Abstract
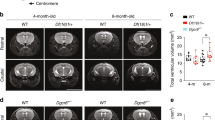
Individuals with 22q11.2 microdeletions show behavioral and cognitive deficits and are at high risk of developing schizophrenia. We analyzed an engineered mouse strain carrying a chromosomal deficiency spanning a segment syntenic to the human 22q11.2 locus. We uncovered a previously unknown alteration in the biogenesis of microRNAs (miRNAs) and identified a subset of brain miRNAs affected by the microdeletion. We provide evidence that the abnormal miRNA biogenesis emerges because of haploinsufficiency of the Dgcr8 gene, which encodes an RNA-binding moiety of the 'microprocessor' complex and contributes to the behavioral and neuronal deficits associated with the 22q11.2 microdeletion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Botto, L.D. et al. A population-based study of the 22q11.2 deletion: phenotype, incidence, and contribution to major birth defects in the population. Pediatrics 112, 101–107 (2003).
McDonald-McGinn, D.M. et al. Phenotype of the 22q11.2 deletion in individuals identified through an affected relative: cast a wide FISHing net!. Genet. Med. 3, 23–29 (2001).
Edelmann, L., Pandita, R.K. & Morrow, B.E. Low-copy repeats mediate the common 3-Mb deletion in patients with velo-cardio-facial syndrome. Am. J. Hum. Genet. 64, 1076–1086 (1999).
Woodin, M. et al. Neuropsychological profile of children and adolescents with the 22q11.2 microdeletion. Genet. Med. 3, 34–39 (2001).
Gothelf, D. et al. Genetic, developmental, and physical factors associated with attention deficit hyperactivity disorder in patients with velocardiofacial syndrome. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 126, 116–121 (2004).
Gothelf, D. et al. Obsessive-compulsive disorder in patients with velocardiofacial (22q11 deletion) syndrome. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 126, 99–105 (2004).
Marshall, C.R. et al. Structural variation of chromosomes in autism spectrum disorder. Am. J. Hum. Genet. 82, 477–488 (2008).
Green, M.F. et al. Approaching a consensus cognitive battery for clinical trials in schizophrenia: the NIMH-MATRICS conference to select cognitive domains and test criteria. Biol. Psychiatry 56, 301–307 (2004).
Sobin, C. et al. Neuropsychological characteristics of children with the 22q11 deletion syndrome: a descriptive analysis. Child Neuropsychol. 11, 39–53 (2005).
Pulver, A.E. et al. Psychotic illness in patients diagnosed with velo-cardio-facial syndrome and their relatives. J. Nerv. Ment. Dis. 182, 476–478 (1994).
Karayiorgou, M. et al. Schizophrenia susceptibility associated with interstitial deletions of chromosome 22q11. Proc. Natl. Acad. Sci. USA 92, 7612–7616 (1995).
Gothelf, D. et al. COMT genotype predicts longitudinal cognitive decline and psychosis in 22q11.2 deletion syndrome. Nat. Neurosci. 8, 1500–1502 (2005).
Liu, H. et al. Genetic variation in the 22q11 locus and susceptibility to schizophrenia. Proc. Natl. Acad. Sci. USA 99, 16859–16864 (2002).
Liu, H. et al. Genetic variation at the 22q11 PRODH2/DGCR6 locus presents an unusual pattern and increases susceptibility to schizophrenia. Proc. Natl. Acad. Sci. USA 99, 3717–3722 (2002).
Mukai, J. et al. Evidence that the gene encoding ZDHHC8 contributes to the risk of schizophrenia. Nat. Genet. 36, 725–731 (2004).
Paterlini, M. et al. Transcriptional and behavioral interaction between 22q11.2 orthologs modulates schizophrenia-related phenotypes in mice. Nat. Neurosci. 8, 1586–1594 (2005).
Raux, G. et al. Involvement of hyperprolinemia in cognitive and psychiatric features of the 22q11 deletion syndrome. Hum. Mol. Genet. 16, 83–91 (2007).
