Abstract
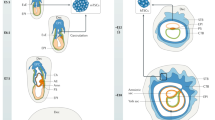
Eutherian placenta, an organ that emerged in the course of mammalian evolution, provides essential architecture, the so-called feto-maternal interface, for fetal development by exchanging nutrition, gas and waste between fetal and maternal blood. Functional defects of the placenta cause several developmental disorders, such as intrauterine growth retardation in humans and mice. A series of new inventions and/or adaptations must have been necessary to form and maintain eutherian chorioallantoic placenta, which consists of capillary endothelial cells and a surrounding trophoblast cell layer(s)1. Although many placental genes have been identified2, it remains unknown how the feto-maternal interface is formed and maintained during development, and how this novel design evolved. Here we demonstrate that retrotransposon-derived Rtl1 (retrotransposon-like 1), also known as Peg11 (paternally expressed 11), is essential for maintenance of the fetal capillaries, and that both its loss and its overproduction cause late-fetal and/or neonatal lethality in mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Rossant, J. & Cross, J.C. Placental development: lessons from mouse mutants. Nat. Rev. Genet. 2, 538–548 (2001).
Watson, E.D. & Cross, J.C. Development of structures and transport functions in the mouse placenta. Physiology (Bethesda) 20, 180–193 (2005).
Seitz, H. et al. Imprinted microRNA genes transcribed antisense to a reciprocally imprinted retrotransposon-like gene. Nat. Genet. 34, 261–262 (2003).
Cattanach, B.M. & Rasverry, C.V. Evidence of imprinting involving the distal region of Chr 12. Mouse Genome 91, 858 (1993).
Georgiades, P., Watkins, M., Surani, M.A. & Ferguson-Smith, A.C. Parental origin-specific developmental defects in mice with uniparental disomy for chromosome 12. Development 127, 4719–4728 (2000).
Tevendale, M., Watkins, M., Rasberry, C., Cattanach, B. & Ferguson-Smith, A.C. Analysis of mouse conceptuses with uniparental duplication/deficiency for distal chromosome 12: comparison with chromosome 12 uniparental disomy and implications for genomic imprinting. Cytogenet. Genome Res. 113, 215–222 (2006).
Davis, E. et al. RNAi-mediated allelic trans-interaction at the imprinted Rtl1/Peg11 locus. Curr. Biol. 15, 743–749 (2005).
Takada, S. et al. Epigenetic analysis of the Dlk1-Gtl2 imprinted domain on mouse chromosome 12: implications for imprinting control from comparison with Igf2-H19. Hum. Mol. Genet. 11, 77–86 (2002).
Lin, S.P. et al. Asymmetric regulation of imprinting on the maternal and paternal chromosomes at the Dlk1-Gtl2 imprinted cluster on mouse chromosome 12. Nat. Genet. 35, 97–102 (2003).
Constância, M. et al. Placental-specific IGF-II is a major modulator of placental and fetal growth. Nature 417, 945–948 (2002).
Sibley, C.P. et al. Placental-specific insulin-like growth factor 2 (Igf2) regulates the diffusional exchange characteristics of the mouse placenta. Proc. Natl. Acad. Sci. USA 101, 8204–8208 (2004).
Angiolini, E. et al. Regulation of placental efficiency for nutrient transport by imprinted genes. Placenta 27 (Suppl. A), S98–S102 (2006).
Moon, Y.S. et al. Mice lacking paternally expressed Pref-1/Dlk1 display growth retardation and accelerated adiposity. Mol. Cell. Biol. 22, 5585–5592 (2002).
Hernandez, A., Martinez, M.E., Fiering, S., Galton, V.A. & St Germain, D. Type 3 deiodinase is critical for the maturation and function of the thyroid axis. J. Clin. Invest. 116, 476–484 (2006).
Georgiades, P., Watkins, M., Burton, G.J. & Ferguson-Smith, A.C. Roles for genomic imprinting and the zygotic genome in placental development. Proc. Natl. Acad. Sci. USA 98, 4522–4527 (2001).
Butler, M., Goodwin, T., Simpson, M., Singh, M. & Poulter, R. Vertebrate LTR retrotransposons of the Tf1/sushi group. J. Mol. Evol. 52, 260–274 (2001).
Lynch, C. & Tristem, M. A co-opted gypsy-type LTR-retrotransposon is conserved in the genomes of humans, sheep, mice, and rats. Curr. Biol. 13, 1518–1523 (2003).
Ono, R. et al. Deletion of Peg10, an imprinted gene acquired from a retrotransposon, causes early embryonic lethality. Nat. Genet. 38, 101–106 (2006).
Brandt, J. et al. Transposable elements as a source of genetic innovation: expression and evolution of a family of retrotransposon-derived neogenes in mammals. Gene 345, 101–111 (2005).
