Abstract
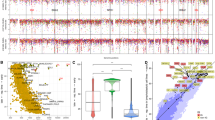
Burkitt lymphoma is a mature aggressive B-cell lymphoma derived from germinal center B cells1. Its cytogenetic hallmark is the Burkitt translocation t(8;14)(q24;q32) and its variants, which juxtapose the MYC oncogene with one of the three immunoglobulin loci2. Consequently, MYC is deregulated, resulting in massive perturbation of gene expression3. Nevertheless, MYC deregulation alone seems not to be sufficient to drive Burkitt lymphomagenesis. By whole-genome, whole-exome and transcriptome sequencing of four prototypical Burkitt lymphomas with immunoglobulin gene (IG)-MYC translocation, we identified seven recurrently mutated genes. One of these genes, ID3, mapped to a region of focal homozygous loss in Burkitt lymphoma4. In an extended cohort, 36 of 53 molecularly defined Burkitt lymphomas (68%) carried potentially damaging mutations of ID3. These were strongly enriched at somatic hypermutation motifs. Only 6 of 47 other B-cell lymphomas with the IG-MYC translocation (13%) carried ID3 mutations. These findings suggest that cooperation between ID3 inactivation and IG-MYC translocation is a hallmark of Burkitt lymphomagenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
Ensembl
NCBI Reference Sequence
Protein Data Bank
References
Swerdlow, S.H. et al. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, Ch. 10, 179–268 (IARC Press, Lyon, France, 2008).
Boerma, E.G., Siebert, R., Kluin, P.M. & Baudis, M. Translocations involving 8q24 in Burkitt lymphoma and other malignant lymphomas: a historical review of cytogenetics in the light of today's knowledge. Leukemia 23, 225–234 (2009).
Klapproth, K. & Wirth, T. Advances in the understanding of MYC-induced lymphomagenesis. Br. J. Haematol. 149, 484–497 (2010).
Scholtysik, R. et al. Detection of genomic aberrations in molecularly defined Burkitt′s lymphoma by array-based, high resolution, single nucleotide polymorphism analysis. Haematologica 95, 2047–2055 (2010).
Hummel, M. et al. A biologic definition of Burkitt′s lymphoma from transcriptional and genomic profiling. N. Engl. J. Med. 354, 2419–2430 (2006).
Dave, S.S. et al. Molecular diagnosis of Burkitt′s lymphoma. N. Engl. J. Med. 354, 2431–2442 (2006).
Schmitz, R. et al. Recurrent oncogenic mutations in CCND3 in aggressive lymphomas. Blood (ASH Annual Meeting Abstracts) 118, 435 (2011).
Wilda, M. et al. Inactivation of the ARF–MDM-2–p53 pathway in sporadic Burkitt′s lymphoma in children. Leukemia 18, 584–588 (2004).
Johnston, J.M. & Carroll, W.L. c-myc hypermutation in Burkitt′s lymphoma. Leuk. Lymphoma 8, 431–439 (1992).
Love, C.L. et al. Whole genome and exome sequencing reveals the genetic landscape of Burkitt lymphoma. Blood (ASH Annual Meeting Abstracts) 118, 433 (2011).
Duan, S. et al. FBXO11 targets BCL6 for degradation and is inactivated in diffuse large B-cell lymphomas. Nature 481, 90–93 (2012).
Wang, L. et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N. Engl. J. Med. 365, 2497–2506 (2011).
Klapper, W. et al. Molecular profiling of pediatric mature B-cell lymphoma treated in population-based prospective clinical trials. Blood 112, 1374–1381 (2008).
Klapper, W. et al. Patient age at diagnosis is associated with the molecular characteristics of diffuse large B-cell lymphoma. Blood 119, 1882–1887 (2012).
Casellas, R. et al. Restricting activation-induced cytidine deaminase tumorigenic activity in B lymphocytes. Immunology 126, 316–328 (2009).
Salaverria, I. & Siebert, R. The gray zone between Burkitt′s lymphoma and diffuse large B-cell lymphoma from a genetics perspective. J. Clin. Oncol. 29, 1835–1843 (2011).
Perk, J., Iavaraone, A. & Benezra, R. Id family of helix-loop-helix proteins in cancer. Nat. Rev. Cancer 5, 603–614 (2005).
Kee, B.L. E and ID proteins branch out. Nat. Rev. Immunol. 9, 175–184 (2009).
Seitz, V. et al. Deep sequencing of MYC DNA-binding sites in Burkitt lymphoma. PLoS ONE 6, e26837 (2011).