Williams, N.M. et al. Strong evidence that GNB1L is associated with schizophrenia. Hum. Mol. Genet. 17, 555–566 (2008).
Swerdlow, N.R., Geyer, M.A. & Braff, D.L. Neural circuit regulation of prepulse inhibition of startle in the rat: current knowledge and future challenges. Psychopharmacology (Berl.) 156, 194–215 (2001).
Gogos, J.A. et al. The gene encoding proline dehydrogenase modulates sensorimotor gating in mice. Nat. Genet. 21, 434–439 (1999).
Kimber, W.L. et al. Deletion of 150 kb in the minimal DiGeorge/velocardiofacial syndrome critical region in mouse. Hum. Mol. Genet. 8, 2229–2237 (1999).
Paylor, R. et al. Tbx1 haploinsufficiency is linked to behavioral disorders in mice and humans: implications for 22q11 deletion syndrome. Proc. Natl. Acad. Sci. USA 103, 7729–7734 (2006).
Lindsay, E.A. et al. Congenital heart disease in mice deficient for the DiGeorge syndrome region. Nature 401, 379–383 (1999).
Merscher, S. et al. TBX1 is responsible for cardiovascular defects in velo-cardio-facial/DiGeorge syndrome. Cell 104, 619–629 (2001).
Zheng, B., Mills, A.A. & Bradley, A. A system for rapid generation of coat color-tagged knockouts and defined chromosomal rearrangements in mice. Nucleic Acids Res. 27, 2354–2360 (1999).
Long, J.M. et al. Behavior of mice with mutations in the conserved region deleted in velocardiofacial/DiGeorge syndrome. Neurogenetics 7, 247–257 (2006).
Paylor, R. et al. Mice deleted for the DiGeorge/velocardiofacial syndrome region show abnormal sensorimotor gating and learning and memory impairments. Hum. Mol. Genet. 10, 2645–2650 (2001).
Phillips, R.G. & LeDoux, J.E. Differential contribution of amygdala and hippocampus to cued and contextual fear conditioning. Behav. Neurosci. 106, 274–285 (1992).
Koike, H., Arguello, P.A., Kvajo, M., Karayiorgou, M. & Gogos, J.A. Disc1 is mutated in the 129S6/SvEv strain and modulates working memory in mice. Proc. Natl. Acad. Sci. USA 103, 3693–3697 (2006).
Jones, M.W. & Wilson, M.A. Theta rhythms coordinate hippocampal-prefrontal interactions in a spatial memory task. PLoS Biol. 3, e402 (2005).
Kellendonk, C. et al. Transient and selective overexpression of dopamine D2 receptors in the striatum causes persistent abnormalities in prefrontal cortex functioning. Neuron 49, 603–615 (2006).
Jurata, L.W. et al. Altered expression of hippocampal dentate granule neuron genes in a mouse model of human 22q11 deletion syndrome. Schizophr. Res. 88, 251–259 (2006).
Meechan, D. et al. Gene dosage in the developing and adult brain in a mouse model of 22q11 deletion syndrome. Mol. Cell. Neurosci. 33, 412–428 (2006).
Sivagnanasundaram, S. et al. Differential gene expression in the hippocampus of the Df1/+ mice: a model for 22q11.2 deletion syndrome and schizophrenia. Brain Res. 1139, 48–59 (2007).
Seitz, H. et al. A large imprinted microRNA gene cluster at the mouse Dlk1-Gtl2 domain. Genome Res. 14, 1741–1748 (2004).
Tomari, Y. & Zamore, P.D. MicroRNA biogenesis: drosha can't cut it without a partner. Curr. Biol. 15, R61–R64 (2005).
Wheeler, G., Ntounia-Fousara, S., Granda, B., Rathjen, T. & Dalmay, T. Identification of new central nervous system specific mouse microRNAs. FEBS Lett. 580, 2195–2200 (2006).
