Abstract
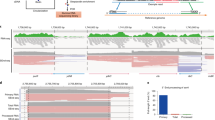
We have determined the high-resolution strand-specific transcriptome of the fission yeast S. pombe under multiple growth conditions using a novel RNA-DNA hybridization mapping (HybMap) technique. HybMap uses an antibody against an RNA-DNA hybrid to detect RNA molecules hybridized to a high-density DNA oligonucleotide tiling microarray. HybMap showed exceptional dynamic range and reproducibility, and allowed us to identify strand-specific coding, noncoding and structural RNAs, as well as previously unknown RNAs conserved in distant yeast species. Notably, we found that virtually the entire euchromatic genome (including intergenics) is transcribed, with heterochromatin dampening intergenic transcription. We identified features including large numbers of condition-specific noncoding RNAs, extensive antisense transcription, new properties of antisense transcripts and induced divergent transcription. Furthermore, our HybMap data informed the efficiency and locations of RNA splicing genome-wide. Finally, we observed strand-specific transcription islands around tRNAs at heterochromatin boundaries inside centromeres. Here, we discuss these new features in terms of organism fitness and transcriptome evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Okazaki, Y. et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420, 563–573 (2002).
David, L. et al. A high-resolution map of transcription in the yeast genome. Proc. Natl. Acad. Sci. USA 103, 5320–5325 (2006).
Washietl, S., Hofacker, I.L., Lukasser, M., Huttenhofer, A. & Stadler, P.F. Mapping of conserved RNA secondary structures predicts thousands of functional noncoding RNAs in the human genome. Nat. Biotechnol. 23, 1383–1390 (2005).
Kampa, D. et al. Novel RNAs identified from an in-depth analysis of the transcriptome of human chromosomes 21 and 22. Genome Res. 14, 331–342 (2004).
Steigele, S., Huber, W., Stocsits, C., Stadler, P.F. & Nieselt, K. Comparative analysis of structured RNAs in S. cerevisiae indicates a multitude of different functions. BMC Biol. 5, 25 (2007).
Struhl, K. Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nat. Struct. Mol. Biol. 14, 103–105 (2007).
Perocchi, F., Xu, Z., Clauder-Munster, S. & Steinmetz, L.M. Antisense artifacts in transcriptome microarray experiments are resolved by actinomycin D. Nucleic Acids Res. 35, e128 (2007).
Hu, Z., Zhang, A., Storz, G., Gottesman, S. & Leppla, S.H. An antibody-based microarray assay for small RNA detection. Nucleic Acids Res. 34, e52 (2006).
Boguslawski, S.J. et al. Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J. Immunol. Methods 89, 123–130 (1986).
Kapranov, P., Willingham, A.T. & Gingeras, T.R. Genome-wide transcription and the implications for genomic organization. Nat. Rev. Genet. 8, 413–423 (2007).
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007).
Wood, V. et al. The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880 (2002).
Leonardi, J., Box, J.A., Bunch, J.T. & Baumann, P. TER1, the RNA subunit of fission yeast telomerase. Nat. Struct. Mol. Biol. 15, 26–33 (2008).
Webb, C.J. & Zakian, V.A. Identification and characterization of the Schizosaccharomyces pombe TER1 telomerase RNA. Nat. Struct. Mol. Biol. 15, 34–42 (2008).
Gordon, M. et al. Genome-wide dynamics of SAPHIRE, an essential complex for gene activation and chromatin boundaries. Mol. Cell. Biol. 27, 4058–4069 (2007).
Chen, D. et al. Global transcriptional responses of fission yeast to environmental stress. Mol. Biol. Cell 14, 214–229 (2003).
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Nicolas, E. et al. Distinct roles of HDAC complexes in promoter silencing, antisense suppression and DNA damage protection. Nat. Struct. Mol. Biol. 14, 372–380 (2007).
Wiren, M. et al. Genomewide analysis of nucleosome density histone acetylation and HDAC function in fission yeast. EMBO J. 24, 2906–2918 (2005).
Noma, K., Cam, H.P., Maraia, R.J. & Grewal, S.I. A role for TFIIIC transcription factor complex in genome organization. Cell 125, 859–872 (2006).
Volpe, T.A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Allshire, R.C., Javerzat, J.P., Redhead, N.J. & Cranston, G. Position effect variegation at fission yeast centromeres. Cell 76, 157–169 (1994).