Youngson, N.A., Kocialkowski, S., Peel, N. & Ferguson-Smith, A.C. A small family of sushi-class retrotransposon-derived genes in mammals and their relation to genomic imprinting. J. Mol. Evol. 61, 481–490 (2005).
Kagami, M. et al. Deletions and epimutations affecting the human 14q32.2 imprinted region in individuals with paternal and maternal upd(14)-like phenotypes. Nat. Genet. advance online publication, 10.1038/ng.2007.56 (6 January 2008).
Gould, S. & Vrba, S. Exaptation—a missing term in the science of form. Paleobiology 8, 4–15 (1982).
Brosius, J. & Gould, S.J. On “genomenclature”: a comprehensive (and respectful) taxonomy for pseudogenes and other “junk DNA”. Proc. Natl. Acad. Sci. USA 89, 10706–10710 (1992).
Smit, A.F. Interspersed repeats and other mementos of transposable elements in mammalian genomes. Curr. Opin. Genet. Dev. 9, 657–663 (1999).
Mi, S. et al. Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature 403, 785–789 (2000).
Kazazian, H.H., Jr. Mobile elements: drivers of genome evolution. Science 303, 1626–1632 (2004).
Dupressoir, A. et al. Syncytin-A and syncytin-B, two fusogenic placenta-specific murine envelope genes of retroviral origin conserved in Muridae. Proc. Natl. Acad. Sci. USA 102, 725–730 (2005).
Bejerano, G. et al. A distal enhancer and an ultraconserved exon are derived from a novel retroposon. Nature 441, 87–90 (2006).
Biémont, C. & Vieira, C. Genetics: junk DNA as an evolutionary force. Nature 443, 521–524 (2006).
Sekita, Y. et al. Aberrant regulation of imprinted gene expression in Gtl2lacZ mice. Cytogenet. Genome Res. 113, 223–229 (2006).
Acknowledgements
We thank S. Aizawa of Center for Developmental Biology, RIKEN for providing the DT-A vector that was used for making Rtl1 KO construct, E. Robertson of University of Oxford for the CCE ES cells, M. Constancia of University of Cambridge for placenta functional assay protocol, Y. Nakahara and M. Takabe of the Mitsubishi Kagaku Institute of Life Sciences for animal breeding and H. Hasegawa, N. Kawabe and A. Akatsuka of the Tokai University and S. Ichinose of the Tokyo Medical and Dental University for their assistance in immunohistochemistry and electron microscopy along with helpful discussion. This work was supported by grants from Creative Science Research, the research program of Japan Society for the Promotion of Science (JSPS), the Uehara Memorial Science Foundation, the Mitsubishi Foundation and the Ministry of Health, Labour and Welfare for Child Health and Development (17C-2) and a Grant-in-Aid for Scientific Research on Priority Areas form the Ministry of Education, Culture, Sports, Science and Technology of Japan (1508023) to F.I., the grant for young investigators from Medical Research Institute to Y.S., and the Asahi Glass Foundation and JSPS, Grants-in Aid for Scientific Research to T.K.-I. Pacific Edit reviewed the manuscript before submission.
Author information
Authors and Affiliations
Contributions
Most analyses in this work were performed by Y.S. with collaboration of H.W., R.O., N.W., T.K. and M.K. in molecular and histological experiments. Construction of Rtl1 targeting vector was done by H.W. under supervision of K.N. and M.Y., and KO mice were produced by T.H., R.S.-M., K.N. and M.Y. The study was designed and coordinated by T.K.-I. and F.I., and the results were discussed by A.O., T.O., T.K.-I. and F.I. The paper was written by Y.S. and F.I.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–3 (PDF 1297 kb)
Rights and permissions
About this article
Cite this article
Sekita, Y., Wagatsuma, H., Nakamura, K. et al. Role of retrotransposon-derived imprinted gene, Rtl1, in the feto-maternal interface of mouse placenta. Nat Genet 40, 243–248 (2008). https://doi.org/10.1038/ng.2007.51
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2007.51
This article is cited by
-
Maternal RNA transcription in Dlk1-Dio3 domain is critical for proper development of the mouse placental vasculature
Communications Biology (2024)
-
Discovery of a novel marker for human granulocytes and tissue macrophages: RTL1 revisited
Cell and Tissue Research (2023)
-
Are there foetal extracellular vesicles in maternal blood? Prospects for diagnostic biomarker discovery
Journal of Molecular Medicine (2023)
-
Epigenetics Beyond Fetal Growth Restriction: A Comprehensive Overview
Molecular Diagnosis & Therapy (2022)
-
AKT signaling is associated with epigenetic reprogramming via the upregulation of TET and its cofactor, alpha-ketoglutarate during iPSC generation
Stem Cell Research & Therapy (2021)