Abstract
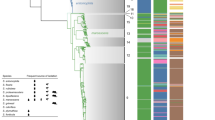
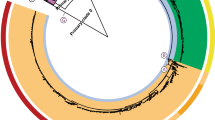
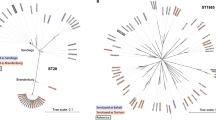
Isolates of Salmonella enterica serovar Typhi (Typhi), a human-restricted bacterial pathogen that causes typhoid, show limited genetic variation. We generated whole-genome sequences for 19 Typhi isolates using 454 (Roche) and Solexa (Illumina) technologies. Isolates, including the previously sequenced CT18 and Ty2 isolates, were selected to represent major nodes in the phylogenetic tree. Comparative analysis showed little evidence of purifying selection, antigenic variation or recombination between isolates. Rather, evolution in the Typhi population seems to be characterized by ongoing loss of gene function, consistent with a small effective population size. The lack of evidence for antigenic variation driven by immune selection is in contrast to strong adaptive selection for mutations conferring antibiotic resistance in Typhi. The observed patterns of genetic isolation and drift are consistent with the proposed key role of asymptomatic carriers of Typhi as the main reservoir of this pathogen, highlighting the need for identification and treatment of carriers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Parry, C.M., Hien, T.T., Dougan, G., White, N.J. & Farrar, J.J. Typhoid fever. N. Engl. J. Med. 347, 1770–1782 (2002).
Coburn, B., Grassl, G.A. & Finlay, B.B. Salmonella, the host and disease: a brief review. Immunol. Cell Biol. 85, 112–118 (2007).
Parkhill, J. et al. Complete genome sequence of a multiple drug resistant Salmonella enterica serovar Typhi CT18. Nature 413, 848–852 (2001).
Pickard, D. et al. Composition, acquisition, and distribution of the Vi exopolysaccharide-encoding Salmonella enterica pathogenicity island SPI-7. J. Bacteriol. 185, 5055–5065 (2003).
Roumagnac, P. et al. Evolutionary history of Salmonella Typhi. Science 314, 1301–1304 (2006).
Chau, T.T. et al. Antimicrobial drug resistance of Salmonella enterica serovar Typhi in Asia and molecular mechanism of reduced susceptibility to the fluoroquinolones. Antimicrob. Agents Chemother. 51, 4315–4323 (2007).
Le, T.A. et al. Clonal expansion and microevolution of quinolone-resistant Salmonella enterica serotype Typhi in Vietnam from 1996 to 2004. J. Clin. Microbiol. 45, 3485–3492 (2007).
Deng, W. et al. Comparative genomics of Salmonella enterica serovar Typhi strains Ty2 and CT18. J. Bacteriol. 185, 2330–2337 (2003).
Hall, N. Advanced sequencing technologies and their wider impact in microbiology. J. Exp. Biol. 210, 1518–1525 (2007).
Pearson, T. et al. Phylogenetic discovery bias in Bacillus anthracis using single-nucleotide polymorphisms from whole-genome sequencing. Proc. Natl. Acad. Sci. USA 101, 13536–13541 (2004).
Baker, S. et al. High-throughput genotyping of Salmonella Typhi allows geographical assignment of haplotypes and pathotypes within an urban district of Jakarta, Indonesia. J. Clin. Microbiol. 46, 1741–1746 (2008).
Rocha, E.P.C. et al. Comparisons of dN/dS are time dependent for closely related bacterial genomes. J. Theor. Biol. 239, 226–235 (2006).
Turner, A.K., Nair, S. & Wain, J. The acquisition of full fluoroquinolone resistance in Salmonella Typhi by accumulation of point mutations in the topoisomerase targets. J. Antimicrob. Chemother. 58, 733–740 (2006).
Haraga, A., Ohlson, M.B. & Miller, S.I. Salmonellae interplay with host cells. Nat. Rev. Microbiol. 6, 53–66 (2008).
Falush, D. & Bowden, R. Genome-wide association mapping in bacteria? Trends Microbiol. 14, 353–355 (2006).
Didelot, X., Achtman, M., Parkhill, J., Thomson, N.R. & Falush, D. A bimodal pattern of relatedness between the Salmonella Paratyphi A and Typhi genomes: convergence or divergence by homologous recombination? Genome Res. 17, 61–68 (2007).
Thomson, N. et al. The role of prophage-like elements in the diversity of Salmonella enterica serovars. J. Mol. Biol. 339, 279–300 (2004).
Vernikos, G.S. & Parkhill, J. Interpolated variable order motifs for identification of horizontally acquired DNA: revisiting the Salmonella pathogenicity islands. Bioinformatics 22, 2196–2203 (2006).
Boyd, E.F., Porwollik, S., Blackmer, F. & McClelland, M. Differences in gene content among Salmonella enterica serovar Typhi isolates. J. Clin. Microbiol. 41, 3823–3828 (2003).
Nair, S. et al. Salmonella enterica serovar Typhi strains from which SPI7, a 134-kilobase island with genes for Vi exopolysaccharide and other functions, has been deleted. J. Bacteriol. 186, 3214–3223 (2004).
Bueno, S.M. et al. Precise excision of the large pathogenicity island, SPI7, in Salmonella enterica serovar Typhi. J. Bacteriol. 186, 3202–3213 (2004).
Bertram, G., Innes, S., Minella, O., Richardson, J.P. & Stansfield, I. Endless possibilities: translation termination and stop codon recognition. Microbiology 147, 255–269 (2001).