Kosik, K.S. The neuronal microRNA system. Nat. Rev. Neurosci. 7, 911–920 (2006).
Lewis, B.P., Shih, I.H., Jones-Rhoades, M.W., Bartel, D.P. & Burge, C.B. Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003).
Krek, A. et al. Combinatorial microRNA target predictions. Nat. Genet. 37, 495–500 (2005).
Paylor, R. & Crawley, J.N. Inbred strain differences in prepulse inhibition of the mouse startle response. Psychopharmacology (Berl.) 132, 169–180 (1997).
Schratt, G.M. et al. A brain-specific microRNA regulates dendritic spine development. Nature 439, 283–289 erratum 441, 902 (2006).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Matsuzaki, M., Honkura, N., Ellis-Davies, G.C. & Kasai, H. Structural basis of long-term potentiation in single dendritic spines. Nature 429, 761–766 (2004).
Arguello, P.A. & Gogos, J.A. Modeling madness in mice: one piece at a time. Neuron 52, 179–196 (2006).
Wang, Y., Medvid, R., Melton, C., Jaenisch, R. & Blelloch, R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat. Genet. 39, 380–385 (2007).
Zhao, Y. et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1–2. Cell 129, 303–317 (2007).
Doench, J.G., Petersen, C.P. & Sharp, P.A. siRNAs can function as miRNAs. Genes Dev. 17, 438–442 (2003).
Ashraf, S.I., McLoon, A.L., Sclarsic, S.M. & Kunes, S. Synaptic protein synthesis associated with memory is regulated by the RISC pathway in Drosophila. Cell 124, 191–205 (2006).
Caudy, A.A., Myers, M., Hannon, G.J. & Hammond, S.M. Fragile X-related protein and VIG associate with the RNA interference machinery. Genes Dev. 16, 2491–2496 (2002).
Acknowledgements
We thank A. Merién for assistance with the dendritic complexity and spine analysis. We thank A. Abrams-Downey, C. Frazier, J. Pellegrino, D. Swor, Y. Sun and M. Sribour for technical support and assistance with the mouse colony. We also thank P.A. Arguello for help and insights with the behavioral assays. This research was supported in part by the US National Institutes of Health (grants MH067068 to M.K. and J.A.G. and MH077235 to J.A.G.) and the New York Academy of Sciences (J.A.G.). Support was also provided in part by a McKnight Brain Disorders award (J.A.G.) and a NARSAD award (J.A.G.), as well as an EJLB grant award (J.A.G.).
Author information
Authors and Affiliations
Contributions
K.L.S., A.B., H.L. and A.A.M. contributed to the generation of the Df16(A)+/− line; K.L.S., B.X. and W.-S.L. performed the behavioral assays; B.X. and R.H. contributed to the generation and validation of the Dgcr8+/− line; B.X. performed the mRNA and miRNA expression analysis and the dendritic spine and complexity analysis; X.W. and P.P. contributed to the bioinformatic analysis of the mRNA expression data; M.K. and J.A.G. contributed to the research design and supervised the project.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–5, Supplementary Note and Supplementary Methods (PDF 1440 kb)
Rights and permissions
About this article
Cite this article
Stark, K., Xu, B., Bagchi, A. et al. Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat Genet 40, 751–760 (2008). https://doi.org/10.1038/ng.138
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.138
This article is cited by
-
DiGeorge syndrome critical region gene 2 (DGCR2), a schizophrenia risk gene, regulates dendritic spine development through cell adhesion
Cell & Bioscience (2023)
-
The molecular pathology of schizophrenia: an overview of existing knowledge and new directions for future research
Molecular Psychiatry (2023)
-
The synaptic hypothesis of schizophrenia version III: a master mechanism
Molecular Psychiatry (2023)
-
Altered corollary discharge signaling in the auditory cortex of a mouse model of schizophrenia predisposition
Nature Communications (2023)
-
The prediction-error hypothesis of schizophrenia: new data point to circuit-specific changes in dopamine activity
Neuropsychopharmacology (2022)