Abstract
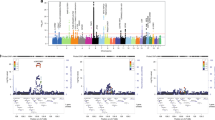
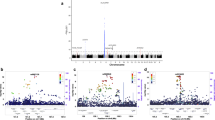
To identify rheumatoid arthritis risk loci in European populations, we conducted a meta-analysis of two published genome-wide association (GWA) studies totaling 3,393 cases and 12,462 controls1,2. We genotyped 31 top-ranked SNPs not previously associated with rheumatoid arthritis in an independent replication of 3,929 autoantibody-positive rheumatoid arthritis cases and 5,807 matched controls from eight separate collections. We identified a common variant at the CD40 gene locus (rs4810485, P = 0.0032 replication, P = 8.2 × 10−9 overall, OR = 0.87). Along with other associations near TRAF1 (refs. 2,3) and TNFAIP3 (refs. 4,5), this implies a central role for the CD40 signaling pathway in rheumatoid arthritis pathogenesis. We also identified association at the CCL21 gene locus (rs2812378, P = 0.00097 replication, P = 2.8 × 10−7 overall), a gene involved in lymphocyte trafficking. Finally, we identified evidence of association at four additional gene loci: MMEL1-TNFRSF14 (rs3890745, P = 0.0035 replication, P = 1.1 × 10−7 overall), CDK6 (rs42041, P = 0.010 replication, P = 4.0 × 10−6 overall), PRKCQ (rs4750316, P = 0.0078 replication, P = 4.4 × 10−6 overall), and KIF5A-PIP4K2C (rs1678542, P = 0.0026 replication, P = 8.8 × 10−8 overall).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
The Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Plenge, R.M. et al. TRAF1–C5 as a risk locus for rheumatoid arthritis–a genomewide study. N. Engl. J. Med. 357, 1199–1209 (2007).
Kurreeman, F.A. et al. A candidate gene approach identifies the TRAF1/C5 region as a risk factor for rheumatoid arthritis. PLoS Med. 4, e278 (2007).
Plenge, R.M. et al. Two independent alleles at 6q23 associated with risk of rheumatoid arthritis. Nat. Genet. 39, 1477–1482 (2007).
Thomson, W. et al. Rheumatoid arthritis association at 6q23. Nat. Genet. 39, 1431–1433 (2007).
Stastny, P. Association of the B-cell alloantigen DRw4 with rheumatoid arthritis. N. Engl. J. Med. 298, 869–871 (1978).
Begovich, A.B. et al. A missense single-nucleotide polymorphism in a gene encoding a protein tyrosine phosphatase (PTPN22) is associated with rheumatoid arthritis. Am. J. Hum. Genet. 75, 330–337 (2004).
Remmers, E.F. et al. STAT4 and the risk of rheumatoid arthritis and systemic lupus erythematosus. N. Engl. J. Med. 357, 977–986 (2007).
Zhernakova, A. et al. Novel association in chromosome 4q27 region with rheumatoid arthritis and confirmation of type 1 diabetes point to a general risk locus for autoimmune diseases. Am. J. Hum. Genet. 81, 1284–1288 (2007).
Plenge, R.M. et al. Replication of putative candidate-gene associations with rheumatoid arthritis in >4,000 samples from North America and Sweden: association of susceptibility with PTPN22, CTLA4, and PADI4. Am. J. Hum. Genet. 77, 1044–1060 (2005).
Suzuki, A. et al. Functional haplotypes of PADI4, encoding citrullinating enzyme peptidylarginine deiminase 4, are associated with rheumatoid arthritis. Nat. Genet. 34, 395–402 (2003).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Jacobson, E.M. et al. A CD40 Kozak sequence polymorphism and susceptibility to antibody-mediated autoimmune conditions: the role of CD40 tissue-specific expression. Genes Immun. 8, 205–214 (2007).
Heward, J.M. et al. A single nucleotide polymorphism in the CD40 gene on chromosome 20q (GD-2) provides no evidence for susceptibility to Graves' disease in UK Caucasians. Clin. Endocrinol. 61, 269–272 (2004).
Houston, F.A. et al. Role of the CD40 locus in Graves' disease. Thyroid 14, 506–509 (2004).
Kawabe, T. et al. The immune responses in CD40-deficient mice: impaired immunoglobulin class switching and germinal center formation. Immunity 1, 167–178 (1994).