Pan, L. et al. Impaired immune responses and B-cell proliferation in mice lacking the Id3 gene. Mol. Cell Biol. 19, 5969–5980 (1999).
Li, J. et al. Mutation of inhibitory helix-loop-helix protein Id3 causes γδ T-cell lymphoma in mice. Blood 116, 5615–5621 (2010).
Schmitz, R. et al. Burkitt lymphoma pathogenesis and therapeutic targets from structural and functional genomics. Nature 490, 116–120 (2012).
Sander, S. et al. Synergy between PI3K signaling and MYC in Burkitt lymphomagenesis. Cancer Cell 22, 167–179 (2012).
Dominguez-Sola, D. & Dalla-Favera, R. Burkitt lymphoma: much more than MYC. Cancer Cell 22, 141–142 (2012).
Medina, P.P. & Sanchez-Cespedes, M. Involvement of the chromatin-remodeling factor BRG1/SMARCA4 in human cancer. Epigenetics 3, 64–68 (2008).
Choi, Y.J. & Lee, S.G. The DEAD-box RNA helicase DDX3 interacts with DDX5, co-localizes with it in the cytoplasm during the G2/M phase of the cycle, and affects its shuttling during mRNP export. J. Cell Biochem. 113, 985–996 (2012).
Zenz, T. et al. Monoallelic TP53 inactivation is associated with poor prognosis in chronic lymphocytic leukemia: results from a detailed genetic characterization with long-term follow-up. Blood 112, 3322–3329 (2008).
van Dongen, J.J. et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4–CT98–3936. Leukemia 17, 2257–2317 (2003).
Lefranc, M.P. et al. IMGT, the international ImMunoGeneTics database. Nucleic Acids Res. 27, 209–212 (1999).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011).
Lamprecht, B. et al. Derepression of an endogenous long terminal repeat activates the CSF1R proto-oncogene in human lymphoma. Nat. Med. 16, 571–579 (2010).
Bibikova, M. et al. High density DNA methylation array with single CpG site resolution. Genomics 98, 288–295 (2011).
Hames, B.D. Gel Electrophoresis of Proteins, Vol. 3 (Oxford University Press, New York, 1998).
Lüschen, S. et al. Sensitization to death receptor cytotoxicity by inhibition of fas-associated death domain protein (FADD)/caspase signaling. Requirement of cell cycle progression. J. Biol. Chem. 275, 24670–24678 (2000).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Jones, D.T. et al. Dissecting the genomic complexity underlying medulloblastoma. Nature 488, 100–105 (2012).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Wang, J. et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nat. Methods 8, 652–654 (2011).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Olshen, A.B. et al. Parent-specific copy number in paired tumor-normal studies using circular binary segmentation. Bioinformatics 27, 2038–2046 (2011).
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425 (2012).
Wang, K. et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674 (2007).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 16, 276–277 (2000).
Ahmadpour, F. et al. Crystal structure of the minimalist Max-E47 protein chimera. PLoS ONE 7, e32136 (2012).
Sali, A. & Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Long, A., Guanga, G.P. & Rose, R.B. Crystal structure of E47-NeuroD1/β2 bHLH domain–DNA complex: heterodimer selectivity and DNA recognition. Biochemistry 47, 218–229 (2008).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Hoffmann, S. et al. Fast mapping of short sequences with mismatches, insertions and deletions using index structures. PLoS Comput. Biol. 5, e1000502 (2009).
Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. A Stat. Soc. 57, 289–300 (1995).
Goeman, J.J., van de Geer, S.A., de Kort, F. & van Houwelingen, H.C. A global test for groups of genes: testing association with a clinical outcome. Bioinformatics 20, 93–99 (2004).
Mansmann, U. & Meister, R. Testing differential gene expression in functional groups. Goeman′s global test versus an ANCOVA approach. Methods Inf. Med. 44, 449–453 (2005).
Läuter, J., Glimm, E. & Kropf, S. New tests for data with an inherent structure. Biometrical J. 38, 5–23 (1996).
Huber, W. et al. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18, S96–S104 (2002).