Thomson, N.R. et al. The complete genome sequence and comparative genome analysis of the high pathogenicity Yersinia enterocolitica strain 8081. PLoS Genet. 2, e206 (2006).
Parkhill, J. et al. Comparative analysis of the genome sequences of Bordetella pertussis, Bordetella parapertussis and Bordetella bronchiseptica. Nat. Genet. 35, 32–40 (2003).
Cole, S.T. et al. Massive gene decay in the leprosy bacillus. Nature 409, 1007–1011 (2001).
Andersson, J.O. & Andersson, S.G.E. Genome degradation is an ongoing process in Rickettsia. Mol. Biol. Evol. 16, 1178–1191 (1999).
Hiller, N. L. et al. Comparative genomic analyses of seventeen Streptococcus pneumoniae strains: insights into the pneumococcal supragenome. J. Bacteriol. 189, 8186–8195 (2007).
Tettelin, H. et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome”. Proc. Natl. Acad. Sci. USA 102, 13950–13955 (2005).
Octavia, S. & Lan, R. Single nucleotide polymorphism typing and genetic relationships of Salmonella enterica serovar Typhi isolates. J. Clin. Microbiol. 45, 3795–3801 (2007).
Saurin, W., Hofnung, M. & Dassa, E. Getting in or out: early segregation between importers and exporters in the evolution of ATP-binding cassette (ABC) transporters. J. Mol. Evol. 48, 22–41 (1999).
Webber, M.A. & Piddock, L.J. The importance of efflux pumps in bacterial antibiotic resistance. J. Antimicrob. Chemother. 51, 9–11 (2003).
Suerbaum, S. et al. Free recombination within Helicobacter pylori. Proc. Natl. Acad. Sci. USA 95, 12619–12624 (1998).
Gomes, J.P. et al. Evolution of Chlamydia trachomatis diversity occurs by widespread interstrain recombination involving hotspots. Genome Res. 17, 50–60 (2007).
Gey Van Pittius, N.C. et al. Evolution and expansion of the Mycobacterium tuberculosis PE and PPE multigene families and their association with the duplication of the ESAT-6 (esx) gene cluster regions. BMC Evol. Biol. 6, 95 (2006).
Vaishnavi, C. et al. Epidemiology of typhoid carriers among blood donors and patients with biliary, gastrointestinal and other related diseases. Microbiol. Immunol. 49, 107–112 (2005).
Levine, M.M., Black, R.E. & Lanata, C. Precise estimation of the number of chronic carriers of Salmonella typhi in Santiago, Chile, an endemic area. J. Infect. Dis. 146, 724–726 (1982).
Lewis, M.D. et al. Typhoid fever: a massive, single-point source, multidrug-resistant outbreak in Nepal. Clin. Infect. Dis. 40, 554–561 (2005).
Sears, S.D., Ferreccio, C. & Levine, M.M. The use of Moore swabs for isolation of Salmonella typhi from irrigation water in Santiago, Chile. J. Infect. Dis. 149, 640–642 (1984).
Cho, J.C. & Kim, S.J. Viable, but non-culturable, state of a green fluorescence protein-tagged environmental isolate of Salmonella typhi in groundwater and pond water. FEMS Microbiol. Lett. 170, 257–264 (1999).
Sokurenko, E.V., Gomulkiewicz, R. & Dykhuizen, D.E. Source-sink dynamics of virulence evolution. Nat. Rev. Microbiol. 4, 548–555 (2006).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Acknowledgements
This work was supported by the Wellcome Trust. M.A. and C.J.M. are supported in Ireland by grant 05/FE1/B882 from the Scientific Foundation Ireland and C.J.M. was supported in Berlin by a Wellcome Trust grant to J. Farrar. We gratefully acknowledge the support of the Sanger Institute core sequencing and informatics groups. Isolates were provided by the Oxford University Clinical Research Unit (CT18, J185SM, AG3); B. Holmes at the National Collection of Type Cultures (M223); the Wellcome Trust Sanger Institute (404ty, Ty2); and F.-X.W. (all other isolates).
Author information
Authors and Affiliations
Contributions
G.D., J.P., M.A., P.R. and J.W. designed the study; F.-X.W. and C.D. contributed isolates for analysis; I.G. and R.R. performed 454 and Solexa sequencing; K.E.H. and S.B. performed validation experiments; D.J.M. co-supervises the PhD studies of K.E.H. and contributed to experimental design; K.E.H. and C.J.M. analysed data and K.E.H., J.P., P.R. and G.D. wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1 and 3, Supplementary Methods, Supplementary Note (PDF 1448 kb)
Rights and permissions
About this article
Cite this article
Holt, K., Parkhill, J., Mazzoni, C. et al. High-throughput sequencing provides insights into genome variation and evolution in Salmonella Typhi. Nat Genet 40, 987–993 (2008). https://doi.org/10.1038/ng.195
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.195
This article is cited by
-
A global resource for genomic predictions of antimicrobial resistance and surveillance of Salmonella Typhi at pathogenwatch
Nature Communications (2021)
-
Population genomics provides insights into the evolution and adaptation to humans of the waterborne pathogen Mycobacterium kansasii
Nature Communications (2021)
-
Comparative genomics and pangenome-oriented studies reveal high homogeneity of the agronomically relevant enterobacterial plant pathogen Dickeya solani
BMC Genomics (2020)
-
Chromosomal assembly and analyses of genome-wide recombination rates in the forest pathogenic fungus Armillaria ostoyae
Heredity (2020)
-
Cryptic speciation of a pelagic Roseobacter population varying at a few thousand nucleotide sites
The ISME Journal (2020)