Abstract
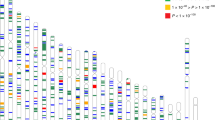
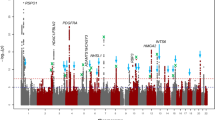
Refractive error is the most common eye disorder worldwide and is a prominent cause of blindness. Myopia affects over 30% of Western populations and up to 80% of Asians. The CREAM consortium conducted genome-wide meta-analyses, including 37,382 individuals from 27 studies of European ancestry and 8,376 from 5 Asian cohorts. We identified 16 new loci for refractive error in individuals of European ancestry, of which 8 were shared with Asians. Combined analysis identified 8 additional associated loci. The new loci include candidate genes with functions in neurotransmission (GRIA4), ion transport (KCNQ5), retinoic acid metabolism (RDH5), extracellular matrix remodeling (LAMA2 and BMP2) and eye development (SIX6 and PRSS56). We also confirmed previously reported associations with GJD2 and RASGRF1. Risk score analysis using associated SNPs showed a tenfold increased risk of myopia for individuals carrying the highest genetic load. Our results, based on a large meta-analysis across independent multiancestry studies, considerably advance understanding of the mechanisms involved in refractive error and myopia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
09 May 2013
In the version of this article initially published, the affiliations of Daniel W.H. Ho were incorrect, and the spelling of Sarayut Janmahasatian in the author list was incorrect. The errors have been corrected in the HTML and PDF versions of this article.
References
Wojciechowski, R. Nature and nurture: the complex genetics of myopia and refractive error. Clin. Genet. 79, 301–320 (2011).
Mutti, D.O. et al. Axial growth and changes in lenticular and corneal power during emmetropization in infants. Invest. Ophthalmol. Vis. Sci. 46, 3074–3080 (2005).
Smith, T.S., Frick, K.D., Holden, B.A., Fricke, T.R. & Naidoo, K.S. Potential lost productivity resulting from the global burden of uncorrected refractive error. Bull. World Health Organ. 87, 431–437 (2009).
Solouki, A.M. et al. A genome-wide association study identifies a susceptibility locus for refractive errors and myopia at 15q14. Nat. Genet. 42, 897–901 (2010).
Shi, Y. et al. Genetic variants at 13q12.12 are associated with high myopia in the Han Chinese population. Am. J. Hum. Genet. 88, 805–813 (2011).
Nakanishi, H. et al. A genome-wide association analysis identified a novel susceptible locus for pathological myopia at 11q24.1. PLoS Genet. 5, e1000660 (2009).
Li, Z. et al. A genome-wide association study reveals association between common variants in an intergenic region of 4q25 and high-grade myopia in the Chinese Han population. Hum. Mol. Genet. 20, 2861–2868 (2011).
Li, Y.J. et al. Genome-wide association studies reveal genetic variants in CTNND2 for high myopia in Singapore Chinese. Ophthalmology 118, 368–375 (2011).
Hysi, P.G. et al. A genome-wide association study for myopia and refractive error identifies a susceptibility locus at 15q25. Nat. Genet. 42, 902–905 (2010).
Fan, Q. et al. Genetic variants on chromosome 1q41 influence ocular axial length and high myopia. PLoS Genet. 8, e1002753 (2012).
Yang, J. et al. Genomic inflation factors under polygenic inheritance. Eur. J. Hum. Genet. 19, 807–812 (2011).
Verhoeven, V.J. et al. Large scale international replication and meta-analysis study confirms association of the 15q14 locus with myopia. The CREAM consortium. Hum. Genet. 131, 1467–1480 (2012).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Estrada, K. et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat. Genet. 44, 491–501 (2012).
ENCODE Project Consortium.. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Rymer, J. & Wildsoet, C.F. The role of the retinal pigment epithelium in eye growth regulation and myopia: a review. Vis. Neurosci. 22, 251–261 (2005).
Fernández-Medarde, A. et al. RasGRF1 disruption causes retinal photoreception defects and associated transcriptomic alterations. J. Neurochem. 110, 641–652 (2009).
Tonini, R. et al. Expression of Ras-GRF in the SK-N-BE neuroblastoma accelerates retinoic-acid-induced neuronal differentiation and increases the functional expression of the IRK1 potassium channel. Eur. J. Neurosci. 11, 959–966 (1999).
Abd-El-Barr, M.M. et al. Genetic dissection of rod and cone pathways in the dark-adapted mouse retina. J. Neurophysiol. 102, 1945–1955 (2009).