Lougaris, V., Badolato, R., Ferrari, S. & Plebani, A. Hyper immunoglobulin M syndrome due to CD40 deficiency: clinical, molecular, and immunological features. Immunol. Rev. 203, 48–66 (2005).
Manzo, A. et al. Systematic microanatomical analysis of CXCL13 and CCL21 in situ production and progressive lymphoid organization in rheumatoid synovitis. Eur. J. Immunol. 35, 1347–1359 (2005).
Ishida, T. et al. CD40 signaling-mediated induction of Bcl-XL, Cdk4, and Cdk6. Implication of their cooperation in selective B cell growth. J. Immunol. 155, 5527–5535 (1995).
Veiga-Fernandes, H. & Rocha, B. High expression of active CDK6 in the cytoplasm of CD8 memory cells favors rapid division. Nat. Immunol. 5, 31–37 (2004).
Carpenter, C.L. Btk-dependent of phosphoinositide synthesis. Biochem. Soc. Trans. 32, 326–329 (2004).
Marsters, S.A. et al. Herpesvirus entry mediator, a member of the tumor necrosis factor receptor (TNFR) family, interacts with members of the TNFR-associated factor family and activates the transcription factors NF-kappaB and AP-1. J. Biol. Chem. 272, 14029–14032 (1997).
Gruber, T. et al. PKCtheta cooperates with atypical PKCzeta and PKCiota in NF-kappaB transactivation of T lymphocytes. Mol. Immunol. 45, 117–126 (2008).
Barton, A. et al. Rheumatoid arthritis susceptibility loci at chromosomes 10p15, 12q13 and 22q13. Nat. Genet. advance online publication, doi:10.1038/ng.218 (14 September 2008).
Harnett, M.M. CD40: a growing cytoplasmic tale. Sci. STKE 2004, pe25 (2004).
Xie, P., Hostager, B.S., Munroe, M.E., Moore, C.R. & Bishop, G.A. Cooperation between TNF receptor-associated factors 1 and 2 in CD40 signaling. J. Immunol. 176, 5388–5400 (2006).
Song, H.Y., Rothe, M. & Goeddel, D.V. The tumor necrosis factor-inducible zinc finger protein A20 interacts with TRAF1/TRAF2 and inhibits NF-kappaB activation. Proc. Natl. Acad. Sci. USA 93, 6721–6725 (1996).
Balasa, B. et al. CD40 ligand-CD40 interactions are necessary for the initiation of insulitis and diabetes in nonobese diabetic mice. J. Immunol. 159, 4620–4627 (1997).
Durie, F.H. et al. Prevention of collagen-induced arthritis with an antibody to gp39, the ligand for CD40. Science 261, 1328–1330 (1993).
MacGregor, A.J. et al. Characterizing the quantitative genetic contribution to rheumatoid arthritis using data from twins. Arthritis Rheum. 43, 30–37 (2000).
Acknowledgements
We thank the WTCCC for making access to genotype data available online, and for providing autoantibody status of the WTCCC rheumatoid arthritis cases. We thank S. Myers and J. Marchini for help with IMPUTE. S.R. is supported by a T32 NIH training grant (AR007530-23), a K08 grant from the NIH (KAR055688A), and through the BWH Rheumatology Fellowship program, directed by S. Helfgott. R.M.P. is supported by a K08 grant from the NIH (AI55314-3), a private donation from the Fox Trot Fund, the William Randolph Hearst Fund of Harvard University, and holds a Career Award for Medical Scientists from the Burroughs Wellcome Fund. M.J.D. is supported by a UO1 NIH grant (UO1 HG004171). The BRASS Registry is supported by a grant from Millennium Pharmaceuticals and Biogen-Idec. The Broad Institute Center for Genotyping and Analysis is supported by grant U54 RR020278 from the National Center for Research Resources. The NARAC is supported by NIH grants RO1-AR44422 and NO1-AR-2-2263 (P.K.G.). This work was also supported in part by the Intramural Research Program of the National Institute of Arthritis and Musculoskeletal and Skin Diseases of the National Institutes of Health. The EIRA study is supported by grants from the Swedish Medical Research Council, the Swedish Council for Working Life and Social Research, King Gustaf V's 80-year Foundation, the Swedish Rheumatic Foundation, the Stockholm County Council, the insurance company Arbetsmarknadens Försäkringsaktiebolag, and the County of Sörmland Research and Development Center. Genotyping of the EIRA cohort was supported by the Agency for Science Technology and Research (Singapore). Genotyping of the GCI and LUMC samples was funded by Celera. D.A. is a Burroughs Wellcome Fund Clinical Scholar in Translational Research, and a Distinguished Clinical Scholar of the