Abstract
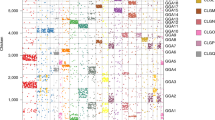
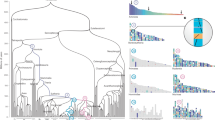
Human segmental duplications are hotspots for nonallelic homologous recombination leading to genomic disorders, copy-number polymorphisms and gene and transcript innovations. The complex structure and history of these regions have precluded a global evolutionary analysis. Combining a modified A-Bruijn graph algorithm with comparative genome sequence data, we identify the origin of 4,692 ancestral duplication loci and use these to cluster 437 complex duplication blocks into 24 distinct groups. The sequence-divergence data between ancestral-derivative pairs and a comparison with the chimpanzee and macaque genome support a 'punctuated' model of evolution. Our analysis reveals that human segmental duplications are frequently organized around 'core' duplicons, which are enriched for transcripts and, in some cases, encode primate-specific genes undergoing positive selection. We hypothesize that the rapid expansion and fixation of some intrachromosomal segmental duplications during great-ape evolution has been due to the selective advantage conferred by these genes and transcripts embedded within these core duplications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bailey, J.A., Yavor, A.M., Massa, H.F., Trask, B.J. & Eichler, E.E. Segmental duplications: organization and impact within the current human genome project assembly. Genome Res. 11, 1005–1017 (2001).
She, X. et al. A preliminary comparative analysis of primate segmental duplications shows elevated substitution rates and a great-ape expansion of intrachromosomal duplications. Genome Res. 16, 576–583 (2006).
Eichler, E.E. et al. Interchromosomal duplications of the adrenoleukodystrophy locus: a phenomenon of pericentromeric plasticity. Hum. Mol. Genet. 6, 991–1002 (1997).
Orti, R. et al. Conservation of pericentromeric duplications of a 200-kb part of the human 21q22.1 region in primates. Cytogenet. Cell Genet. 83, 262–265 (1998).
Jackson, M.S. et al. Sequences flanking the centromere of human chromosome 10 are a complex patchwork of arm-specific sequences, stable duplications, and unstable sequences with homologies to telomeric and other centromeric locations. Hum. Mol. Genet. 8, 205–215 (1999).
Horvath, J., Schwartz, S. & Eichler, E. The mosaic structure of a 2p11 pericentromeric segment: a strategy for characterizing complex regions of the human genome. Genome Res. 10, 839–852 (2000).
Horvath, J. et al. Molecular structure and evolution of an alpha/non-alpha satellite junction at 16p11. Hum. Mol. Genet. 9, 113–123 (2000).
Johnson, M.E. et al. Positive selection of a gene family during the emergence of humans and African apes. Nature 413, 514–519 (2001).
Stankiewicz, P. & Lupski, J.R. Genome architecture, rearrangements and genomic disorders. Trends Genet. 18, 74–82 (2002).
Horvath, J.E. et al. Punctuated duplication seeding events during the evolution of human chromosome 2p11. Genome Res. 15, 914–927 (2005).
Locke, D.P. et al. Molecular evolution of the human chromosome 15 pericentromeric region. Cytogenet. Genome Res. 108, 73–82 (2005).
Linardopoulou, E.V. et al. Human subtelomeres are hot spots of interchromosomal recombination and segmental duplication. Nature 437, 94–100 (2005).
Bailey, J.A. et al. Recent segmental duplications in the human genome. Science 297, 1003–1007 (2002).
She, X. et al. Shotgun sequence assembly and recent segmental duplications within the human genome. Nature 431, 927–930 (2004).
Pevzner, P.A., Tang, H. & Tesler, G. De novo repeat classification and fragment assembly. Genome Res. 14, 1786–1796 (2004).
Gibbs, R.A. et al. Evolutionary and biomedical insights from the rhesus macaque genome. Science 316, 222–234 (2007).
Waterston, R. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Gibbs, R.A. et al. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428, 493–521 (2004).
Lindblad-Toh, K. et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 438, 803–819 (2005).
Eichler, E.E. et al. Duplication of a gene-rich cluster between 16p11.1 and Xq28: a novel pericentromeric-directed mechanism for paralogous genome evolution. Hum. Mol. Genet. 5, 899–912 (1996).
Regnier, V. et al. Emergence and scattering of multiple neurofibromatosis (NF1)-related sequences during hominoid evolution suggest a process of pericentromeric interchromosomal transposition. Hum. Mol. Genet. 6, 9–16 (1997).
Potier, M. et al. Two sequence-ready contigs spanning the two copies of a 200-kb duplication on human 21q: partial sequence and polymorphisms. Genomics 51, 417–426 (1998).
She, X. et al. The structure and evolution of centromeric transition regions within the human genome. Nature 430, 857–864 (2004).
