Abstract
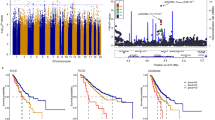
Genome-wide association studies (GWAS) have identified four susceptibility loci for epithelial ovarian cancer (EOC), with another two suggestive loci reaching near genome-wide significance. We pooled data from a GWAS conducted in North America with another GWAS from the UK. We selected the top 24,551 SNPs for inclusion on the iCOGS custom genotyping array. We performed follow-up genotyping in 18,174 individuals with EOC (cases) and 26,134 controls from 43 studies from the Ovarian Cancer Association Consortium. We validated the two loci at 3q25 and 17q21 that were previously found to have associations close to genome-wide significance and identified three loci newly associated with risk: two loci associated with all EOC subtypes at 8q21 (rs11782652, P = 5.5 × 10−9) and 10p12 (rs1243180, P = 1.8 × 10−8) and another locus specific to the serous subtype at 17q12 (rs757210, P = 8.1 × 10−10). An integrated molecular analysis of genes and regulatory regions at these loci provided evidence for functional mechanisms underlying susceptibility and implicated CHMP4C in the pathogenesis of ovarian cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lichtenstein, P. et al. Environmental and heritable factors in the causation of cancer—analyses of cohorts of twins from Sweden, Denmark and Finland. N. Engl. J. Med. 343, 78–85 (2000).
Stratton, J.F., Pharoah, P., Smith, S.K., Easton, D. & Ponder, B.A. A systematic review and meta-analysis of family history and risk of ovarian cancer. Br. J. Obstet. Gynaecol. 105, 493–499 (1998).
Antoniou, A.C. & Easton, D.F. Risk prediction models for familial breast cancer. Future Oncol. 2, 257–274 (2006).
Song, H. et al. A genome-wide association study identifies a new ovarian cancer susceptibility l ocus on 9p22.2. Nat. Genet. 41, 996–1000 (2009).
Bolton, K.L. et al. Common variants at 19p13 are associated with susceptibility to ovarian cancer. Nat. Genet. 42, 880–884 (2010).
Goode, E.L. et al. A genome-wide association study identifies susceptibility loci for ovarian cancer at 2q31 and 8q24. Nat. Genet. 42, 874–879 (2010).
Shen, H. et al. Epigenetic analysis leads to identification of HNF1B as a subtype-specific susceptibility gene for ovarian cancer. Nat. Comm. published online; doi:10.1038/ncomms2629 (27 March 2013).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Jia, L. et al. Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet. 5, e1000597 (2009).
Kim, M.J. et al. Functional characterization of liver enhancers that regulate drug-associated transporters. Clin. Pharmacol. Ther. 89, 571–578 (2011).
Wasserman, N.F., Aneas, I. & Nobrega, M.A. An 8q24 gene desert variant associated with prostate cancer risk confers differential in vivo activity to a MYC enhancer. Genome Res. 20, 1191–1197 (2010).
Wright, J.B., Brown, S.J. & Cole, M.D. Upregulation of c-MYC in cis through a large chromatin loop linked to a cancer risk-associated single-nucleotide polymorphism in colorectal cancer cells. Mol. Cell Biol. 30, 1411–1420 (2010).
McCullough, J., Fisher, R.D., Whitby, F.G., Sundquist, W.I. & Hill, C.P. ALIX-CHMP4 interactions in the human ESCRT pathway. Proc. Natl. Acad. Sci. USA 105, 7687–7691 (2008).
Carlton, J.G., Caballe, A., Agromayor, M., Kloc, M. & Martin-Serrano, J. ESCRT-III governs the Aurora B-mediated abscission checkpoint through CHMP4C. Science 336, 220–225 (2012).
Yu, X., Riley, T. & Levine, A.J. The regulation of the endosomal compartment by p53 the tumor suppressor gene. FEBS J. 276, 2201–2212 (2009).
Nikolova, D.N. et al. Genome-wide gene expression profiles of ovarian carcinoma: identification of molecular targets for the treatment of ovarian carcinoma. Mol. Med. Report 2, 365–384 (2009).
Caudell, D. & Aplan, P.D. The role of CALM-AF10 gene fusion in acute leukemia. Leukemia 22, 678–685 (2008).
Cóser, V.M. et al. Nebulette is the second member of the nebulin family fused to the MLL gene in infant leukemia. Cancer Genet. Cytogenet. 198, 151–154 (2010).
Ram, R. & Blaxall, B.C. Nebulette mutations in cardiac remodeling: big effects from a small mechanosensor. J. Am. Coll. Cardiol. 56, 1503–1505 (2010).
