Abstract
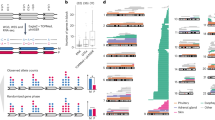
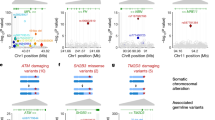
We detected clonal mosaicism for large chromosomal anomalies (duplications, deletions and uniparental disomy) using SNP microarray data from over 50,000 subjects recruited for genome-wide association studies. This detection method requires a relatively high frequency of cells with the same abnormal karyotype (>5–10%; presumably of clonal origin) in the presence of normal cells. The frequency of detectable clonal mosaicism in peripheral blood is low (<0.5%) from birth until 50 years of age, after which it rapidly rises to 2–3% in the elderly. Many of the mosaic anomalies are characteristic of those found in hematological cancers and identify common deleted regions with genes previously associated with these cancers. Although only 3% of subjects with detectable clonal mosaicism had any record of hematological cancer before DNA sampling, those without a previous diagnosis have an estimated tenfold higher risk of a subsequent hematological cancer (95% confidence interval = 6–18).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Miller, O.J. & Therman, E. Human Chromosomes (Springer-Verlag, New York, 2001).
Strachan, T. & Read, A.P. Human Molecular Genetics (Wiley-Liss, New York, 1996).
Nowell, P.C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Delhanty, J.D. Mechanisms of aneuploidy induction in human oogenesis and early embryogenesis. Cytogenet. Genome Res. 111, 237–244 (2005).
Vanneste, E. et al. Chromosome instability is common in human cleavage-stage embryos. Nat. Med. 15, 577–583 (2009).
Hassold, T. Mosaic trisomies in human spontaneous abortions. Hum. Genet. 61, 31–35 (1982).
Conlin, L.K. et al. Mechanisms of mosaicism, chimerism and uniparental disomy identified by single nucleotide polymorphism array analysis. Hum. Mol. Genet. 19, 1263–1275 (2010).
Ballif, B.C. et al. Detection of low-level mosaicism by array CGH in routine diagnostic specimens. Am. J. Med. Genet. A. 140, 2757–2767 (2006).
Heim, S. & Mitelman, F. Nonrandom chromosome abnormalities in cancer—an overview. in Cancer Cytogenetics (eds. Mitelman, F. & Heim, S.) 25–44 (John Wiley & Sons, Hoboken, New Jersey, 2009).
Gardner, R.J.M. & Sutherland, G.R. Chromosome Abnormalities and Genetic Counseling (Oxford University Press, Oxford, 2004).
Maciejewski, J.P., Tiu, R.V. & O'Keefe, C. Application of array-based whole genome scanning technologies as a cytogenetic tool in haematological malignancies. Br. J. Haematol. 146, 479–488 (2009).
Dougherty, M.J. et al. Implementation of high resolution single nucleotide polymorphism array analysis as a clinical test for patients with hematologic malignancies. Cancer Genet. 204, 26–38 (2011).
McCarroll, S.A. & Altshuler, D.M. Copy-number variation and association studies of human disease. Nat. Genet. 39, S37–S42 (2007).
Conrad, D.F. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010).
Itsara, A. et al. Population analysis of large copy number variants and hotspots of human genetic disease. Am. J. Hum. Genet. 84, 148–161 (2009).
Cornelis, M.C. et al. The Gene, Environment Association Studies consortium (GENEVA): maximizing the knowledge obtained from GWAS by collaboration across studies of multiple conditions. Genet. Epidemiol. 34, 364–372 (2010).
Peiffer, D.A. et al. High-resolution genomic profiling of chromosomal aberrations using Infinium whole-genome genotyping. Genome Res. 16, 1136–1148 (2006).
Jacobs, K.B. et al. Detectable clonal mosaicism and its relationship to aging and cancer. Nat. Genet. published online, doi:10.1038/ng.2270 (6 May 2012).
Pinto, D. et al. Comprehensive assessment of array-based platforms and calling algorithms for detection of copy number variants. Nat. Biotechnol. 29, 512–520 (2011).
Pekarsky, Y., Zanesi, N. & Croce, C.M. Molecular basis of CLL. Semin. Cancer Biol. 20, 370–376 (2010).