Beyer, B. et al. Absence seizures in C3H/HeJ and knockout mice caused by mutation of the AMPA receptor subunit Gria4. Hum. Mol. Genet. 17, 1738–1749 (2008).
Connaughton, V. Glutamate and glutamate receptors in the vertebrate retina. in Webvision: The Organization of the Retina and Visual System (eds. Kolb, H., Fernandez, E. & Nelson, R.) (Natural Library of Medicine, Salt Lake City, Utah, 1995).
Yang, J., Nemargut, J.P. & Wang, G.Y. The roles of ionotropic glutamate receptors along the On and Off signaling pathways in the light-adapted mouse retina. Brain Res. 1390, 70–79 (2011).
Smith, E.L. III, Fox, D.A. & Duncan, G.C. Refractive-error changes in kitten eyes produced by chronic on-channel blockade. Vision Res. 31, 833–844 (1991).
Fogel, B.L. et al. RBFOX1 regulates both splicing and transcriptional networks in human neuronal development. Hum. Mol. Genet. 21, 4171–4186 (2012).
Zhang, X., Yang, D. & Hughes, B.A. KCNQ5/Kv7.5 potassium channel expression and subcellular localization in primate retinal pigment epithelium and neural retina. Am. J. Physiol. Cell Physiol. 301, C1017–C1026 (2011).
Pattnaik, B.R. & Hughes, B.A. Effects of KCNQ channel modulators on the M-type potassium current in primate retinal pigment epithelium. Am. J. Physiol. Cell Physiol. 302, C821–C833 (2012).
Troilo, D., Nickla, D.L., Mertz, J.R. & Summers Rada, J.A. Change in the synthesis rates of ocular retinoic acid and scleral glycosaminoglycan during experimentally altered eye growth in marmosets. Invest. Ophthalmol. Vis. Sci. 47, 1768–1777 (2006).
Mertz, J.R. & Wallman, J. Choroidal retinoic acid synthesis: a possible mediator between refractive error and compensatory eye growth. Exp. Eye Res. 70, 519–527 (2000).
McFadden, S.A., Howlett, M.H. & Mertz, J.R. Retinoic acid signals the direction of ocular elongation in the guinea pig eye. Vision Res. 44, 643–653 (2004).
Parker, R.O. & Crouch, R.K. Retinol dehydrogenases (RDHs) in the visual cycle. Exp. Eye Res. 91, 788–792 (2010).
Schéele, S. et al. Laminin isoforms in development and disease. J. Mol. Med. (Berl.) 85, 825–836 (2007).
Zhang, Y., Liu, Y. & Wildsoet, C.F. Bidirectional, optical sign-dependent regulation of BMP2 gene expression in chick retinal pigment epithelium. Invest. Ophthalmol. Vis. Sci. 53, 6072–6080 (2012).
Ramdas, W.D. et al. A genome-wide association study of optic disc parameters. PLoS Genet. 6, e1000978 (2010).
Gallardo, M.E. et al. Analysis of the developmental SIX6 homeobox gene in patients with anophthalmia/microphthalmia. Am. J. Med. Genet. A. 129A, 92–94 (2004).
Gal, A. et al. Autosomal-recessive posterior microphthalmos is caused by mutations in PRSS56, a gene encoding a trypsin-like serine protease. Am. J. Hum. Genet. 88, 382–390 (2011).
Orr, A. et al. Mutations in a novel serine protease PRSS56 in families with nanophthalmos. Mol. Vis. 17, 1850–1861 (2011).
Nair, K.S. et al. Alteration of the serine protease PRSS56 causes angle-closure glaucoma in mice and posterior microphthalmia in humans and mice. Nat. Genet. 43, 579–584 (2011).
Rossin, E.J. et al. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 7, e1001273 (2011).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Aulchenko, Y.S., Struchalin, M.V. & van Duijn, C.M. ProbABEL package for genome-wide association analysis of imputed data. BMC Bioinformatics 11, 134 (2010).
Chen, W.M. & Abecasis, G.R. Family-based association tests for genomewide association scans. Am. J. Hum. Genet. 81, 913–926 (2007).
Mägi, R. & Morris, A.P. GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11, 288 (2010).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Booij, J.C. et al. Functional annotation of the human retinal pigment epithelium transcriptome. BMC Genomics 10, 164 (2009).
Edgar, R., Domrachev, M. & Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
van Soest, S.S. et al. Comparison of human retinal pigment epithelium gene expression in macula and periphery highlights potential topographic differences in Bruch's membrane. Mol. Vis. 13, 1608–1617 (2007).