Salzman, J. et al. ESRRA-C11orf20 is a recurrent gene fusion in serous ovarian carcinoma. PLoS Biol. 9, e1001156 (2011).
Voight, B.F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 42, 579–589 (2010).
Spurdle, A.B. et al. Genome-wide association study identifies a common variant associated with risk of endometrial cancer. Nat. Genet. 43, 451–454 (2011).
Elliott, K.S. et al. Evaluation of association of HNF1B variants with diverse cancers: collaborative analysis of data from 19 genome-wide association studies. PLoS ONE 5, e10858 (2010).
Kato, N., Sasou, S. & Motoyama, T. Expression of hepatocyte nuclear factor-1β (HNF-1β) in clear cell tumors and endometriosis of the ovary. Mod. Pathol. 19, 83–89 (2006).
Tsuchiya, A. et al. Expression profiling in ovarian clear cell carcinoma: identification of hepatocyte nuclear factor-1β as a molecular marker and a possible molecular target for therapy of ovarian clear cell carcinoma. Am. J. Pathol. 163, 2503–2512 (2003).
Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Kato, N. & Motoyama, T. Hepatocyte nuclear factor-1β (HNF-1β) in human urogenital organs: its expression and role in embryogenesis and tumorigenesis. Histol. Histopathol. 24, 1479–1486 (2009).
Hindorff, L.A., Junkins, H.A., Hall, P.A., Mehta, J.P. & Manolio, T.A. A catalogue of published genome-wide association studies. Genome.gov: National Institutes of Health National Human Genome Research Institute. <http://www.genome.gov/gwastudies/> (2013).
Antoniou, A.C. et al. A locus on 19p13 modifies risk of breast cancer in BRCA1 mutation carriers and is associated with hormone receptor-negative breast cancer in the general population. Nat. Genet. 42, 885–892 (2010).
Michailidou, K. et al. Large-scale genotyping identifies 41 new breast cancer susceptibility loci. Nat. Genet. published online; doi:10.1038/ng.2563 (27 March 2013).
Lawrenson, K. et al. Senescent fibroblasts promote neoplastic transformation of partially transformed ovarian epithelial cells in a three-dimensional model of early stage ovarian cancer. Neoplasia 12, 317–325 (2010).
Permuth-Wey, J. et al. LIN28B polymorphisms influence susceptibility to epithelial ovarian cancer. Cancer Res. 71, 3896–3903 (2011).
Kermani, B.G. Artificial intelligence and global normalization methods for genotyping. US patent 7,035,740 (2008).
Teo, Y.Y. et al. A genotype calling algorithm for the Illumina BeadArray platform. Bioinformatics 23, 2741–2746 (2007).
Giannoulatou, E., Yau, C., Colella, S., Ragoussis, J. & Holmes, C.C. GenoSNP: a variational Bayes within-sample SNP genotyping algorithm that does not require a reference population. Bioinformatics 24, 2209–2214 (2008).
Sankararaman, S., Sridhar, S., Kimmel, G. & Halperin, E. Estimating local ancestry in admixed populations. Am. J. Hum. Genet. 82, 290–303 (2008).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Xing, G., Lin, C.Y., Wooding, S.P. & Xing, C. Blindly using Wald's test can miss rare disease-causal variants in case-control association studies. Ann. Hum. Genet. 76, 168–177 (2012).
Breslow, N.E. & Day, N.E. Statistical Methods in Cancer Research. Volume 1—The Analysis of Case-Control Studies (International Agency for Research on Cancer, Lyon, 1980).
Forbes, S.A. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945–D950 (2011).
Mostafavi, S., Ray, D., Warde-Farley, D., Grouios, C. & Morris, Q. GeneMANIA: a real-time multiple association network integration algorithm for predicting gene function. Genome Biol. 9 (suppl. 1), S4 (2008).
Heintzman, N.D. et al. Histone modifications at human enhancers reflect global cell-type–specific gene expression. Nature 459, 108–112 (2009).