Döhner, H. et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N. Engl. J. Med. 343, 1910–1916 (2000).
Bejar, R., Levine, R. & Ebert, B.L. Unraveling the molecular pathophysiology of myelodysplastic syndromes. J. Clin. Oncol. 29, 504–515 (2011).
Yan, X.J. et al. Exome sequencing identifies somatic mutations of DNA methyltransferase gene DNMT3A in acute monocytic leukemia. Nat. Genet. 43, 309–315 (2011).
Gunn, S.R. et al. Array CGH analysis of chronic lymphocytic leukemia reveals frequent cryptic monoallelic and biallelic deletions of chromosome 22q11 that include the PRAME gene. Leuk. Res. 33, 1276–1281 (2009).
Gurvich, N. et al. L3MBTL1 polycomb protein, a candidate tumor suppressor in del(20q12) myeloid disorders, is essential for genome stability. Proc. Natl. Acad. Sci. USA 107, 22552–22557 (2010).
Tuna, M., Knuutila, S. & Mills, G.B. Uniparental disomy in cancer. Trends Mol. Med. 15, 120–128 (2009).
O′Keefe, C., McDevitt, M.A. & Maciejewski, J.P. Copy neutral loss of heterozygosity: a novel chromosomal lesion in myeloid malignancies. Blood 115, 2731–2739 (2010).
Raghavan, M., Gupta, M., Molloy, G., Chaplin, T. & Young, B.D. Mitotic recombination in haematological malignancy. Adv. Enzyme Regul. 50, 96–103 (2010).
Forsberg, L.A. et al. Age-related somatic structural changes in the nuclear genome of human blood cells. Am. J. Hum. Genet. 90, 217–228 (2012).
Vorobtsova, I., Semenov, A., Timofeyeva, N., Kanayeva, A. & Zvereva, I. An investigation of the age-dependency of chromosome abnormalities in human populations exposed to low-dose ionising radiation. Mech. Ageing Dev. 122, 1373–1382 (2001).
Mukherjee, A.B. & Thomas, S. A longitudinal study of human age-related chromosomal analysis in skin fibroblasts. Exp. Cell Res. 235, 161–169 (1997).
Rossi, D.J. et al. Hematopoietic stem cell quiescence attenuates DNA damage response and permits DNA damage accumulation during aging. Cell Cycle 6, 2371–2376 (2007).
Lindstrom, D.L., Leverich, C.K., Henderson, K.A. & Gottschling, D.E. Replicative age induces mitotic recombination in the ribosomal RNA gene cluster of Saccharomyces cerevisiae. PLoS Genet. 7, e1002015 (2011).
Sahin, E. & Depinho, R.A. Linking functional decline of telomeres, mitochondria and stem cells during ageing. Nature 464, 520–528 (2010).
Sharpless, N.E. & DePinho, R.A. How stem cells age and why this makes us grow old. Nat. Rev. Mol. Cell Biol. 8, 703–713 (2007).
Crow, J.F. & Kimura, M. An Introduction to Population Genetics Theory (Harper and Row, New York, 1970).
Prchal, J.T. et al. Clonal stability of blood cell lineages indicated by X-chromosomal transcriptional polymorphism. J. Exp. Med. 183, 561–567 (1996).
Swierczek, S.I. et al. Hematopoiesis is not clonal in healthy elderly women. Blood 112, 3186–3193 (2008).
Fischbach, F. & Dunning, M.B. A Manual of Laboratory and Diagnostic Tests (Lippincott, Williams and Wilkins, Philadelphia, 1992).
Vandewoestyne, M.L. et al. Laser microdissection for the assessment of the clonal relationship between chronic lymphocytic leukemia/small lymphocytic lymphoma and proliferating B cells within lymph node pseudofollicles. Leukemia 25, 883–888 (2011).
Marti, G.E. et al. Diagnostic criteria for monoclonal B-cell lymphocytosis. Br. J. Haematol. 130, 325–332 (2005).
Landgren, O. et al. B-cell clones as early markers for chronic lymphocytic leukemia. N. Engl. J. Med. 360, 659–667 (2009).