Kent, W.J., Baertsch, R., Hinrichs, A., Miller, W. & Haussler, D. Evolution's cauldron: duplication, deletion, and rearrangement in the mouse and human genomes. Proc. Natl. Acad. Sci. USA 100, 11484–11489 (2003).
Eichler, E.E. et al. Divergent origins and concerted expansion of two segmental duplications on chromosome 16. J. Hered. 92, 462–468 (2001).
Jackson, M.S. et al. Evidence for widespread reticulate evolution within human duplicons. Am. J. Hum. Genet. 77, 824–840 (2005).
Hurles, M.E. Gene conversion homogenizes the CMT1A paralogous repeats. BMC Genomics 2, 11 (2001).
Pavlicek, A., House, R., Gentles, A.J., Jurka, J. & Morrow, B.E. Traffic of genetic information between segmental duplications flanking the typical 22q11.2 deletion in velo-cardio-facial syndrome/DiGeorge syndrome. Genome Res. 15, 1487–1495 (2005).
Cheng, Z. et al. A genome-wide comparison of recent chimpanzee and human segmental duplications. Nature 437, 88–93 (2005).
Bowers, P.M., Cokus, S.J., Eisenberg, D. & Yeates, T.O. Use of logic relationships to decipher protein network organization. Science 306, 2246–2249 (2004).
Rivera, M.C. & Lake, J.A. The ring of life provides evidence for a genome fusion origin of eukaryotes. Nature 431, 152–155 (2004).
Lake, J.A. & Rivera, M.C. Deriving the genomic tree of life in the presence of horizontal gene transfer: conditioned reconstruction. Mol. Biol. Evol. 21, 681–690 (2004).
Rual, J.F. et al. Towards a proteome-scale map of the human protein-protein interaction network. Nature 437, 1173–1178 (2005).
Paulding, C.A., Ruvolo, M. & Haber, D.A. The Tre2 (USP6) oncogene is a hominoid-specific gene. Proc. Natl. Acad. Sci. USA 100, 2507–2511 (2003).
Vandepoele, K., Van Roy, N., Staes, K., Speleman, F. & van Roy, F. A novel gene family NBPF: intricate structure generated by gene duplications during primate evolution. Mol. Biol. Evol. 22, 2265–2274 (2005).
Ciccarelli, F.D. et al. Complex genomic rearrangements lead to novel primate gene function. Genome Res. 15, 343–351 (2005).
Gu, X., Wang, Y. & Gu, J. Age distribution of human gene families shows significant roles of both large- and small-scale duplications in vertebrate evolution. Nat. Genet. 31, 205–209 (2002).
Lynch, M. & Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 290, 1151–1155 (2000).
Horvath, J.E. et al. Using a pericentromeric interspersed repeat to recapitulate the phylogeny and expansion of human centromeric segmental duplications. Mol. Biol. Evol. 20, 1463–1479 (2003).
Johnson, M.E. et al. Recurrent duplication-driven transposition of DNA during hominoid evolution. Proc. Natl. Acad. Sci. USA 103, 17626–17631 (2006).
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Acknowledgements
We thank P. Green, J. Felsenstein, T. Newman, C. Alkan and Z. Bao for useful comments and valuable discussions in the preparation of this manuscript, and E. Tüzün and Z. Cheng for computational assistance. This work was supported by a US National Institutes of Health grant GM58815 to E.E.E. and a Rosetta Inpharmatics fellowship (Merck Laboratories) to Z.J. T.M.-B. is a research fellow supported by Departament d'Educació i Universitats de la Generalitat de Catalunya. E.E.E. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Z.J. performed the analyses and drafted the manuscript. H.T. implemented the program package and performed part of the analyses. M.V. and M.F.C. performed the FISH validation experiment. T.M.-B. performed the positive selection analysis on the core genes. X.S. was involved in part of the fusion gene analysis. P.A.P. and E.E.E. designed the study, and E.E.E. finalized the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Note (PDF 900 kb)
Rights and permissions
About this article
Cite this article
Jiang, Z., Tang, H., Ventura, M. et al. Ancestral reconstruction of segmental duplications reveals punctuated cores of human genome evolution. Nat Genet 39, 1361–1368 (2007). https://doi.org/10.1038/ng.2007.9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2007.9
This article is cited by
-
Multiscale analysis of pangenomes enables improved representation of genomic diversity for repetitive and clinically relevant genes
Nature Methods (2023)
-
Increased mutation and gene conversion within human segmental duplications
Nature (2023)
-
Fast characterization of segmental duplication structure in multiple genome assemblies
Algorithms for Molecular Biology (2022)
-
Modelling segmental duplications in the human genome
BMC Genomics (2021)
-
Systemic and rapid restructuring of the genome: a new perspective on punctuated equilibrium
Current Genetics (2021)