Acknowledgements
This manuscript is dedicated to the memory of Karl Lennert, the founder of the Kiel classification of lymphoma, who died during the preparation of the manuscript. The authors thank O. Batic, C. Becher, C. Botz-von Drathen, A. Dietsch, J. Eils, M. Friskovec, K. Göbel, T. Grieb, S. Hengst, U. Jacobsen, T. Kaacksteen, H. Lammert, C. von der Lancken, H.-H. Müller, S. Radomski, J. Schieferstein, M. Schlapkohl, D. Schuster, S. Ölmez and L. Valles for their excellent technical support. We are grateful to G. Richter for excellent work in the coordination of the ICGC MMML-Seq Consortium and to the members of the MMML Consortium for providing extensive data of the sample analyzed within that project. We are also very grateful to all individuals who participate in this study. The project was funded by the Federal Ministry of Education and Research in Germany (BMBF) within the Program for Medical Genome Research (01KU1002A to 01KU1002J). The authors are responsible for the content of this publication. The MMML Consortium was funded by the Deutsche Krebshilfe from 2003 to 2011. Support of sequencing infrastructure by the Deutsche Forschungsgemeinschaft (DFG) and the KinderKrebsInitiative Buchholz/Holm-Seppensen is gratefully acknowledged. R.W. is the recipient of a Christoph-Schubert-Award from the KinderKrebsInitiative Buchholz/Holm-Seppensen.
Author information
Authors and Affiliations
Consortia
Contributions
B. Burkhardt, A.C., A.B., M. Rohde, J.L. and N.H. provided subject samples and clinical data. W.K., M.H. and P.M. coordinated and performed pathology review. D. Lenze, M. Szczepanowski, M.H. and W.K. stained and reviewed cryomaterial, prepared analytes and performed analyte quality control. D. Lenze and M.H. performed and interpreted immunoglobulin gene PCR. R.A.F.M. and H.G.D. characterized and provided cell lines. E.L., M. Schilhabel, V.H., S.P. and B.R. performed next-generation sequencing analyses, and A.R., P.R., P.L. and S.S. supervised next-generation sequencing analysis and interpreted data. C.L. coordinated transfer and data management of the sequences. M. Schlesner, B. Brors, S.H., S.H.B., P.F.S., D. Langenberger, V.H., J.O.K., T.R. and J.P. performed analysis of next-generation sequencing data, and B. Brors and R.E. conceived the statistical analysis. J.R., E.L., O.A., S.A.-K., M.P., S.L., R.B.R. and G.A. performed validation analyses. J.R. and R.W. were responsible for ID3 mutation and expression analyses. M.K., M. Rosolowski, M.L., H.T., C.P., R. Spang, K.M., R.K., R. Scholtysik, T.Z., D.K., L.T., D.H. and R. Siebert provided and analyzed data from the MMML cohort. M.K. and M. Rosolowski performed correlative and biometric analyses of the MMML cohort. J.R., M. Schlesner, M.K., S.H., R.E. and R. Siebert interpreted data and wrote the manuscript. R.E., M.H., W.K., P.R., A.R., B. Brors, O.A. and R. Siebert designed the study and coordinated the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The author declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–4, 6–8, 10, 11 and 15–21 and Supplementary Figures 1–18 (PDF 2842 kb)
Supplementary Table 5
Somatic structural variants (XLSX 14 kb)
Supplementary Table 9
Potentially protein changing somatic mutations in the four index patients (XLSX 411 kb)
Supplementary Table 12
U133A gene expression data of the potentially protein changing somatic mutations (XLSX 169 kb)
Supplementary Table 13
450K DNA methylation pattern of the potentially protein changing somatic mutations (XLSX 426 kb)
Supplementary Table 14
Results of ID3 and CCND3 mutation analysis of primary lymphoma (XLSX 17 kb)
Supplementary Table 22
Primer sequences (XLSX 14 kb)
Rights and permissions
About this article
Cite this article
the ICGC MMML-Seq Project. Recurrent mutation of the ID3 gene in Burkitt lymphoma identified by integrated genome, exome and transcriptome sequencing. Nat Genet 44, 1316–1320 (2012). https://doi.org/10.1038/ng.2469
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2469
This article is cited by
-
Lymphoid clonal hematopoiesis: implications for malignancy, immunity, and treatment
Blood Cancer Journal (2023)
-
Mosaic chromosomal alterations in peripheral blood leukocytes of children in sub-Saharan Africa
Nature Communications (2023)
-
Burkitt lymphoma
Nature Reviews Disease Primers (2022)
-
Complex genetic and histopathological study of 15 patient-derived xenografts of aggressive lymphomas
Laboratory Investigation (2022)
-
Clinical relevance of molecular characteristics in Burkitt lymphoma differs according to age
Nature Communications (2022)