. Biol. 24, 8872–8883 (2004).
Jha, K.N. et al. Biochemical and structural characterization of apolipoprotein A-I binding protein, a novel phosphoprotein with a potential role in sperm capacitation. Endocrinology 149, 2108–2120 (2008).
Gouveia, R. et al. Expression of glycogenes in differentiating human NT2N neurons. Downregulation of fucosyltransferase 9 leads to decreased Lewisx levels and impaired neurite outgrowth. Biochim. Biophys. Acta 1820, 2007–2019 (2012).
Lieberoth, A. et al. Lewisx and α2,3-sialyl glycans and their receptors TAG-1, Contactin, and L1 mediate CD24-dependent neurite outgrowth. J. Neurosci. 29, 6677–6690 (2009).
Nishihara, S. α1,3-fucosyltransferase IX (Fut9) determines Lewis X expression in brain. Glycobiology 13, 445–455 (2003).
Lawrence, T. & Natoli, G. Transcriptional regulation of macrophage polarization: enabling diversity with identity. Nat. Rev. Immunol. 11, 750–761 (2011).
Park, S.J. et al. Astrocytes, but not microglia, rapidly sense H2O2 via STAT6 phosphorylation, resulting in cyclooxygenase-2 expression and prostaglandin release. J. Immunol. 188, 5132–5141 (2012).
Dibble, C.C. et al. TBC1D7 is a third subunit of the TSC1-TSC2 complex upstream of mTORC1. Mol. Cell 47, 535–546 (2012).
Han, J.M. & Sahin, M. TSC1/TSC2 signaling in the CNS. FEBS Lett. 585, 973–980 (2011).
Russell, M.B. & Olesen, J. Increased familial risk and evidence of genetic factor in migraine. BMJ 311, 541–544 (1995).
Frazer, K.A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Altshuler, D., Daly, M.J. & Lander, E.S. Genetic mapping in human disease. Science 322, 881–888 (2008).
Hindorff, L.A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc., B 57, 289–300 (1995).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Magi, R., Lindgren, C.M. & Morris, A.P. Meta-analysis of sex-specific genome-wide association studies. Genet. Epidemiol. 34, 846–853 (2010).
Mägi, R. & Morris, A.P. GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11, 288 (2010).
Segrè, A.V., Groop, L., Mootha, V.K., Daly, M.J. & Altshuler, D. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Lage, K. et al. A human phenome-interactome network of protein complexes implicated in genetic disorders. Nat. Biotechnol. 25, 309–316 (2007).
Lage, K. et al. A large-scale analysis of tissue-specific pathology and gene expression of human disease genes and complexes. Proc. Natl. Acad. Sci. USA 105, 20870–20875 (2008).
Rossin, E.J. et al. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 7, e1001273 (2011).
Raychaudhuri, S. et al. Identifying relationships among genomic disease regions: predicting genes at pathogenic SNP associations and rare deletions. PLoS Genet. 5, e1000534 (2009).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Boyle, A.P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Li, C. & Wong, W.H. Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc. Natl. Acad. Sci. USA 98, 31–36 (2001).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Acknowledgements
Study-specific acknowledgments appear in the Supplementary Note. We wish to thank A. Coffey, S. Hunt, R. Gwillian, P. Whittaker, S. Potter and A. Tashakkori-Ghanbarian for their invaluable help with this study, and we collectively thank everyone who has contributed to the collection, genotyping and analysis of the individual cohorts, as well as all the study participants.
