Abstract
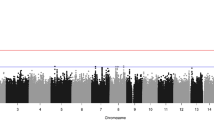
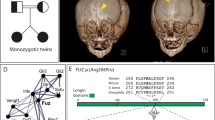
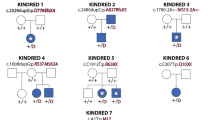
Craniosynostosis, the premature fusion of the cranial sutures, is a heterogeneous disorder with a prevalence of ∼1 in 2,200 (refs. 1,2). A specific genetic etiology can be identified in ∼21% of cases3, including mutations of TWIST1, which encodes a class II basic helix-loop-helix (bHLH) transcription factor, and causes Saethre-Chotzen syndrome, typically associated with coronal synostosis4,5,6. Using exome sequencing, we identified 38 heterozygous TCF12 mutations in 347 samples from unrelated individuals with craniosynostosis. The mutations predominantly occurred in individuals with coronal synostosis and accounted for 32% and 10% of subjects with bilateral and unilateral pathology, respectively. TCF12 encodes one of three class I E proteins that heterodimerize with class II bHLH proteins such as TWIST1. We show that TCF12 and TWIST1 act synergistically in a transactivation assay and that mice doubly heterozygous for loss-of-function mutations in Tcf12 and Twist1 have severe coronal synostosis. Hence, the dosage of TCF12-TWIST1 heterodimers is critical for normal coronal suture development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Primary accessions
NCBI Reference Sequence
Referenced accessions
Protein Data Bank
Change history
05 September 2013
In the version of this article initially published, numbering and spacing for the exon structure of TCF12 in Figure 2a was incorrect. The error has been corrected in the HTML and PDF versions of the article.
References
Lajeunie, E., Le Merrer, M., Bonaïti-Pellie, C., Marchac, D. & Renier, D. Genetic study of nonsyndromic coronal craniosynostosis. Am. J. Med. Genet. 55, 500–504 (1995).
Boulet, S.L., Rasmussen, S.A. & Honein, M.A. A population-based study of craniosynostosis in metropolitan Atlanta, 1989–2003. Am. J. Med. Genet. 146A, 984–991 (2008).
Wilkie, A.O.M. et al. Prevalence and complications of single-gene and chromosomal disorders in craniosynostosis. Pediatrics 126, e391–e400 (2010).
Howard, T.D. et al. Mutations in TWIST, a basic helix-loop-helix transcription factor, in Saethre-Chotzen syndrome. Nat. Genet. 15, 36–41 (1997).
El Ghouzzi, V. et al. Mutations of the TWIST gene in the Saethre-Chotzen syndrome. Nat. Genet. 15, 42–46 (1997).
Jabs, E.W. TWIST and the Saethre-Chotzen syndrome. in Inborn Errors of Development. The Molecular Basis of Clinical Disorders of Morphogenesis, 2nd edn (eds. Epstein, C.J., Erickson, R.P. & Wynshaw-Boris, A.) 474–481 (Oxford University Press, Oxford, 2008).
Ng, S.B. et al. Exome sequencing identifies the cause of a mendelian disorder. Nat. Genet. 42, 30–35 (2010).
Johnson, D. & Wilkie, A.O.M. Craniosynostosis. Eur. J. Hum. Genet. 19, 369–376 (2011).
Hu, J.S., Olson, E.N. & Kingston, R.E. HEB, a helix-loop-helix protein related to E2A and ITF2, that can modulate the DNA-binding ability of myogenic regulatory factors. Mol. Cell Biol. 12, 1031–1042 (1992).
Nielsen, A.L., Pallisgaard, N., Pedersen, F.S. & Jørgensen, P. Murine helix-loop-helix transcriptional activator proteins binding to the E-box motif of the Akv murine leukemia-virus enhancer identified by cDNA cloning. Mol. Cell Biol. 12, 3449–3459 (1992).
Gan, T.-I. Genomic organization of human TCF12 gene and spliced mRNA variants producing isoforms of transcription factor HTF4. Cytogenet. Genome Res. 98, 245–248 (2002).
Hiraki, Y. et al. Craniosynostosis in a patient with a de novo 15q15-q22 deletion. Am. J. Med. Genet. 146A, 1462–1465 (2008).
Wang, D. et al. The basic helix-loop-helix transcription factor HEBAlt is expressed in pro-T cells and enhances the generation of T cell precursors. J. Immunol. 177, 109–119 (2006).
Whalen, S. et al. Novel comprehensive diagnostic strategy in Pitt-Hopkins syndrome: clinical score and further delineation of the TCF4 mutational spectrum. Hum. Mutat. 33, 64–72 (2012).
Laursen, K.B., Mielke, E., Iannaccone, P. & Fuchtbauer, E.M. Mechanism of transcriptional activation by the proto-oncogene Twist1. J. Biol. Chem. 282, 34623–34633 (2007).
Wojciechowski, J., Lai, A., Kondo, M. & Zhuang, Y. E2A and HEB are required to block thymocyte proliferation prior to pre-TCR expression. J. Immunol. 178, 5717–5726 (2007).
