Abstract
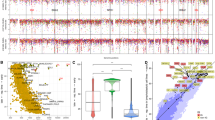
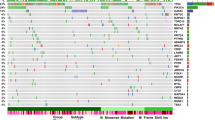
Burkitt lymphoma is characterized by deregulation of MYC, but the contribution of other genetic mutations to the disease is largely unknown. Here, we describe the first completely sequenced genome from a Burkitt lymphoma tumor and germline DNA from the same affected individual. We further sequenced the exomes of 59 Burkitt lymphoma tumors and compared them to sequenced exomes from 94 diffuse large B-cell lymphoma (DLBCL) tumors. We identified 70 genes that were recurrently mutated in Burkitt lymphomas, including ID3, GNA13, RET, PIK3R1 and the SWI/SNF genes ARID1A and SMARCA4. Our data implicate a number of genes in cancer for the first time, including CCT6B, SALL3, FTCD and PC. ID3 mutations occurred in 34% of Burkitt lymphomas and not in DLBCLs. We show experimentally that ID3 mutations promote cell cycle progression and proliferation. Our work thus elucidates commonly occurring gene-coding mutations in Burkitt lymphoma and implicates ID3 as a new tumor suppressor gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Dave, S.S. et al. Molecular diagnosis of Burkitt's lymphoma. N. Engl. J. Med. 354, 2431–2442 (2006).
Schiffman, J.D. et al. Genome wide copy number analysis of paediatric Burkitt lymphoma using formalin-fixed tissues reveals a subset with gain of chromosome 13q and corresponding miRNA over expression. Br. J. Haematol. 155, 477–486 (2011).
Hummel, M. et al. A biologic definition of Burkitt's lymphoma from transcriptional and genomic profiling. N. Engl. J. Med. 354, 2419–2430 (2006).
Swerdlow, S.H. et al. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues 4th edn. (IARC Press, Lyon, France, 2008).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Chen, K. et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat. Methods 6, 677–681 (2009).
Pleasance, E.D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, 184–190 (2010).
Sherry, S.T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Li, Y. et al. Resequencing of 200 human exomes identifies an excess of low-frequency non-synonymous coding variants. Nat. Genet. 42, 969–972 (2010).
Ng, S.B. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, 272–276 (2009).
Siva, N. 1000 Genomes project. Nat. Biotechnol. 26, 256 (2008).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Morin, R.D. et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476, 298–303 (2011).
Pasqualucci, L. et al. Analysis of the coding genome of diffuse large B-cell lymphoma. Nat. Genet. 43, 830–837 (2011).
Lohr, J.G. et al. Discovery and prioritization of somatic mutations in diffuse large B-cell lymphoma (DLBCL) by whole-exome sequencing. Proc. Natl. Acad. Sci. USA 109, 3879–3884 (2012).
Ngo, V.N. et al. Oncogenically active MYD88 mutations in human lymphoma. Nature 470, 115–119 (2011).
Wright, G. et al. A gene expression–based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc. Natl. Acad. Sci. USA 100, 9991–9996 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Dalla-Favera, R., Martinotti, S., Gallo, R.C., Erikson, J. & Croce, C.M. Translocation and rearrangements of the c-myc oncogene locus in human undifferentiated B-cell lymphomas. Science 219, 963–967 (1983).
Little, C.D., Nau, M.M., Carney, D.N., Gazdar, A.F. & Minna, J.D. Amplification and expression of the c-myc oncogene in human lung cancer cell lines. Nature 306, 194–196 (1983).
Münzel, P., Marx, D., Kochel, H., Schauer, A. & Bock, K.W. Genomic alterations of the c-myc protooncogene in relation to the overexpression of c-erbB2 and Ki-67 in human breast and cervix carcinomas. J. Cancer Res. Clin. Oncol. 117, 603–607 (1991).
Wang, Z.R., Liu, W., Smith, S.T., Parrish, R.S. & Young, S.R. c-myc and chromosome 8 centromere studies of ovarian cancer by interphase FISH. Exp. Mol. Pathol. 66, 140–148 (1999).
