Abstract
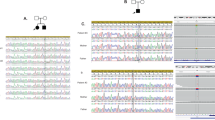
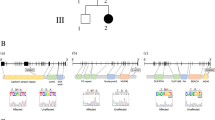
Paroxysmal kinesigenic dyskinesia is the most common type of paroxysmal movement disorder and is often misdiagnosed clinically as epilepsy. Using whole-exome sequencing followed by Sanger sequencing, we identified three truncating mutations within PRRT2 (NM_145239.2) in eight Han Chinese families with histories of paroxysmal kinesigenic dyskinesia: c.514_517delTCTG (p.Ser172Argfs*3) in one family, c.649dupC (p.Arg217Profs*8) in six families and c.972delA (p.Val325Serfs*12) in one family. These truncating mutations co-segregated exactly with the disease in these families and were not observed in 1,000 control subjects of matched ancestry. PRRT2 is a newly discovered gene consisting of four exons encoding the proline-rich transmembrane protein 2, which encompasses 340 amino acids and contains two predicted transmembrane domains. PRRT2 is highly expressed in the developing nervous system, and a truncating mutation alters the subcellular localization of the PRRT2 protein. The function of PRRT2 and its role in paroxysmal kinesigenic dyskinesia should be further investigated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Kertesz, A. Paroxysmal kinesigenic choreoathetosis. An entity within the paroxysmal choreoathetosis syndrome. Description of 10 cases, including 1 autopsied. Neurology 17, 680–690 (1967).
Bruno, M.K. et al. Clinical evaluation of idiopathic paroxysmal kinesigenic dyskinesia: new diagnostic criteria. Neurology 63, 2280–2287 (2004).
Goodenough, D.J., Fariello, R.G., Annis, B.L. & Chun, R.W. Familial and acquired paroxysmal dyskinesias. A proposed classification with delineation of clinical features. Arch. Neurol. 35, 827–831 (1978).
Tomita, H. et al. Paroxysmal kinesigenic choreoathetosis locus maps to chromosome 16p11.2-q12.1. Am. J. Hum. Genet. 65, 1688–1697 (1999).
Swoboda, K.J. et al. Paroxysmal kinesigenic dyskinesia and infantile convulsions: clinical and linkage studies. Neurology 55, 224–230 (2000).
Bennett, L.B., Roach, E.S. & Bowcock, A.M. A locus for paroxysmal kinesigenic dyskinesia maps to human chromosome 16. Neurology 54, 125–130 (2000).
Valente, E.M. et al. A second paroxysmal kinesigenic choreoathetosis locus (EKD2) mapping on 16q13-q22.1 indicates a family of genes which give rise to paroxysmal disorders on human chromosome 16. Brain 123, 2040–2045 (2000).
Kikuchi, T. et al. Paroxysmal kinesigenic choreoathetosis (PKC): confirmation of linkage to 16p11-q21, but unsuccessful detection of mutations among 157 genes at the PKC-critical region in seven PKC families. J. Hum. Genet. 52, 334–341 (2007).
Ono, S. et al. Mutation and copy number analysis in paroxysmal kinesigenic dyskinesia families. Mov. Disord. 26, 761–763 (2011).
Ng, S.B. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, 272–276 (2009).
Wang, J. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008).
Ng, P.C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Su, A.I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
Bhatia, K.P., Griggs, R.C. & Ptácek, L.J. Episodic movement disorders as channelopathies. Mov. Disord. 15, 429–433 (2000).
Celesia, G.G. Disorders of membrane channels or channelopathies. Clin. Neurophysiol. 112, 2–18 (2001).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Kruglyak, L., Daly, M.J., Reeve-Daly, M.P. & Lander, E.S. Parametric and nonparametric linkage analysis: a unified multipoint approach. Am. J. Hum. Genet. 58, 1347–1363 (1996).
Hu, X.L. et al. Conditional deletion of NRSF in forebrain neurons accelerates epileptogenesis in the kindling model. Cereb. Cortex 21, 2158–2165 (2011).
Acknowledgements
The authors sincerely thank the participants for their help and willingness to take part in this study. The authors also thank the Beijing Genomics Institute (BGI)-Shenzhen for assistance in the analysis of exome sequence data. This work was supported by a grant from the National Natural Science Foundation (China; 81125009 to Z.-Y.W.) and a grant from Huashan Hospital for the special professorship of Fudan University (to Z.-Y.W.) and by a key program of scientific research of Fujian Medical University (2009D064 to N.W.).
Author information
Authors and Affiliations
Contributions
N.W. and Z.-Y.W. planned the project. Z.-Q.X., J.X., N.W. and Z.-Y.W. conceived of and designed the study. W.-J.C., Yu Lin, Yi Lin, S.-X.M., N.W. and Z.-Y.W. performed the sample collection. W.-J.C., Yu Lin, W.W., W.N., J.H., Y.-F.C., Q.-J.Z. and H.-F.L. performed sequence analysis. J.-F.X. and Z.-Y.W. performed linkage and haplotype analyses. Z.-Q.X., G.-H.T. and S.-L.G. performed the expression analysis. W.-J.C., Z.-Q.X., J.X., N.W. and Z.-Y.W. analyzed the data. W.-J.C., Z.-Q.X. and Z.-Y.W. wrote the manuscript, and J.X. and Z.-Y.W. revised it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Tables 1 and 2 (PDF 1391 kb)
Rights and permissions
About this article
Cite this article
Chen, WJ., Lin, Y., Xiong, ZQ. et al. Exome sequencing identifies truncating mutations in PRRT2 that cause paroxysmal kinesigenic dyskinesia. Nat Genet 43, 1252–1255 (2011). https://doi.org/10.1038/ng.1008
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.1008
This article is cited by
-
Levetiracetam may be an unsuitable choice for patients with PRRT2-associated self-limited infantile epilepsy
BMC Pediatrics (2023)
-
A Push–Pull Mechanism Between PRRT2 and β4-subunit Differentially Regulates Membrane Exposure and Biophysical Properties of NaV1.2 Sodium Channels
Molecular Neurobiology (2023)
-
Paroxysmal Kinesigenic Dyskinesia: Genetics and Pathophysiological Mechanisms
Neuroscience Bulletin (2023)
-
Pain-causing stinging nettle toxins target TMEM233 to modulate NaV1.7 function
Nature Communications (2023)
-
Sequencing individual genomes with recurrent genomic disorder deletions: an approach to characterize genes for autosomal recessive rare disease traits
Genome Medicine (2022)