Abstract
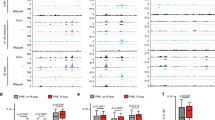
It is widely assumed that the key rate-limiting step in gene activation is the recruitment of RNA polymerase II (Pol II) to the core promoter1. Although there are well-documented examples in which Pol II is recruited to a gene but stalls2,3,4,5,6,7,8,9,10,11,12, a general role for Pol II stalling in development has not been established. We have carried out comprehensive Pol II chromatin immunoprecipitation microarray (ChIP-chip) assays in Drosophila embryos and identified three distinct Pol II binding behaviors: active (uniform binding across the entire transcription unit), no binding, and stalled (binding at the transcription start site). The notable feature of the ∼10% genes that are stalled is that they are highly enriched for developmental control genes, which are either repressed or poised for activation during later stages of embryogenesis. We propose that Pol II stalling facilitates rapid temporal and spatial changes in gene activity during development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Ptashne, M. & Gann, A. Transcriptional activation by recruitment. Nature 386, 569–577 (1997).
Conaway, J.W., Shilatifard, A., Dvir, A. & Conaway, R.C. Control of elongation by RNA polymerase II. Trends Biochem. Sci. 25, 375–380 (2000).
Saunders, A., Core, L.J. & Lis, J.T. Breaking barriers to transcription elongation. Nat. Rev. Mol. Cell Biol. 7, 557–567 (2006).
Gilmour, D.S. & Lis, J.T. RNA polymerase II interacts with the promoter region of the noninduced hsp70 gene in Drosophila melanogaster cells. Mol. Cell. Biol. 6, 3984–3989 (1986).
Bender, T.P., Thompson, C.B. & Kuehl, W.M. Differential expression of c-myb mRNA in murine B lymphomas by a block to transcription elongation. Science 237, 1473–1476 (1987).
Strobl, L.J. & Eick, D. Hold back of RNA polymerase II at the transcription start site mediates down-regulation of c-myc in vivo. EMBO J. 11, 3307–3314 (1992).
Krumm, A., Meulia, T., Brunvand, M. & Groudine, M. The block to transcriptional elongation within the human c-myc gene is determined in the promoter-proximal region. Genes Dev. 6, 2201–2213 (1992).
Laspia, M.F., Wendel, P. & Mathews, M.B. HIV-1 Tat overcomes inefficient transcriptional elongation in vitro. J. Mol. Biol. 232, 732–746 (1993).
Cheng, C. & Sharp, P.A. RNA polymerase II accumulation in the promoter-proximal region of the dihydrofolate reductase and gamma-actin genes. Mol. Cell. Biol. 23, 1961–1967 (2003).
Aida, M. et al. Transcriptional pausing caused by NELF plays a dual role in regulating immediate-early expression of the junB gene. Mol. Cell. Biol. 26, 6094–6104 (2006).
Radonjic, M. et al. Genome-wide analyses reveal RNA polymerase II located upstream of genes poised for rapid response upon S. cerevisiae stationary phase exit. Mol. Cell 18, 171–183 (2005).
Wang, X., Lee, C., Gilmour, D.S. & Gergen, J.P. Transcription elongation controls cell fate specification in the Drosophila embryo. Genes Dev. 21, 1031–1036 (2007).
Lis, J. & Wu, C. Protein traffic on the heat shock promoter: parking, stalling, and trucking along. Cell 74, 1–4 (1993).
Reppas, N.B., Wade, J.T., Church, G.M. & Struhl, K. The transition between transcriptional initiation and elongation in E. coli is highly variable and often rate limiting. Mol. Cell 24, 747–757 (2006).
Agalioti, T. et al. Ordered recruitment of chromatin modifying and general transcription factors to the IFN-beta promoter. Cell 103, 667–678 (2000).
Soutoglou, E. & Talianidis, I. Coordination of PIC assembly and chromatin remodeling during differentiation-induced gene activation. Science 295, 1901–1904 (2002).
Schneider, D.S., Hudson, K.L., Lin, T.Y. & Anderson, K.V. Dominant and recessive mutations define functional domains of Toll, a transmembrane protein required for dorsal-ventral polarity in the Drosophila embryo. Genes Dev. 5, 797–807 (1991).