Acknowledgements
We thank all the individuals who took part in this study and all the researchers, clinicians and technical and administrative staff who made possible the many studies contributing to this work (a full list is provided in the Supplementary Note). The COGS project is funded through a European Commission's Seventh Framework Programme grant (agreement number 223175 - HEALTH-F2-2009-223175). The Ovarian Cancer Association Consortium is supported by a grant from the Ovarian Cancer Research Fund thanks to donations by the family and friends of Kathryn Sladek Smith (PPD/RPCI.07). The scientific development and funding for this project were supported in part by the Genetic Associations and Mechanisms in Oncology (GAME-ON) and a National Cancer Institute Cancer Post-GWAS Initiative (U19-CA148112). Details of the funding of individual investigators and studies are provided in the Supplementary Note. This study made use of data generated by the Wellcome Trust Case Control consortium; funding for the project was provided by the Wellcome Trust under award 076113. A full list of the investigators who contributed to the generation of the data is available from the website (see URLs). The results published here are based in part on data generated by The Cancer Genome Atlas Pilot Project established by the National Cancer Institute and National Human Genome Research Institute; information about The Cancer Genome Atlas (TCGA) and the investigators and institutions who constitute the TCGA research network can be found on the website (see URLs).
Author information
Authors and Affiliations
Consortia
Contributions
Writing group: P.D.P.P., Y.-Y.T., C.M.P., S.J.R., J.M.S., T.A.S., B.L.F., E.L.G., A.N.A.M. and S.A.G. All authors read and approved the final manuscript. Provision of samples and data from contributing studies: K.L., M.P., J.P.T., H. Shen, R.W., R.K., M.C.L., H. Song, D.C.T., F.B., D.V., J.M.C., J.D., E. Dicks, K.K.A., H.A.-C., N.A., S.M.A., L.B., E.V.B., M.W.B., M.J.B., G.B., N.B., J.D.B., L.A.B., A.B.-W., R. Brown, R. Butzow, I.C., M.E.C., R.S.C., J.C.-C., Y.A.C., Z.C., A.D.-M., E. Despierre, J.A.D., T.D., A.d.B., M.D., D.E., R.E., A.B.E., P.A.F., D.F., J.F., Y.-T.G., M.G.-C., A.G.-M., G.G., A.G., M.G., J.G., Q.G., M.K.H., P. Harter, A.H., F.H., P. Hillemanns, M.H., E.H., C.K.H., S.H., A. Jakubowska, A. Jensen, K.R.K., B.Y.K., L.E.K., L.A.K., S.K.K., G.E.K., C.K., J.K., D.L., S.L., N.D.L., N.L., J. Lee, A.L., B.K.L., J. Lissowska, J. Lubinński, L.L., G.L., L.F.A.G.M., K.M., V.M., J.R.M., U.M., F.M., K.B.M., T.N., S.A.N., R.B.N., H. Nevanlinna, S.N., H. Noushmehr, K.O., S.O., I.O., J.P., T.P., L.M.P., J.P.-W., M.C.P., E.M.P., X.Q., H.A.R., L.R.-R., M.A.R., A.R., I.R., I.K.R., H.B.S., I.S., G.S., H. Shen, V.S., X.-O.S., W.S., M.C.S., P.S., K.T., S.-H.T., K.L.T., P.J.T., A.T., S.S.T., A.M.v.A., D.v.d.B., I.V., R.A.V., A.F.V., S.W.-G., N.W., A.S.W., E.W., B.W., Y.L.W., A.H.W., H.P.Y., W.Z., A.Z., F.Z., M.T.G., P. Hall, D.F.E., C.L.P., A.B., G.C.-T., E.I. and J.M.S. Bioinformatics and data management: J.D., E. Dicks, Z.C. and R.W. Data analysis: J.P.T., Q.G., Y.-Y.T. and B.L.F. Preparation of samples for genotyping: S.J.R. and C.M.P. Genotyping: J.M.C., D.C.T., F.B. and D.V. Functional analyses: S.A.G., M.B., A.N.A.M., B.L.F., K.L., H. Shen, E.L.G., S.J.R., Y.A.C. and M.L.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–5, Supplementary Figures 1–13 and Supplementary Note (PDF 43425 kb)
Rights and permissions
About this article
Cite this article
Pharoah, P., Tsai, YY., Ramus, S. et al. GWAS meta-analysis and replication identifies three new susceptibility loci for ovarian cancer. Nat Genet 45, 362–370 (2013). https://doi.org/10.1038/ng.2564
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2564
This article is cited by
-
Extensive germline-somatic interplay contributes to prostate cancer progression through HNF1B co-option of TMPRSS2-ERG
Nature Communications (2022)
-
CLIMB: High-dimensional association detection in large scale genomic data
Nature Communications (2022)
-
Considering hormone-sensitive cancers as a single disease in the UK biobank reveals shared aetiology
Communications Biology (2022)
-
Fine-mapping and association analysis of candidate genes for papilla number in sea cucumber, Apostichopus japonicus
Marine Life Science & Technology (2022)
-
Constructing germline research cohorts from the discarded reads of clinical tumor sequences
Genome Medicine (2021)