Abstract
Interactions between environmental and genetic factors are proposed to explain why autoimmunity afflicts certain individuals and not others. Genes and genetic loci predisposing to autoimmunity are being identified, but theories as to how the environment contributes to autoimmunity still rely largely on examples such as drug-induced systemic lupus erythematosus (SLE) and epidemiologic evidence of occupational exposure, without clear mechanistic explanations or identification of specific environmental agents. Eukaryotic gene expression requires not only transcription factor activation but also regional modification of chromatin structure into a transcriptionally permissive configuration through epigenetic mechanisms, including DNA methylation and histone modifications. The realization that epigenetic mechanisms can alter gene expression and, therefore, cellular function has led to new insights into how environmental agents might contribute to the development of diseases in genetically predisposed individuals. The observation that some SLE-inducing drugs, such as procainamide and hydralazine, affect T cell DNA methylation and thereby cellular function, and that identical changes in T cell DNA methylation and cellular function are found in patients with SLE, implicates epigenetic mechanisms in the pathogenesis of human SLE, and perhaps other autoimmune diseases. In this Review we discuss how epigenetic mechanisms affect gene expression, how environmental agents can affect epigenetic mechanisms, and how epigenetic changes in gene expression can contribute to autoimmunity. Similar mechanisms might also contribute to the pathogenesis of other poorly understood human diseases.
Key Points
-
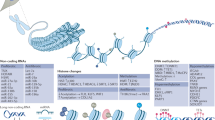
DNA is packaged as chromatin in eukaryotic cells, and regional chromatin structure is determined by epigenetic mechanisms including DNA methylation and histone modifications
-
Gene expression requires two events: activation of transcription factors to permit binding to their recognition sequence, and localized remodeling of chromatin to permit assembly of the transcription initiation machinery
-
Environmental factors can modify chromatin structure, causing changes in gene expression
-
The systemic-lupus-erythematosus-inducing drugs procainamide and hydralazine inhibit T cell DNA methylation, which alters gene expression to cause autoreactivity and a lupus-like disease in animal models
-
T cells from patients with active systemic lupus erythematosus have identical changes in DNA methylation patterns, gene expression and function as T cells treated with DNA methylation inhibitors
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Robertson KD (2005) DNA methylation and human disease. Nat Rev Genet 6: 597–610
Harley JB et al. (2006) Unraveling the genetics of systemic lupus erythematosus. Springer Semin Immunopathol 28: 119–130
Rao T and Richardson B (1999) Environmentally induced autoimmune diseases: potential mechanisms. Environ Health Perspect 107 (Suppl 5): 737–742
Cooper GS et al. (2004) Occupational risk factors for the development of systemic lupus erythematosus. J Rheumatol 31: 1928–1933
Järvinen P and Aho K (1994) Twin studies in rheumatic diseases. Semin Arthritis Rheum 24: 19–28
Felsenfeld G and Groudine M (2003) Controlling the double helix. Nature 421: 448–453
Bird AP and Wolffe AP (1999) Methylation-induced repression—belts, braces, and chromatin. Cell 99: 451–454
Antequera F and Bird A (1993) In DNA Methylation: Molecular Biology and Biologic Significance, 169–185 (eds Jost J and Saluz H) Boston: Birkhauser
Reik W et al. (2001) Epigenetic reprogramming in mammalian development. Science 293: 1089–1093
Carrel L and Willard HF (2005) X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 434: 400–404
Jaenisch R and Bird A (2003) Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet 33 (Suppl): 245–254
Ehrlich M (2003) The ICF syndrome, a DNA methyltransferase 3B deficiency and immunodeficiency disease. Clin Immunol 109: 17–28
Amir RE et al. (1999) Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat Genet 23: 185–188
Yoder JA et al. (1997) Cytosine methylation and the ecology of intragenomic parasites. Trends Genet 13: 335–340
Robertson KD (2000) The role of DNA methylation in modulating Epstein–Barr virus gene expression. Curr Top Microbiol Immunol 249: 21–34
Reiner SL (2005) Epigenetic control in the immune response. Hum Mol Genet 14 (Spec No 1): R41–46
Toyota M et al. (1999) CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA 96: 8681–8686
Ehrlich M (2002) DNA methylation in cancer: too much, but also too little. Oncogene 21: 5400–5413
Richardson B (2003) Impact of aging on DNA methylation. Ageing Res Rev 2: 245–261
Tra J et al. (2002) Infrequent occurrence of age-dependent changes in CpG island methylation as detected by restriction landmark genome scanning. Mech Ageing Dev 123: 1487–1503
Golbus J et al. (1990) Quantitative changes in T cell DNA methylation occur during differentiation and ageing. Eur J Immunol 20: 1869–1872
Zhang Z et al. (2002) Age-dependent DNA methylation changes in the ITGAL (CD11a) promoter. Mech Ageing Dev 123: 1257–1268
Bruniquel D and Schwartz RH (2003) Selective, stable demethylation of the interleukin-2 gene enhances transcription by an active process. Nat Immunol 4: 235–240