Shanafelt, T.D., Ghia, P., Lanasa, M.C., Landgren, O. & Rawstron, A.C. Monoclonal B-cell lymphocytosis (MBL): biology, natural history and clinical management. Leukemia 24, 512–520 (2010).
Cogle, C.R., Craig, B.M., Rollison, D.E. & List, A.F. Incidence of the myelodysplastic syndromes using a novel claims-based algorithm: high number of uncaptured cases by cancer registries. Blood 117, 7121–7125 (2011).
Neukirchen, J. et al. Incidence and prevalence of myelodysplastic syndromes: data from the Dusseldorf MDS-registry. Leuk. Res. 35, 1591–1596 (2011).
Ma, X., Vanasse, G., Cartmel, B., Wang, Y. & Selinger, H.A. Prevalence of polycythemia vera and essential thrombocythemia. Am. J. Hematol. 83, 359–362 (2008).
Simon-Sanchez, J. et al. Genome-wide SNP assay reveals structural genomic variation, extended homozygosity and cell-line induced alterations in normal individuals. Hum. Mol. Genet. 16, 1–14 (2007).
Diskin, S.J. et al. Adjustment of genomic waves in signal intensities from whole-genome SNP genotyping platforms. Nucleic Acids Res. 36, e126 (2008).
Olshen, A.B., Venkatraman, E.S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Lin, P. et al. Copy number variation accuracy in genome-wide association studies. Hum. Hered. 71, 141–147 (2011).
Wang, K. et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674 (2007).
Tracy, N.D., Young, J.C. & Mason, R.L. Multivariate control charts for individual observations. Journal of Quality Technology 24, 88–95 (1992).
Rodríguez-Santiago, B. et al. Mosaic uniparental disomies and aneuploidies as large structural variants of the human genome. Am. J. Hum. Genet. 87, 129–138 (2010).
Acknowledgements
The GENEVA Consortium thanks the subjects and the staff of all GENEVA studies for their important contributions. We thank the following state cancer registries for their help: Alaska, Arizona, Arkansas, California, Colorado, Connecticut, Delaware, Florida, Georgia, Idaho, Illinois, Indiana, Iowa, Kentucky, Louisiana, Maine, Maryland, Massachusetts, Michigan, Nebraska, New Hampshire, New Jersey, New York, North Carolina, North Dakota, Ohio, Oklahoma, Oregon, Pennsylvania, Rhode Island, South Carolina, Tennessee, Texas, Virginia, Washington and Wyoming. We thank C. Laird and G. Marti for helpful comments on the manuscript and B. Wakimoto and D. Gottschling for enlightening discussions. We also thank K. Jacobs for exchanging ideas and for working with us to estimate cross-method concordance of mosaic detection using the PLCO/GENEVA Lung Cancer study. Support for the GENEVA genome-wide association studies was provided through the US National Institutes of Health (NIH) Genes, Environment and Health Initiative (GEI). Some studies also received support from individual NIH Institutes. The grant numbers are: Melanoma (NCI R29CA70334, R01CA100264 and P50CA093459); Lung Health (U01HG004738); Cleft Lip/Palate (National Institute Dental and Craniofacial Research (NIDCR): U01DE018993 and NIH contract: HHSN268200782096C); Addiction (U01HG004422, National Institute on Alcohol Abuse and Alcoholism (NIAAA): U10AA008401, National Cancer Institute (NCI): P01CA089392, National Institute on Drug Abuse (NIDA): R01DA013423 and R01DA019963); Lung Cancer (Z01CP010200); Blood Clotting (R37 HL 039693); Prostate Cancer (U01HG004726, NCI: CA63464, CA54281, CA1326792 and RC2 CA148085); Venous Thromboembolism (U01HG004735); Birth Weight (U01HG004415); Dental Caries (NIDCR: U01DE018903 and R01DE014899, NIH Center for Inherited Disease Research (CIDR) contract: HHSN268200-782096C); Prematurity (U01HG004423); Glaucoma (U01HG004728, National Eye Institute (NEI): R01EY015473 and R01EY015872); GENEVA Coordinating Center (U01 HG004446); CIDR (U01HG004438 and HHSN268200782096C); Broad Center for Genotyping and Analysis (U01HG04424); the Intramural Research Program of the NIH, the National Library of Medicine; and the Intramural Research Program of the Division of Cancer Epidemiology and Genetics, NCI, NIH. L.R.P. was also supported by a Physician Scientist award from Research to Prevent Blindness in NYC and an Ophthalmology Scholar Award from Harvard Medical School and from the Harvard Glaucoma Center of Excellence. L.R.Z. was supported by the NCI (T32 CA09168).