Takahashi, K. et al. A low copy number central sequence with strict symmetry and unusual chromatin structure in fission yeast centromere. Mol. Biol. Cell 3, 819–835 (1992).
Scott, K.C., White, C.V. & Willard, H.F. An RNA polymerase III-dependent heterochromatin barrier at fission yeast centromere 1. PLoS ONE 2, e1099 (2007).
Baum, M., Ngan, V.K. & Clarke, L. The centromeric K-type repeat and the central core are together sufficient to establish a functional Schizosaccharomyces pombe centromere. Mol. Biol. Cell 5, 747–761 (1994).
Partridge, J.F., Scott, K.S., Bannister, A.J., Kouzarides, T. & Allshire, R.C. cis-acting DNA from fission yeast centromeres mediates histone H3 methylation and recruitment of silencing factors and cohesin to an ectopic site. Curr. Biol. 12, 1652–1660 (2002).
Steiner, N.C., Hahnenberger, K.M. & Clarke, L. Centromeres of the fission yeast Schizosaccharomyces pombe are highly variable genetic loci. Mol. Cell. Biol. 13, 4578–4587 (1993).
Clarke, L., Amstutz, H., Fishel, B. & Carbon, J. Analysis of centromeric DNA in the fission yeast Schizosaccharomyces pombe. Proc. Natl. Acad. Sci. USA 83, 8253–8257 (1986).
Nakaseko, Y., Kinoshita, N. & Yanagida, M. A novel sequence common to the centromere regions of Schizosaccharomyces pombe chromosomes. Nucleic Acids Res. 15, 4705–4715 (1987).
Nakaseko, Y., Adachi, Y., Funahashi, S., Niwa, O. & Yanagida, M. Chromosome walking shows a highly homologous repetitive sequence present in all the centromere regions of fission yeast. EMBO J. 5, 1011–1021 (1986).
Motamedi, M.R. et al. Two RNAi complexes, RITS and RDRC, physically interact and localize to noncoding centromeric RNAs. Cell 119, 789–802 (2004).
Lindsey-Boltz, L.A. & Sancar, A. RNA polymerase: the most specific damage recognition protein in cellular responses to DNA damage? Proc. Natl. Acad. Sci. USA 104, 13213–13214 (2007).
Cam, H.P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37, 809–819 (2005).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Reinhart, B.J. & Bartel, D.P. Small RNAs correspond to centromere heterochromatic repeats. Science 297, 1831 (2002).
Kato, H. et al. RNA polymerase II is required for RNAi-dependent heterochromatin assembly. Science 309, 467–469 (2005).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Acknowledgements
We thank S. Leppla (US National Institutes of Health) for generously providing the S9.6 antibody and Bob Schackmann (University of Utah) for oligo synthesis. The work was supported by the Howard Hughes Medical Institute (B.R.C., D.H., T.J.P., and supplies), US National Institutes of Health Genetics Training Grant T32 GM007464 (N.D.), the Huntsman Cancer Institute (D.A.N.), and CA24014 (for core facilities).
Author information
Authors and Affiliations
Contributions
N.D., D.A.N., and B.R.C.; system design and experimental approaches. B.D.; array method optimization. D.A.N., N.D., B.M., T.J.P., D.H., and B.R.C.; array design, data analysis methods and data analysis. E.W.; feature computation. N.D., D.A.N., and D.H.; figures. B.R.C., N.D. and D.A.N. wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1–11, Supplementary Note, Supplementary Methods (PDF 1308 kb)
Rights and permissions
About this article
Cite this article
Dutrow, N., Nix, D., Holt, D. et al. Dynamic transcriptome of Schizosaccharomyces pombe shown by RNA-DNA hybrid mapping. Nat Genet 40, 977–986 (2008). https://doi.org/10.1038/ng.196
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.196
This article is cited by
-
S9.6-based hybrid capture immunoassay for pathogen detection
Scientific Reports (2023)
-
Structural basis of R-loop recognition by the S9.6 monoclonal antibody
Nature Communications (2022)
-
The Antisense Transcriptome and the Human Brain
Journal of Molecular Neuroscience (2016)
-
Highly condensed chromatins are formed adjacent to subtelomeric and decondensed silent chromatin in fission yeast
Nature Communications (2015)
-
RNAi triggered by specialized machinery silences developmental genes and retrotransposons
Nature (2013)