Augenlicht, L.H. et al. Low-level c-myc amplification in human colonic carcinoma cell lines and tumors: a frequent, p53-independent mutation associated with improved outcome in a randomized multi-institutional trial. Cancer Res. 57, 1769–1775 (1997).
Kee, B.L. E and ID proteins branch out. Nat. Rev. Immunol. 9, 175–184 (2009).
Morin, R.D. et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat. Genet. 42, 181–185 (2010).
Jima, D.D. et al. Deep sequencing of the small RNA transcriptome of normal and malignant human B cells identifies hundreds of novel microRNAs. Blood 116, e118–e127 (2010).
Cock, P.J., Fields, C.J., Goto, N., Heuer, M.L. & Rice, P.M. The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res. 38, 1767–1771 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Parmigiani, G. et al. Design and analysis issues in genome-wide somatic mutation studies of cancer. Genomics 93, 17–21 (2009).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Ge, D. et al. SVA: software for annotating and visualizing sequenced human genomes. Bioinformatics 27, 1998–2000 (2011).
Reva, B., Antipin, Y. & Sander, C. Determinants of protein function revealed by combinatorial entropy optimization. Genome Biol. 8, R232 (2007).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Kumar, P., Henikoff, S. & Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Needleman, S.B. & Wunsch, C.D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J. Mol. Biol. 48, 443–453 (1970).
Pruitt, K.D. et al. The consensus coding sequence (CCDS) project: identifying a common protein-coding gene set for the human and mouse genomes. Genome Res. 19, 1316–1323 (2009).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Wood, L.D. et al. The genomic landscapes of human breast and colorectal cancers. Science 318, 1108–1113 (2007).
Acknowledgements
The authors thank S. Sunay and the Georgia Cancer Coalition for support in sample collection. A.B.M. was supported by the Hertz Foundation. This work was supported through grants R21CA1561686 and R01CA136895 from the National Cancer Institute (S.S.D.). S.S.D. was also supported by the American Cancer Society. We gratefully acknowledge the generous support of C. Stiefel and D. Stiefel.
Author information
Authors and Affiliations
Contributions
C.L., R.M., K.L.R., C.H.D., W.W.L.C., G.S., P.L.L., D.A.R., A.S.L., L.B.-M., K.P.M., C.R.F., K.N.N., A.M.E., A.C., L.I.G., M.B.C., J.I.G., E.D.H., J.Z., G.L., A.G., M.M., S.L. and S.S.D. performed research and edited the manuscript. C.L., J.Z., A.B.M., D.J., Z.S., V.G., A.B., D.B.D. and S.S.D. analyzed data. C.L. and S.S.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note and Supplementary Figures 1–10 (PDF 4090 kb)
Supplementary Table 1
Burkitt lymphoma whole genome sequencing variants (XLS 2765 kb)
Supplementary Table 2
Sanger sequence validation (XLS 66 kb)
Supplementary Table 3
Recurrently mutated genes in Burkitt lymphoma (XLS 50 kb)
Supplementary Table 4
Individual variants found in Burkitt lymphoma (XLS 217 kb)
Supplementary Table 5
Patient data (XLS 42 kb)
Supplementary Table 6
Primer sequences (XLS 30 kb)
Rights and permissions
About this article
Cite this article
Love, C., Sun, Z., Jima, D. et al. The genetic landscape of mutations in Burkitt lymphoma. Nat Genet 44, 1321–1325 (2012). https://doi.org/10.1038/ng.2468
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2468
This article is cited by
-
SWI/SNF complexes in hematological malignancies: biological implications and therapeutic opportunities
Molecular Cancer (2023)
-
The role of chromatin remodeler SMARCA4/BRG1 in brain cancers: a potential therapeutic target
Oncogene (2023)
-
Successful pseudo-autologous stem cell transplantation for donor-derived Burkitt lymphoma occurring 9 years after allogeneic transplantation
International Journal of Hematology (2023)
-
Mosaic chromosomal alterations in peripheral blood leukocytes of children in sub-Saharan Africa
Nature Communications (2023)
-
SMARCA4 biology in alveolar rhabdomyosarcoma
Oncogene (2022)