Author information
Authors and Affiliations
Contributions
K.F.D., H.L., K.N.H. and E.W.P. initiated the detection of chromosomal anomalies in GENEVA GWAS data. C.A.L. developed the automated methods of anomaly detection, with assistance from C.C.L., L.R.Z., C.P.M., V.E.S. and A.N.M. C.C.L., C.A.L., K.R., L.R.Z., C.P.M., J.S., D.R.C., D.M.L., X.Z., S.C.N., S.M.G., M.P.C., J.I.U. and S.B. performed data analyses. C.A., Q.W., L.W., J.E.L., K.C.B., N.N.H., R.M., T.H.B., A.F.S., L.J.B., M.T.L., L.R.G., D.G., K.C.D., S.S.S., W.J.B., L.B.S., S.A.I., S.J.C., S.I.B., L.L.M., B.E.H., J.A.H., S.M.A., C.R., W.L.L., M.L.M., J.C.M., M.M., B.F., J.H.K., J.L.W., L.R.P., C.A.H. and N.C. contributed sample collections and phenotypic data. K.F.D., H.L., K.N.H., E.W.P., D.B.M. and A.C. performed genotyping. L.R.P., J.H.K., N.C., C.A.H., B.E.H. and K.R.M. provided data and interpretation for analysis of incident hematological cancer. C.C.L., C.A.L., L.R.Z., K.F.D., K.R., C.A., D.D., T.H.B., A.F.S., I.R., R.B.S., L.J.B., S.M.H., N.D.F., J.L., B.E.H., K.R.M., M.d.A., W.L.L., M.G.H., M.L.M., E.F., J.C.M., M.M., B.F., J.L.W., A.W., C.P.M., J.S., D.R.C., D.M.L., X.Z., J.I.U., S.B., S.C.N., S.M.G., P.H., G.P.J., A.N.M., V.E.S., H.L., K.N.H., E.W.P., D.B.M., A.C., N.S., T.M., L.R.P., C.A.H., N.C. and B.S.W. contributed ideas and advice during regular discussions of the project. C.C.L. coordinated the study and wrote the first draft of the manuscript, with guidance from a writing committee consisting of C.A.L., K.R., K.F.D., T.M., L.R.P., N.C. and B.S.W. All authors contributed to review and revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
L.J.B. served as a consultant for Pfizer Inc. in 2008 and is an inventor on the patent Markers for Addiction (US 20070258898) covering the use of certain SNPs in determining the diagnosis, prognosis and treatment of addiction.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1, 2, 4, 5 and 7 and Supplementary Figures 1–13 (PDF 5308 kb)
Supplementary Table 3
Breakpoints and other characteristics of autosomal mosaic anomalies (XLSX 64 kb)
Supplementary Table 6
Site and histology descriptions and hematological cancer category (XLSX 13 kb)
Rights and permissions
About this article
Cite this article
Laurie, C., Laurie, C., Rice, K. et al. Detectable clonal mosaicism from birth to old age and its relationship to cancer. Nat Genet 44, 642–650 (2012). https://doi.org/10.1038/ng.2271
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2271
This article is cited by
-
Driver mutation zygosity is a critical factor in predicting clonal hematopoiesis transformation risk
Blood Cancer Journal (2024)
-
A pan-tissue survey of mosaic chromosomal alterations in 948 individuals
Nature Genetics (2023)
-
Lymphoid clonal hematopoiesis: implications for malignancy, immunity, and treatment
Blood Cancer Journal (2023)
-
Mutation rates and fitness consequences of mosaic chromosomal alterations in blood
Nature Genetics (2023)
-
High-throughput electron tomography identifies centriole over-elongation as an early event in plasma cell disorders
Leukemia (2023)