Connerney, J. et al. Twist1 dimer selection regulates cranial suture patterning and fusion. Dev. Dyn. 235, 1345–1357 (2006).
Lakso, M. et al. Efficient in vivo manipulation of mouse genomic sequences at the zygote stage. Proc. Natl. Acad. Sci. USA 93, 5860–5865 (1996).
Chen, Z.F. & Behringer, R.R. twist is required in head mesenchyme for cranial neural tube morphogenesis. Genes Dev. 9, 686–699 (1995).
Merrill, A.E. et al. Cell mixing at a neural crest–mesoderm boundary and deficient ephrin-Eph signaling in the pathogenesis of craniosynostosis. Hum. Mol. Genet. 15, 1319–1328 (2006).
Chai, Y. & Maxson, R.E. Jr. Recent advances in craniofacial morphogenesis. Dev. Dyn. 235, 2353–2375 (2006).
Bialek, P. et al. A Twist code determines the onset of osteoblast differentiation. Dev. Cell 6, 423–435 (2004).
Hayashi, M. et al. Comparative roles of Twist-1 and Id1 in transcriptional regulation by BMP signaling. J. Cell Sci. 120, 1350–1357 (2007).
Aronheim, A., Shiran, R., Rosen, A. & Walker, M.D. The E2A gene-product contains two separable and functionally distinct transcription activation domains. Proc. Natl. Acad. Sci. USA 90, 8063–8067 (1993).
Markus, M., Du, Z.M. & Benezra, R. Enhancer-specific modulation of E protein activity. J. Biol. Chem. 277, 6469–6477 (2002).
Longo, A., Guanga, G.P. & Rose, R.B. Crystal structure of E47-NeuroD1/β2 bHLH domain-DNA complex: heterodimer selectivity and DNA recognition. Biochemistry 47, 218–229 (2008).
Acknowledgements
We thank all the families for their participation, S. Butler for cell culture, J. Frankland and T. Rostron for DNA sequencing, S. Knight for coordinating array–comparative genomic hybridization (aCGH), L. Gregory and the High-Throughput Genomics core at the Wellcome Trust Centre for Human Genetics for exome sequencing, R. Evans for review of anesthetic records, W. Baggley for clinical photography, A. van den Ouweland for genetic testing, E.-M. Füchtbauer (Aarhus University) for constructs and Y. Zhuang (Duke University) for the gift of the Tcf12flox mutant. This work was funded by the National Institute for Health Research (NIHR) Biomedical Research Centre Oxford (V.P.S. and R.J.C.), the Oxford University Clinical Academic Graduate School and the Oxfordshire Health Services Research Committee (V.P.S.), the Oxford Craniofacial Unit Charitable Fund (V.P.S.), the Thames Valley Comprehensive Local Research Network (J.M.P.), The Dutch Center for Translational Molecular Medicine (P.J.v.d.S.), the Carolien Bijl Foundation (J.A.C.G.), the US National Institutes of Health (NIH; R01DE016320 and R01DE019650 to R.E.M.) and the Wellcome Trust (093329 to S.R.F.T. and A.O.M.W.).
Author information
Authors and Affiliations
Consortia
Contributions
S.R.F.T., R.E.M. and A.O.M.W. conceived the project. V.P.S., A.L.F., M.S.B. and S.R.F.T. performed experimental analyses. S.B. and R.J.C. performed immune function tests. S.J.M., J.B., A.K. and WGS500 coordinated or performed bioinformatics analyses. V.P.S., J.A.C.G., A.J.M.H., A.F.B., N.O.J., S.A.L., J.B.M., D.J.M., J.M.P., E.S., S.E.T., L.C.W., D.J., S.A.W., P.J.v.d.S., I.M.J.M. and A.O.M.W. recruited samples from subjects and collected clinical information. V.P.S., A.L.F., R.E.M., S.R.F.T. and A.O.M.W. drafted the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of members and affiliations is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–6 and Supplementary Tables 1–6 (PDF 2360 kb)
Rights and permissions
About this article
Cite this article
Sharma, V., Fenwick, A., Brockop, M. et al. Mutations in TCF12, encoding a basic helix-loop-helix partner of TWIST1, are a frequent cause of coronal craniosynostosis. Nat Genet 45, 304–307 (2013). https://doi.org/10.1038/ng.2531
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2531
This article is cited by
-
The CMS19 disease model specifies a pivotal role for collagen XIII in bone homeostasis
Scientific Reports (2022)
-
Craniofacial morphology and growth in Muenke syndrome, Saethre-Chotzen syndrome, and TCF12-related craniosynostosis
Clinical Oral Investigations (2022)
-
MACF1 promotes osteoblast differentiation by sequestering repressors in cytoplasm
Cell Death & Differentiation (2021)
-
Approaches to characterize the transcriptional trajectory of human myogenesis
Cellular and Molecular Life Sciences (2021)
-
Temporal transcriptome analysis of neuronal commitment reveals the preeminent role of the divergent lncRNA biotype and a critical candidate gene during differentiation
Cell Death Discovery (2020)