Barreto G et al. (2007) Gadd45a promotes epigenetic gene activation by repair-mediated DNA demethylation. Nature 445: 671–675
Ingrosso D et al. (2003) Folate treatment and unbalanced methylation and changes of allelic expression induced by hyperhomocysteinaemia in patients with uraemia. Lancet 361: 1693–1699
Niculescu MD and Zeisel SH (2002) Diet, methyl donors and DNA methylation: interactions between dietary folate, methionine and choline. J Nutr 132: 2333S–2335S
Deng C et al. (1998) Role of the ras-MAPK signaling pathway in the DNA methyltransferase response to DNA hypomethylation. Biol Chem 379: 1113–1120
MacLeod AR et al. (1995) Regulation of DNA methylation by the Ras signaling pathway. J Biol Chem 270: 11327–11337
Lieberman MW et al. (1983) Ultraviolet radiation-induced metallothionein-I gene activation is associated with extensive DNA demethylation. Cell 35: 207–214
Li-Weber M et al. (2005) Ultraviolet irradiation suppresses T cell activation via blocking TCR-mediated ERK and NF-kappa B signaling pathways. J Immunol 175: 2132–2143
Lu Q et al. (2005) Demethylation of the same promoter sequence increases CD70 expression in lupus T cells and T cells treated with lupus-inducing drugs. J Immunol 174: 6212–6219
Cooper GS and Stroehla BC (2003) The epidemiology of autoimmune diseases. Autoimmun Rev 2: 119–125
Richardson B (1986) Effect of an inhibitor of DNA methylation on T cells. II. 5-Azacytidine induces self-reactivity in antigen-specific T4+ cells. Hum Immunol 17: 456–470
Richardson BC et al. (1992) Phenotypic and functional similarities between 5-azacytidine-treated T cells and a T cell subset in patients with active systemic lupus erythematosus. Arthritis Rheum 35: 647–662
Lu Q et al. (2002) Demethylation of ITGAL (CD11a) regulatory sequences in systemic lupus erythematosus. Arthritis Rheum 46: 1282–1291
Yung R et al. (1996) Mechanisms of drug-induced lupus. II. T cells overexpressing lymphocyte function-associated antigen 1 become autoreactive and cause a lupuslike disease in syngeneic mice. J Clin Invest 97: 2866–2871
van der Veen FM et al. (1982) Autoimmune disease strongly resembling systemic lupus erythematosus (SLE) in F1 mice undergoing graft-versus-host reaction (GVHR). Adv Exp Med Biol 149: 669–677
Kaplan MJ et al. (2002) The apoptotic ligands TRAIL, TWEAK, and Fas ligand mediate monocyte death induced by autologous lupus T cells. J Immunol 169: 6020–6029
Lu Q et al. (2003) DNA methylation and chromatin structure regulate T cell perforin gene expression. J Immunol 170: 5124–5132
Kaplan MJ et al. (2004) Demethylation of promoter regulatory elements contributes to perforin overexpression in CD4+ lupus T cells. J Immunol 172: 3652–3661
Denny MF et al. (2006) Accelerated macrophage apoptosis induces autoantibody formation and organ damage in systemic lupus erythematosus. J Immunol 176: 2095–2104
Oelke K et al. (2004) Overexpression of CD70 and overstimulation of IgG synthesis by lupus T cells and T cells treated with DNA methylation inhibitors. Arthritis Rheum 50: 1850–1860
Lu Q et al. (2005) Demethylation of the same promoter sequence increases CD70 expression in lupus T cells and T cells treated with lupus-inducing drugs. J Immunol 174: 6212–6219
Richardson B et al. (2004) In Autoimmunity. Methods and Protocols, 285–294 (ed Perl A) Totowa, NJ: Humana
Richardson B et al. (1990) Evidence for impaired T cell DNA methylation in systemic lupus erythematosus and rheumatoid arthritis. Arthritis Rheum 33: 1665–1673
Deng C et al. (2001) Decreased Ras-mitogen-activated protein kinase signaling may cause DNA hypomethylation in T lymphocytes from lupus patients. Arthritis Rheum 44: 397–407
Deng C et al. (2003) Hydralazine may induce autoimmunity by inhibiting extracellular signal-regulated kinase pathway signaling. Arthritis Rheum 48: 746–756
Park BL et al. (2004). Association analyses of DNA methyltransferase-1 (DNMT1) polymorphisms with systemic lupus erythematosus. J Hum Genet 49: 642–646
Ashkar AA and Rosenthal KL (2002) Toll-like receptor 9, CpG DNA and innate immunity. Curr Mol Med 2: 545–556
Perl A (2003) Role of endogenous retroviruses in autoimmune diseases. Rheum Dis Clin North Am 29: 123–143
Wang Y et al. (2006) Association between enhanced type I collagen expression and epigenetic repression of the FLI1 gene in scleroderma fibroblasts. Arthritis Rheum 54: 2271–2279
Acknowledgements
The author thanks Dr Amr Sawalha for assistance with illustrations, and Ms Cindy Bourke for her expert secretarial assistance. This work was supported by PHS grants and a Merit grant from the Veterans Administration.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Rights and permissions
About this article
Cite this article
Richardson, B. Primer: epigenetics of autoimmunity. Nat Rev Rheumatol 3, 521–527 (2007). https://doi.org/10.1038/ncprheum0573
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ncprheum0573
This article is cited by
-
The multifaceted functional role of DNA methylation in immune-mediated rheumatic diseases
Clinical Rheumatology (2021)
-
The emerging role of epigenetics in human autoimmune disorders
Clinical Epigenetics (2019)
-
Analysis of methylation datasets identified significantly changed genes and functional pathways in osteoarthritis
Clinical Rheumatology (2019)
-
Translating epigenetics into clinic: focus on lupus
Clinical Epigenetics (2017)
-
The role of epigenetics in personalized medicine: challenges and opportunities
BMC Medical Genomics (2015)