Leptin, M. twist and snail as positive and negative regulators during Drosophila mesoderm development. Genes Dev. 5, 1568–1576 (1991).
Kosman, D., Ip, Y.T., Levine, M. & Arora, K. Establishment of the mesoderm-neuroectoderm boundary in the Drosophila embryo. Science 254, 118–122 (1991).
Furlong, E.E., Andersen, E.C., Null, B., White, K.P. & Scott, M.P. Patterns of gene expression during Drosophila mesoderm development. Science 293, 1629–1633 (2001).
Stathopoulos, A., Van Drenth, M., Erives, A., Markstein, M. & Levine, M. Whole-genome analysis of dorsal-ventral patterning in the Drosophila embryo. Cell 111, 687–701 (2002).
Biemar, F. et al. Comprehensive identification of Drosophila dorsal-ventral patterning genes using a whole-genome tiling array. Proc. Natl. Acad. Sci. USA 103, 12763–12768 (2006).
Zeitlinger, J. et al. Whole-genome ChIP-chip analysis of Dorsal, Twist, and Snail suggests integration of diverse patterning processes in the Drosophila embryo. Genes Dev. 21, 385–390 (2007).
Markstein, M., Markstein, P., Markstein, V. & Levine, M.S. Genome-wide analysis of clustered Dorsal binding sites identifies putative target genes in the Drosophila embryo. Proc. Natl. Acad. Sci. USA 99, 763–768 (2002).
Tomancak, P. et al. Systematic determination of patterns of gene expression during Drosophila embryogenesis. Genome Biol. 3, research0088.1–research0088.14 (2002).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Huang, A.M., Rusch, J. & Levine, M. An anteroposterior Dorsal gradient in the Drosophila embryo. Genes Dev. 11, 1963–1973 (1997).
Boulay, J.L., Dennefeld, C. & Alberga, A. The Drosophila developmental gene snail encodes a protein with nucleic acid binding fingers. Nature 330, 395–398 (1987).
Sandmann, T. et al. A temporal map of transcription factor activity: mef2 directly regulates target genes at all stages of muscle development. Dev. Cell 10, 797–807 (2006).
Furlong, E.E. Integrating transcriptional and signalling networks during muscle development. Curr. Opin. Genet. Dev. 14, 343–350 (2004).
Acknowledgements
We would like to thank R. Zinzen for collecting the Toll10b, Tollrm9/rm10 and gd7 embryos, T. Volkert and J. Love for microarray experimental support and members of the Young laboratory for critical reading of the manuscript. This research was supported in part by the Intramural Research Program of the US National Institutes of Health, National Institute of Environmental Health Sciences (K.A.), US National Institutes of Health grants HG002668 and GM069676 to R.A.Y. and GM34431 to M.L., a grant by the Moore Foundation and a postdoctoral fellowship by the Human Frontier Science Program Organization (HFSPO, A.S.).
Author information
Authors and Affiliations
Contributions
J.Z. and M.L. designed the experiment. J.Z. designed the arrays and carried out the experiments and analysis. A.S., M.K. and J.Z. analyzed expression data and functional categories. J.-W.H., S.N. and K.A. carried out the permanganate footprint assays. J.Z., M.L. and R.A.Y. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
One of the authors (R.A.Y.) consults for Agilent Technologies.
Supplementary information
Supplementary Text and Figures
Supplementary Note (PDF 713 kb)
Rights and permissions
About this article
Cite this article
Zeitlinger, J., Stark, A., Kellis, M. et al. RNA polymerase stalling at developmental control genes in the Drosophila melanogaster embryo. Nat Genet 39, 1512–1516 (2007). https://doi.org/10.1038/ng.2007.26
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2007.26
This article is cited by
-
Tissue-specific RNA Polymerase II promoter-proximal pause release and burst kinetics in a Drosophila embryonic patterning network
Genome Biology (2024)
-
Lola-I is a promoter pioneer factor that establishes de novo Pol II pausing during development
Nature Communications (2023)
-
Regulation, functions and transmission of bivalent chromatin during mammalian development
Nature Reviews Molecular Cell Biology (2023)
-
PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
Nature Communications (2023)
-
Genomic regulation of transcription and RNA processing by the multitasking Integrator complex
Nature Reviews Molecular Cell Biology (2023)