Author information
Authors and Affiliations
Consortia
Contributions
A. Palotie, D.I.C., D.R.N., M.J.D., K.S., G.D.S., M.W., M.D., A.M.J.M.v.d.M., M.D.F., C.K., T.K., D.P.S., L.C., J.-A.Z., M.-R.J., C.v.D., D.I.B., J.K., L.Q. and G.T. jointly supervised research. A. Palotie, D.I.C., D.R.N., V. Anttila, B.S.W., M.J.D., M.W., M.D., A.M.J.M.v.d.M., C.K., L.C., J.-A.Z., M.-R.J., C.v.D., D.I.B., J.K., T.K., M. Kallela, R.M., B.d.V., G.T., L.Q., M.A.I., L.L., E.H., M.S., H.S., K.S., F.J., T.F. and B.M.-M. conceived and designed the study. V. Anttila, B.S.W., A. Palotie, D.R.N., G.D.S., M.W., M.D., A.M.J.M.v.d.M., C.K., J.-A.Z., M.-R.J., O.R., C.v.D., D.I.B., J.K., E.B., M. Kallela, B.d.V., G.T., E.H., T.F., R.R.F., N.G.M., A.G.U., T.M. and J.G.E. performed the experiments. V. Anttila, B.S.W., P.G., D.I.C., D.R.N., M.J.D., D.P.S., E.B., J.R.G., S.B.R.J., T.K., F.B., G.M., R.M., B.d.V., L.Q., M.A.I., L.L., I.D., P.P., M.S., S. Steinberg, T.F. and B.M.-M. performed statistical analysis. V. Anttila, B.S.W., P.G., A. Palotie, D.I.C., D.R.N., L.C., J.R.G., S.B.R.J., K.L., T.K., F.B., G.M., R.M., B.d.V., S.E.M., L.Q., M.A.I., L.L., J.W., P.P., M.S., S. Steinberg, H.S., T.F., N.A., B.M.-M. and D.T. analyzed the data. D.I.C., D.R.N., M.J.D., A. Palotie, G.D.S., M.W., M.D., A.M.J.M.v.d.M., M.D.F., C.K., D.P.S., L.C., J.-A.Z., M.-R.J., O.R., C.v.D., D.I.B., B.W.P., J.K., E.B., J.R.G., K.L., E.R., V. Anttila, B.S.W., P.G., T.K., F.B., G.M., M. Kallela, R.M., B.d.V., G.T., U.T., W.L.M., L.Q., M. Koiranen, M.A.I., T.L., A.H.S., L.L., I.D., B.M.N., M.S., L.M.R., J.E.B., P.M.R., S. Steinberg, H.S., F.J., D.A.L., D.M.E., S.M.R., M.F., V. Artto, M.A.K., T.F., J.S., R.R.F., N.P., C.M.W., R.Z., A.C.H., P.A.F.M., G.W.M., N.G.M., G.B., H.G., A. Heinze, K.H.-K., F.M.K.W., A.-L.H., A. Pouta, J.v.d.E., A.G.U., A. Hofman, J.-J.H., J.M.V., K.H., M.A., B.M.-M., S. Schreiber, T.M., H.E.W., A.A., J.G.E., B.J.T. and D.T. contributed reagents and/or materials and/or analysis tools. V. Anttila, B.S.W., A. Palotie, D.I.C., D.R.N., A.M.J.M.v.d.M. and C.K. wrote the manuscript. All authors contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–12 and Supplementary Note (PDF 3257 kb)
Rights and permissions
About this article
Cite this article
Anttila, V., Winsvold, B., Gormley, P. et al. Genome-wide meta-analysis identifies new susceptibility loci for migraine. Nat Genet 45, 912–917 (2013). https://doi.org/10.1038/ng.2676
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2676
This article is cited by
-
Galcanezumab effects on incidence of headache after occurrence of triggers, premonitory symptoms, and aura in responders, non-responders, super-responders, and super non-responders
The Journal of Headache and Pain (2023)
-
Proteomics profiling reveals mitochondrial damage in the thalamus in a mouse model of chronic migraine
The Journal of Headache and Pain (2023)
-
Genetics of migraine: where are we now?
The Journal of Headache and Pain (2023)
-
Current Update on Categorization of Migraine Subtypes on the Basis of Genetic Variation: a Systematic Review
Molecular Neurobiology (2023)
-
A Meta-Analysis of the Genome-Wide Association Studies on Two Genetically Correlated Phenotypes Suggests Four New Risk Loci for Headaches
Phenomics (2023)