Doris Duke Charitable Foundation. B.D.K. was supported by the NIH Clinical Research Training Program, a public-private partnership between the Foundation for the National Institutes of Health and Pfizer Inc. E.W.K. is supported by NIH grants R01 AR49880, CA87969, P60 AR047782, K24 AR0524-01 and BIRCWH K12 HD051959 (supported by US National Institute of Mental Health, National Institute of Allergy and Infectious Diseases, National Institute of Child Health and Human Development and the Office of the Director). K.H.C. is the recipient of an Arthritis Foundation/American College of Rheumatology Arthritis Investigator Award and a Katherine Swan Ginsburg Memorial Award. L.A.C. is a Kirkland Scholar Awardee and her work on this project was supported by K24 AR02175, R01 AI065841 and 5 M01 RR-00079. P.P.T. and N.d.V. were supported by European Community's FP6 funding (Autocure). We acknowledge the help of C. Ellen van der Schoot for healthy control samples for GENRA and the help of B.A.C. Dijkmans, D. van Schaardenburg, A.S. Peña, P.L. Klarenbeek, Z. Zhang, M.T. Nurmohammed, W.F. Lems, R.R.J. van de Stadt, W.H. Bos, J. Ursum, M.G.M. Bartelds, D.M. Gerlag, M.G.H. van der Sande, C.A. Wijbrandts and M.M.J. Herenius in gathering GENRA subject samples and data. We thank Y. Li, S. Schrodi, and J. Sninsky (Celera); and B. Voight and C. Cotsapas (Broad Institute) for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
S.R., M.J.D. and R.M.P. designed the study, conducted the statistical analysis, interpreted the primary data, and wrote the initial manuscript. E.F.R. and B.D.K. generated Sequenom genotype data on the replication samples from North America (at NIAMS); N.P.B., R.H., L.G., S.W. and C.G. generated Sequenom genotype data on the replication samples from North America and NHS (at the Broad Institute); A.T.L. and P.K.G. generated the NARAC genome-wide association genotype data; A.B.B., M.C., K.G.A. and J.J.C. generated genotype data on GCI and LUMC samples (at Celera); F.A.S.K., R.E.M.T., A.H.M.M., and T.W.J.H. contributed LUMC samples; G.J.W., P.P.T., J.B.A.C., I.E.V.D.H.-B. and N.d.V. contributed GENRA samples; K.H.C., J. Cui and E.W.K. contributed NHS samples and interpretation of the study data; L.K., L.P., B.D. and L.A. contributed EIRA samples and helped interpret the data; J.W. is principle investigator of the rheumatoid arthritis WTCCC study and contributed to the study design; N.A.S. and M.E.W. are principle investigators of BRASS, and J. Coblyn contributed BRASS subject samples. P.K.G. is principle investigator of NARAC, provided replication samples and guidance on the study design, and helped to interpret the data. D.A. provided guidance on study design, interpretation of data and initial draft of manuscript. B.M.N. contributed statistical analysis. M.S. generated the genome-wide genotype data on EIRA. L.A.C., C.I.A., M.F.S., D.L.K., E.F.R., A.T.L., R.M.P. and P.K.G. are all members of NARAC and have contributed to the study design. All authors contributed to writing the final manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1 and 2, Supplementary Tables 1–5 (PDF 613 kb)
Rights and permissions
About this article
Cite this article
Raychaudhuri, S., Remmers, E., Lee, A. et al. Common variants at CD40 and other loci confer risk of rheumatoid arthritis. Nat Genet 40, 1216–1223 (2008). https://doi.org/10.1038/ng.233
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.233
This article is cited by
-
Could NCOA5 a novel candidate gene for multiple sclerosis susceptibility?
Molecular Biology Reports (2023)
-
Genome-wide exploratory analysis for NARAC dataset with preparation for haplotype block partitioning through minor allele frequency quality control viewpoint
Iran Journal of Computer Science (2023)
-
Gene selection by incorporating genetic networks into case-control association studies
European Journal of Human Genetics (2022)
-
Transcriptome-wide association study identifies susceptibility genes for rheumatoid arthritis
Arthritis Research & Therapy (2021)
-
Integrative analysis of genome-wide association studies identifies novel loci associated with neuropsychiatric disorders
Translational Psychiatry (2021)