Hotelling, H. The generalization of Student's ratio. Ann. Math. Stat. 2, 360–378 (1931).
Irizarry, R.A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Booij, J.C. et al. A new strategy to identify and annotate human RPE-specific gene expression. PLoS One 5, e9341 (2010).
Acknowledgements
We gratefully thank the invaluable contributions of all study participants, their relatives and staff at the recruitment centers. Complete funding information and acknowledgments by study can be found in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
V.J.M.V., P.G.H., R.W., C.J.H., C.C.W.K., A.W.H., D.A.M., T.L.Y. and C.M.v.D. performed analyses and drafted the manuscript. C.C.W.K., D.S., C.J.H., J.E.B.-W., S.-M.S., C.M.v.D., A.H., D.A.M., S.M., A.D.P., V.V., C.W., P.N.B., T.-Y.W., J.S.R., T.L.Y., K.O., O. Pärssinen, S.P.Y., J.A.G., A. Metspalu, M.P., S.K.I. and N.P. jointly conceived the project and supervised the work. J.E.B.W., S.-M.S., D.A.M., T.L.Y., C.J.H., C.C.W.K., D.S., J.E.B.-W., C.M.v.D., R.W., P.G.H., V.J.M.V., K.O., Y.-Y.T., T.-Y.W., P.N.B., V.V., N.A., B.A.O., A.H., J.R.V., F.R., A.G.U., N.P., C.M., A. Mirshahi, T.Z., B.F., J.F.W., Z.V., O. Polasek, A.F.W., C.H., I.R., S.K.I., E.C., J.H.L., R.P.I., S.J., M.S., J.J.W., P.M., I.C., J.S.R., P.M.C., C.E.P., G.W.M., A. Mishra, W.A., F.M., M.P., L.C.K., T.D.S., E.Y.-D., A.N., O.R., C.-C.K., T.M., A.D., R.T.O., Y.Z., J.L., R.L., P.C., V.A.B., W.-T.T., E.V., T.A., E.-S.T., A. Metspalu, T.H., R.K., B.E.K.K., J.E.C., K.P.B., L.J.C., C.P.P., D.W.H.H., S.P.Y., J.W., O. Pärssinen, J.B.J., L.X., H.S.W., S.M.H., A.D.P., M.K., T.L., K.-M.M., C.L.S., C.W., N.J.T., D.M.E., B.S.P., J.P.K., G.M., G.H.S.B., M.K.I., X.Z., C.-Y.C., A.W.H., S.M., R.H., J.A.G. and Q.F. were responsible for study-specific data. G.H.S.B., V.J.M.V., Q.F. and J.A.G. were involved in the genetic risk score analysis. T.L.Y., A.A.B.B., T.G.M.F.G. and F.H. performed the data expression experiments. A.A.B.B., T.G.M.F.G., A.M. and S.M. were involved in pathway analyses. J.E.B.-W., S.-M.S., D.A.M., T.L.Y., K.O., T.-Y.W., P.N.B., T.G.M.F.G., S.K.I., E.C., J.J.W., A.J.M.H.V., C.-C.K., B.E.K.K., S.P.Y., C.W., N.J.T., G.H.S.B., M.K.I., A.W.H. and J.A.G. critically reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members appears in the Supplementary Note.
A full list of members appears in the Supplementary Note.
A full list of members appears in the Supplementary Note.
A full list of members appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–4, Supplementary Figures 1–5 and Supplementary Note (PDF 4615 kb)
Rights and permissions
About this article
Cite this article
Verhoeven, V., Hysi, P., Wojciechowski, R. et al. Genome-wide meta-analyses of multiancestry cohorts identify multiple new susceptibility loci for refractive error and myopia. Nat Genet 45, 314–318 (2013). https://doi.org/10.1038/ng.2554
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2554
This article is cited by
-
Impact of text contrast polarity on the retinal activity in myopes and emmetropes using modified pattern ERG
Scientific Reports (2023)
-
Targeting the cochlin/SFRP1/CaMKII axis in the ocular posterior pole prevents the progression of nonpathologic myopia
Communications Biology (2023)
-
Post-GWAS screening of candidate genes for refractive error in mutant zebrafish models
Scientific Reports (2023)
-
Overlapping pathogenic de novo CNVs in neurodevelopmental disorders and congenital anomalies impacting constraint genes regulating early development
Human Genetics (2023)
-
Rare variant analyses across multiethnic cohorts identify novel genes for refractive error
Communications Biology (2023)